grnsight 6.0.4 → 6.0.7
This diff represents the content of publicly available package versions that have been released to one of the supported registries. The information contained in this diff is provided for informational purposes only and reflects changes between package versions as they appear in their respective public registries.
- package/database/README.md +40 -16
- package/database/expression-database/README.md +2 -2
- package/database/expression-database/schema.sql +14 -14
- package/database/expression-database/scripts/loader.py +6 -6
- package/database/network-database/README.md +36 -4
- package/database/network-database/schema.sql +7 -7
- package/database/network-database/scripts/filter_genes.py +1 -1
- package/database/network-database/scripts/generate_network.py +1 -1
- package/database/network-database/scripts/generate_new_network_version.py +223 -0
- package/database/network-database/scripts/loader.py +3 -3
- package/database/network-database/scripts/loader_updates.py +99 -0
- package/package.json +2 -2
- package/server/controllers/custom-workbook-controller.js +4 -3
- package/server/dals/expression-dal.js +8 -7
- package/server/dals/network-dal.js +3 -3
- package/web-client/public/js/api/grnsight-api.js +56 -55
- package/web-client/public/js/generateNetwork.js +15 -7
- package/web-client/public/js/setup-load-and-import-handlers.js +171 -170
- package/web-client/public/js/upload.js +74 -95
- package/.bundle/config +0 -3
- package/GRNsight - Beta.html +0 -194
- package/Gemfile.lock +0 -259
- package/_gh_pages/about.html +0 -807
- package/_gh_pages/assets/css/bootstrap.min.css +0 -10
- package/_gh_pages/assets/css/footer.css +0 -3
- package/_gh_pages/assets/css/main.css +0 -363
- package/_gh_pages/assets/images/21-genes_31-edges_Schade-data_estimation_output_binary-no-targetless-genes_sif.png +0 -0
- package/_gh_pages/assets/images/21-genes_31-edges_Schade-data_estimation_output_binary_sif.png +0 -0
- package/_gh_pages/assets/images/21-genes_31-edges_Schade-data_graphml_3-edges-and-footer.png +0 -0
- package/_gh_pages/assets/images/21-genes_31-edges_Schade-data_graphml_header-and-3-nodes.png +0 -0
- package/_gh_pages/assets/images/21-genes_31-edges_Schade-data_graphml_output_3-edges-and-footer.png +0 -0
- package/_gh_pages/assets/images/21-genes_31-edges_Schade-data_graphml_output_header-and-3-nodes.png +0 -0
- package/_gh_pages/assets/images/21-genes_31-edges_Schade-data_input_binary-no-targetless-genes_sif.png +0 -0
- package/_gh_pages/assets/images/21-genes_31-edges_Schade-data_input_binary_sif.png +0 -0
- package/_gh_pages/assets/images/21-genes_31-edges_Schade-data_input_concatenated-no-targetless-genes_sif.png +0 -0
- package/_gh_pages/assets/images/21-genes_31-edges_Schade-data_input_concatenated_sif.png +0 -0
- package/_gh_pages/assets/images/Carrillo_Roque_LMU_Symposium_2015.pptx +0 -0
- package/_gh_pages/assets/images/Choe-Shin_CMSI402-poster-session_20180430.jpg +0 -0
- package/_gh_pages/assets/images/Choe_SCCUR_2017.jpg +0 -0
- package/_gh_pages/assets/images/Dahlquist-BMB-2015-Fig1.jpg +0 -0
- package/_gh_pages/assets/images/Dahlquist-BMB-2015-Fig8.jpg +0 -0
- package/_gh_pages/assets/images/Dahlquist-Choe-Shin_CMSI402-poster-session_20180430.jpg +0 -0
- package/_gh_pages/assets/images/Demo-3-auto-layout.jpg +0 -0
- package/_gh_pages/assets/images/Demo-3-paper-layout.jpg +0 -0
- package/_gh_pages/assets/images/Demo-4-auto-layout.jpg +0 -0
- package/_gh_pages/assets/images/Demo-4-paper-layout.jpg +0 -0
- package/_gh_pages/assets/images/Dionisio-Dahlquist_GRNsight-shades_20170506.jpg +0 -0
- package/_gh_pages/assets/images/GRNsight_logo_20140710_main.jpg +0 -0
- package/_gh_pages/assets/images/GRNsight_logo_20140710_main_resized.jpg +0 -0
- package/_gh_pages/assets/images/GRNsight_logo_20140710_rollover.jpg +0 -0
- package/_gh_pages/assets/images/GRNsight_logo_20140710_rollover_resized.jpg +0 -0
- package/_gh_pages/assets/images/Johnson_Williams_LMU_Symposium_2015.pptx +0 -0
- package/_gh_pages/assets/images/Klein_Samdarshi_TriBeta_2018_20180317.jpg +0 -0
- package/_gh_pages/assets/images/LMULogoHQ2.png +0 -0
- package/_gh_pages/assets/images/LMUSymposium_Group_20150321-2.jpg +0 -0
- package/_gh_pages/assets/images/LMUSymposium_Group_20150321-2_resized.jpg +0 -0
- package/_gh_pages/assets/images/Sample_network_optimized_weights_worksheet.jpg +0 -0
- package/_gh_pages/assets/images/Sample_network_worksheet.jpg +0 -0
- package/_gh_pages/assets/images/Shin_SCCUR_2017.jpg +0 -0
- package/_gh_pages/assets/images/demo-3_network-sheet.png +0 -0
- package/_gh_pages/assets/images/demo-4_network-optimized-weights-sheet.png +0 -0
- package/_gh_pages/assets/images/gene-pages-0.png +0 -0
- package/_gh_pages/assets/images/gene-pages-1.png +0 -0
- package/_gh_pages/assets/images/gene-pages-2.png +0 -0
- package/_gh_pages/assets/images/gene-pages-3.png +0 -0
- package/_gh_pages/assets/images/grnsight2020.png +0 -0
- package/_gh_pages/assets/images/lmu logo.gif +0 -0
- package/_gh_pages/assets/images/v3demo2-grid+nodecoloring.png +0 -0
- package/_gh_pages/assets/images/v3demo2-nodecoloring.png +0 -0
- package/_gh_pages/assets/images/v3demo2.png +0 -0
- package/_gh_pages/assets/js/ga-report.js +0 -35
- package/_gh_pages/assets/js/iframeResizer.min.js +0 -9
- package/_gh_pages/assets/js/main.js +0 -132
- package/_gh_pages/beta.html +0 -144
- package/_gh_pages/contact.html +0 -178
- package/_gh_pages/coverage/coverage.json +0 -1
- package/_gh_pages/coverage/coverage.raw.json +0 -1
- package/_gh_pages/coverage/lcov-report/base.css +0 -223
- package/_gh_pages/coverage/lcov-report/block-navigation.js +0 -63
- package/_gh_pages/coverage/lcov-report/controllers/additional-sheet-parser.js.html +0 -330
- package/_gh_pages/coverage/lcov-report/controllers/constants.js.html +0 -243
- package/_gh_pages/coverage/lcov-report/controllers/export-controller.js.html +0 -285
- package/_gh_pages/coverage/lcov-report/controllers/exporters/graphml.js.html +0 -405
- package/_gh_pages/coverage/lcov-report/controllers/exporters/index.html +0 -110
- package/_gh_pages/coverage/lcov-report/controllers/exporters/sif.js.html +0 -150
- package/_gh_pages/coverage/lcov-report/controllers/helpers.js.html +0 -114
- package/_gh_pages/coverage/lcov-report/controllers/import-controller.js.html +0 -233
- package/_gh_pages/coverage/lcov-report/controllers/importers/graphml.js.html +0 -716
- package/_gh_pages/coverage/lcov-report/controllers/importers/index.html +0 -106
- package/_gh_pages/coverage/lcov-report/controllers/importers/sif.js.html +0 -488
- package/_gh_pages/coverage/lcov-report/controllers/index.html +0 -162
- package/_gh_pages/coverage/lcov-report/controllers/semantic-checker.js.html +0 -810
- package/_gh_pages/coverage/lcov-report/controllers/spreadsheet-controller.js.html +0 -1779
- package/_gh_pages/coverage/lcov-report/index.html +0 -136
- package/_gh_pages/coverage/lcov-report/prettify.css +0 -1
- package/_gh_pages/coverage/lcov-report/prettify.js +0 -1
- package/_gh_pages/coverage/lcov-report/server/controllers/additional-sheet-parser.js.html +0 -330
- package/_gh_pages/coverage/lcov-report/server/controllers/constants.js.html +0 -243
- package/_gh_pages/coverage/lcov-report/server/controllers/export-controller.js.html +0 -285
- package/_gh_pages/coverage/lcov-report/server/controllers/exporters/graphml.js.html +0 -405
- package/_gh_pages/coverage/lcov-report/server/controllers/exporters/index.html +0 -110
- package/_gh_pages/coverage/lcov-report/server/controllers/exporters/sif.js.html +0 -150
- package/_gh_pages/coverage/lcov-report/server/controllers/graphml-constants.js.html +0 -585
- package/_gh_pages/coverage/lcov-report/server/controllers/helpers.js.html +0 -114
- package/_gh_pages/coverage/lcov-report/server/controllers/import-controller.js.html +0 -237
- package/_gh_pages/coverage/lcov-report/server/controllers/importers/graphml.js.html +0 -585
- package/_gh_pages/coverage/lcov-report/server/controllers/importers/index.html +0 -110
- package/_gh_pages/coverage/lcov-report/server/controllers/importers/sif.js.html +0 -492
- package/_gh_pages/coverage/lcov-report/server/controllers/index.html +0 -188
- package/_gh_pages/coverage/lcov-report/server/controllers/semantic-checker.js.html +0 -810
- package/_gh_pages/coverage/lcov-report/server/controllers/spreadsheet-controller.js.html +0 -1779
- package/_gh_pages/coverage/lcov-report/sort-arrow-sprite.png +0 -0
- package/_gh_pages/coverage/lcov-report/sorter.js +0 -158
- package/_gh_pages/coverage/lcov-report/web-client/public/js/grnstate.js.html +0 -225
- package/_gh_pages/coverage/lcov-report/web-client/public/js/index.html +0 -97
- package/_gh_pages/coverage/lcov.info +0 -49
- package/_gh_pages/documentation.html +0 -1132
- package/_gh_pages/documents/abstracts/SIGGRAPH 2017 Abstract/siggraph-abstract-review.aux +0 -47
- package/_gh_pages/documents/abstracts/SIGGRAPH 2017 Abstract/siggraph-abstract-review.bbl +0 -73
- package/_gh_pages/documents/abstracts/SIGGRAPH 2017 Abstract/siggraph-abstract-review.blg +0 -52
- package/_gh_pages/documents/abstracts/SIGGRAPH 2017 Abstract/siggraph-abstract-review.log +0 -1056
- package/_gh_pages/documents/abstracts/SIGGRAPH 2017 Abstract/siggraph-abstract-review.out +0 -7
- package/_gh_pages/documents/abstracts/SIGGRAPH 2017 Abstract/siggraph-abstract-review.synctex.gz +0 -0
- package/_gh_pages/documents/manuscripts/peerj-computerscience-2016/revisions/GRNsight_PeerJ-CS_manuscript_2016_text-only_revised-Dondi.docx +0 -0
- package/_gh_pages/encryption/server.cert +0 -21
- package/_gh_pages/encryption/server.key +0 -28
- package/_gh_pages/favicon.ico +0 -0
- package/_gh_pages/googlea1a858c5bc91c87e.html +0 -1
- package/_gh_pages/index.html +0 -163
- package/_gh_pages/links.html +0 -177
- package/_gh_pages/news.html +0 -367
- package/_gh_pages/onlyfooter.html +0 -78
- package/_gh_pages/onlyheader.html +0 -64
- package/_gh_pages/onlysidebar.html +0 -73
- package/_gh_pages/package-lock.json +0 -14048
- package/_gh_pages/params.json +0 -1
- package/_gh_pages/people.html +0 -260
- package/_gh_pages/privacy.html +0 -147
- package/_gh_pages/publications.html +0 -378
- package/_gh_pages/robots.txt +0 -1
- package/_gh_pages/server/API Project-7d45e9f24a96.p12 +0 -0
- package/_gh_pages/server/grnmap.pem +0 -31
- package/_gh_pages/server/grnsight.pem +0 -19
- package/_gh_pages/sitemap.xml +0 -174
- package/_gh_pages/test-files/import-samples/attributes.graphml +0 -40
- package/_gh_pages/test-files/import-samples/port.graphml +0 -32
- package/_gh_pages/test-files/import-samples/simple.graphml +0 -31
- package/_gh_pages/web-client/public/js/grnsight.min.js +0 -2347
- package/_gh_pages/web-client/public/stylesheets/grnsight.css +0 -443
- package/coverage/coverage.json +0 -1
- package/coverage/coverage.raw.json +0 -1
- package/coverage/lcov-report/base.css +0 -223
- package/coverage/lcov-report/block-navigation.js +0 -63
- package/coverage/lcov-report/controllers/additional-sheet-parser.js.html +0 -330
- package/coverage/lcov-report/controllers/constants.js.html +0 -243
- package/coverage/lcov-report/controllers/export-controller.js.html +0 -285
- package/coverage/lcov-report/controllers/exporters/graphml.js.html +0 -405
- package/coverage/lcov-report/controllers/exporters/index.html +0 -110
- package/coverage/lcov-report/controllers/exporters/sif.js.html +0 -150
- package/coverage/lcov-report/controllers/helpers.js.html +0 -114
- package/coverage/lcov-report/controllers/import-controller.js.html +0 -233
- package/coverage/lcov-report/controllers/importers/graphml.js.html +0 -716
- package/coverage/lcov-report/controllers/importers/index.html +0 -106
- package/coverage/lcov-report/controllers/importers/sif.js.html +0 -488
- package/coverage/lcov-report/controllers/index.html +0 -162
- package/coverage/lcov-report/controllers/semantic-checker.js.html +0 -810
- package/coverage/lcov-report/controllers/spreadsheet-controller.js.html +0 -1779
- package/coverage/lcov-report/index.html +0 -136
- package/coverage/lcov-report/prettify.css +0 -1
- package/coverage/lcov-report/prettify.js +0 -1
- package/coverage/lcov-report/server/controllers/additional-sheet-parser.js.html +0 -330
- package/coverage/lcov-report/server/controllers/constants.js.html +0 -243
- package/coverage/lcov-report/server/controllers/export-controller.js.html +0 -285
- package/coverage/lcov-report/server/controllers/exporters/graphml.js.html +0 -405
- package/coverage/lcov-report/server/controllers/exporters/index.html +0 -110
- package/coverage/lcov-report/server/controllers/exporters/sif.js.html +0 -150
- package/coverage/lcov-report/server/controllers/graphml-constants.js.html +0 -585
- package/coverage/lcov-report/server/controllers/helpers.js.html +0 -114
- package/coverage/lcov-report/server/controllers/import-controller.js.html +0 -237
- package/coverage/lcov-report/server/controllers/importers/graphml.js.html +0 -585
- package/coverage/lcov-report/server/controllers/importers/index.html +0 -110
- package/coverage/lcov-report/server/controllers/importers/sif.js.html +0 -492
- package/coverage/lcov-report/server/controllers/index.html +0 -188
- package/coverage/lcov-report/server/controllers/semantic-checker.js.html +0 -810
- package/coverage/lcov-report/server/controllers/spreadsheet-controller.js.html +0 -1779
- package/coverage/lcov-report/sort-arrow-sprite.png +0 -0
- package/coverage/lcov-report/sorter.js +0 -158
- package/coverage/lcov-report/web-client/public/js/grnstate.js.html +0 -225
- package/coverage/lcov-report/web-client/public/js/index.html +0 -97
- package/coverage/lcov.info +0 -1952
- package/documents/abstracts/SIGGRAPH 2017 Abstract/siggraph-abstract-review.aux +0 -47
- package/documents/abstracts/SIGGRAPH 2017 Abstract/siggraph-abstract-review.bbl +0 -73
- package/documents/abstracts/SIGGRAPH 2017 Abstract/siggraph-abstract-review.blg +0 -52
- package/documents/abstracts/SIGGRAPH 2017 Abstract/siggraph-abstract-review.log +0 -1056
- package/documents/abstracts/SIGGRAPH 2017 Abstract/siggraph-abstract-review.out +0 -7
- package/documents/abstracts/SIGGRAPH 2017 Abstract/siggraph-abstract-review.synctex.gz +0 -0
- package/documents/manuscripts/peerj-computerscience-2016/revisions/GRNsight_PeerJ-CS_manuscript_2016_text-only_revised-Dondi.docx +0 -0
- package/encryption/server.cert +0 -21
- package/encryption/server.key +0 -28
- package/gdb/PG_VERSION +0 -1
- package/gdb/base/1/112 +0 -0
- package/gdb/base/1/113 +0 -0
- package/gdb/base/1/1247 +0 -0
- package/gdb/base/1/1247_fsm +0 -0
- package/gdb/base/1/1247_vm +0 -0
- package/gdb/base/1/1249 +0 -0
- package/gdb/base/1/1249_fsm +0 -0
- package/gdb/base/1/1249_vm +0 -0
- package/gdb/base/1/1255 +0 -0
- package/gdb/base/1/1255_fsm +0 -0
- package/gdb/base/1/1255_vm +0 -0
- package/gdb/base/1/1259 +0 -0
- package/gdb/base/1/1259_fsm +0 -0
- package/gdb/base/1/1259_vm +0 -0
- package/gdb/base/1/13852 +0 -0
- package/gdb/base/1/13852_fsm +0 -0
- package/gdb/base/1/13852_vm +0 -0
- package/gdb/base/1/13855 +0 -0
- package/gdb/base/1/13856 +0 -0
- package/gdb/base/1/13857 +0 -0
- package/gdb/base/1/13857_fsm +0 -0
- package/gdb/base/1/13857_vm +0 -0
- package/gdb/base/1/13860 +0 -0
- package/gdb/base/1/13861 +0 -0
- package/gdb/base/1/13862 +0 -0
- package/gdb/base/1/13862_fsm +0 -0
- package/gdb/base/1/13862_vm +0 -0
- package/gdb/base/1/13865 +0 -0
- package/gdb/base/1/13866 +0 -0
- package/gdb/base/1/13867 +0 -0
- package/gdb/base/1/13867_fsm +0 -0
- package/gdb/base/1/13867_vm +0 -0
- package/gdb/base/1/13870 +0 -0
- package/gdb/base/1/13871 +0 -0
- package/gdb/base/1/1417 +0 -0
- package/gdb/base/1/1418 +0 -0
- package/gdb/base/1/174 +0 -0
- package/gdb/base/1/175 +0 -0
- package/gdb/base/1/2187 +0 -0
- package/gdb/base/1/2224 +0 -0
- package/gdb/base/1/2228 +0 -0
- package/gdb/base/1/2328 +0 -0
- package/gdb/base/1/2336 +0 -0
- package/gdb/base/1/2337 +0 -0
- package/gdb/base/1/2579 +0 -0
- package/gdb/base/1/2600 +0 -0
- package/gdb/base/1/2600_fsm +0 -0
- package/gdb/base/1/2600_vm +0 -0
- package/gdb/base/1/2601 +0 -0
- package/gdb/base/1/2601_fsm +0 -0
- package/gdb/base/1/2601_vm +0 -0
- package/gdb/base/1/2602 +0 -0
- package/gdb/base/1/2602_fsm +0 -0
- package/gdb/base/1/2602_vm +0 -0
- package/gdb/base/1/2603 +0 -0
- package/gdb/base/1/2603_fsm +0 -0
- package/gdb/base/1/2603_vm +0 -0
- package/gdb/base/1/2604 +0 -0
- package/gdb/base/1/2605 +0 -0
- package/gdb/base/1/2605_fsm +0 -0
- package/gdb/base/1/2605_vm +0 -0
- package/gdb/base/1/2606 +0 -0
- package/gdb/base/1/2606_fsm +0 -0
- package/gdb/base/1/2606_vm +0 -0
- package/gdb/base/1/2607 +0 -0
- package/gdb/base/1/2607_fsm +0 -0
- package/gdb/base/1/2607_vm +0 -0
- package/gdb/base/1/2608 +0 -0
- package/gdb/base/1/2608_fsm +0 -0
- package/gdb/base/1/2608_vm +0 -0
- package/gdb/base/1/2609 +0 -0
- package/gdb/base/1/2609_fsm +0 -0
- package/gdb/base/1/2609_vm +0 -0
- package/gdb/base/1/2610 +0 -0
- package/gdb/base/1/2610_fsm +0 -0
- package/gdb/base/1/2610_vm +0 -0
- package/gdb/base/1/2611 +0 -0
- package/gdb/base/1/2612 +0 -0
- package/gdb/base/1/2612_fsm +0 -0
- package/gdb/base/1/2612_vm +0 -0
- package/gdb/base/1/2613 +0 -0
- package/gdb/base/1/2615 +0 -0
- package/gdb/base/1/2615_fsm +0 -0
- package/gdb/base/1/2615_vm +0 -0
- package/gdb/base/1/2616 +0 -0
- package/gdb/base/1/2616_fsm +0 -0
- package/gdb/base/1/2616_vm +0 -0
- package/gdb/base/1/2617 +0 -0
- package/gdb/base/1/2617_fsm +0 -0
- package/gdb/base/1/2617_vm +0 -0
- package/gdb/base/1/2618 +0 -0
- package/gdb/base/1/2618_fsm +0 -0
- package/gdb/base/1/2618_vm +0 -0
- package/gdb/base/1/2619 +0 -0
- package/gdb/base/1/2619_fsm +0 -0
- package/gdb/base/1/2619_vm +0 -0
- package/gdb/base/1/2620 +0 -0
- package/gdb/base/1/2650 +0 -0
- package/gdb/base/1/2651 +0 -0
- package/gdb/base/1/2652 +0 -0
- package/gdb/base/1/2653 +0 -0
- package/gdb/base/1/2654 +0 -0
- package/gdb/base/1/2655 +0 -0
- package/gdb/base/1/2656 +0 -0
- package/gdb/base/1/2657 +0 -0
- package/gdb/base/1/2658 +0 -0
- package/gdb/base/1/2659 +0 -0
- package/gdb/base/1/2660 +0 -0
- package/gdb/base/1/2661 +0 -0
- package/gdb/base/1/2662 +0 -0
- package/gdb/base/1/2663 +0 -0
- package/gdb/base/1/2664 +0 -0
- package/gdb/base/1/2665 +0 -0
- package/gdb/base/1/2666 +0 -0
- package/gdb/base/1/2667 +0 -0
- package/gdb/base/1/2668 +0 -0
- package/gdb/base/1/2669 +0 -0
- package/gdb/base/1/2670 +0 -0
- package/gdb/base/1/2673 +0 -0
- package/gdb/base/1/2673_fsm +0 -0
- package/gdb/base/1/2674 +0 -0
- package/gdb/base/1/2674_fsm +0 -0
- package/gdb/base/1/2675 +0 -0
- package/gdb/base/1/2678 +0 -0
- package/gdb/base/1/2679 +0 -0
- package/gdb/base/1/2680 +0 -0
- package/gdb/base/1/2681 +0 -0
- package/gdb/base/1/2682 +0 -0
- package/gdb/base/1/2683 +0 -0
- package/gdb/base/1/2684 +0 -0
- package/gdb/base/1/2685 +0 -0
- package/gdb/base/1/2686 +0 -0
- package/gdb/base/1/2687 +0 -0
- package/gdb/base/1/2688 +0 -0
- package/gdb/base/1/2689 +0 -0
- package/gdb/base/1/2690 +0 -0
- package/gdb/base/1/2691 +0 -0
- package/gdb/base/1/2692 +0 -0
- package/gdb/base/1/2693 +0 -0
- package/gdb/base/1/2696 +0 -0
- package/gdb/base/1/2699 +0 -0
- package/gdb/base/1/2701 +0 -0
- package/gdb/base/1/2702 +0 -0
- package/gdb/base/1/2703 +0 -0
- package/gdb/base/1/2704 +0 -0
- package/gdb/base/1/2753 +0 -0
- package/gdb/base/1/2753_fsm +0 -0
- package/gdb/base/1/2753_vm +0 -0
- package/gdb/base/1/2754 +0 -0
- package/gdb/base/1/2755 +0 -0
- package/gdb/base/1/2756 +0 -0
- package/gdb/base/1/2757 +0 -0
- package/gdb/base/1/2830 +0 -0
- package/gdb/base/1/2831 +0 -0
- package/gdb/base/1/2832 +0 -0
- package/gdb/base/1/2833 +0 -0
- package/gdb/base/1/2834 +0 -0
- package/gdb/base/1/2835 +0 -0
- package/gdb/base/1/2836 +0 -0
- package/gdb/base/1/2836_fsm +0 -0
- package/gdb/base/1/2836_vm +0 -0
- package/gdb/base/1/2837 +0 -0
- package/gdb/base/1/2838 +0 -0
- package/gdb/base/1/2838_fsm +0 -0
- package/gdb/base/1/2838_vm +0 -0
- package/gdb/base/1/2839 +0 -0
- package/gdb/base/1/2840 +0 -0
- package/gdb/base/1/2840_fsm +0 -0
- package/gdb/base/1/2840_vm +0 -0
- package/gdb/base/1/2841 +0 -0
- package/gdb/base/1/2995 +0 -0
- package/gdb/base/1/2996 +0 -0
- package/gdb/base/1/3079 +0 -0
- package/gdb/base/1/3079_fsm +0 -0
- package/gdb/base/1/3079_vm +0 -0
- package/gdb/base/1/3080 +0 -0
- package/gdb/base/1/3081 +0 -0
- package/gdb/base/1/3085 +0 -0
- package/gdb/base/1/3118 +0 -0
- package/gdb/base/1/3119 +0 -0
- package/gdb/base/1/3164 +0 -0
- package/gdb/base/1/3256 +0 -0
- package/gdb/base/1/3257 +0 -0
- package/gdb/base/1/3258 +0 -0
- package/gdb/base/1/3350 +0 -0
- package/gdb/base/1/3351 +0 -0
- package/gdb/base/1/3379 +0 -0
- package/gdb/base/1/3380 +0 -0
- package/gdb/base/1/3381 +0 -0
- package/gdb/base/1/3394 +0 -0
- package/gdb/base/1/3394_fsm +0 -0
- package/gdb/base/1/3394_vm +0 -0
- package/gdb/base/1/3395 +0 -0
- package/gdb/base/1/3429 +0 -0
- package/gdb/base/1/3430 +0 -0
- package/gdb/base/1/3431 +0 -0
- package/gdb/base/1/3433 +0 -0
- package/gdb/base/1/3439 +0 -0
- package/gdb/base/1/3440 +0 -0
- package/gdb/base/1/3455 +0 -0
- package/gdb/base/1/3456 +0 -0
- package/gdb/base/1/3456_fsm +0 -0
- package/gdb/base/1/3456_vm +0 -0
- package/gdb/base/1/3466 +0 -0
- package/gdb/base/1/3467 +0 -0
- package/gdb/base/1/3468 +0 -0
- package/gdb/base/1/3501 +0 -0
- package/gdb/base/1/3502 +0 -0
- package/gdb/base/1/3503 +0 -0
- package/gdb/base/1/3534 +0 -0
- package/gdb/base/1/3541 +0 -0
- package/gdb/base/1/3541_fsm +0 -0
- package/gdb/base/1/3541_vm +0 -0
- package/gdb/base/1/3542 +0 -0
- package/gdb/base/1/3574 +0 -0
- package/gdb/base/1/3575 +0 -0
- package/gdb/base/1/3576 +0 -0
- package/gdb/base/1/3596 +0 -0
- package/gdb/base/1/3597 +0 -0
- package/gdb/base/1/3598 +0 -0
- package/gdb/base/1/3599 +0 -0
- package/gdb/base/1/3600 +0 -0
- package/gdb/base/1/3600_fsm +0 -0
- package/gdb/base/1/3600_vm +0 -0
- package/gdb/base/1/3601 +0 -0
- package/gdb/base/1/3601_fsm +0 -0
- package/gdb/base/1/3601_vm +0 -0
- package/gdb/base/1/3602 +0 -0
- package/gdb/base/1/3602_fsm +0 -0
- package/gdb/base/1/3602_vm +0 -0
- package/gdb/base/1/3603 +0 -0
- package/gdb/base/1/3603_fsm +0 -0
- package/gdb/base/1/3603_vm +0 -0
- package/gdb/base/1/3604 +0 -0
- package/gdb/base/1/3605 +0 -0
- package/gdb/base/1/3606 +0 -0
- package/gdb/base/1/3607 +0 -0
- package/gdb/base/1/3608 +0 -0
- package/gdb/base/1/3609 +0 -0
- package/gdb/base/1/3712 +0 -0
- package/gdb/base/1/3764 +0 -0
- package/gdb/base/1/3764_fsm +0 -0
- package/gdb/base/1/3764_vm +0 -0
- package/gdb/base/1/3766 +0 -0
- package/gdb/base/1/3767 +0 -0
- package/gdb/base/1/3997 +0 -0
- package/gdb/base/1/4143 +0 -0
- package/gdb/base/1/4144 +0 -0
- package/gdb/base/1/4145 +0 -0
- package/gdb/base/1/4146 +0 -0
- package/gdb/base/1/4147 +0 -0
- package/gdb/base/1/4148 +0 -0
- package/gdb/base/1/4149 +0 -0
- package/gdb/base/1/4150 +0 -0
- package/gdb/base/1/4151 +0 -0
- package/gdb/base/1/4152 +0 -0
- package/gdb/base/1/4153 +0 -0
- package/gdb/base/1/4154 +0 -0
- package/gdb/base/1/4155 +0 -0
- package/gdb/base/1/4156 +0 -0
- package/gdb/base/1/4157 +0 -0
- package/gdb/base/1/4158 +0 -0
- package/gdb/base/1/4159 +0 -0
- package/gdb/base/1/4160 +0 -0
- package/gdb/base/1/4163 +0 -0
- package/gdb/base/1/4164 +0 -0
- package/gdb/base/1/4165 +0 -0
- package/gdb/base/1/4166 +0 -0
- package/gdb/base/1/4167 +0 -0
- package/gdb/base/1/4168 +0 -0
- package/gdb/base/1/4169 +0 -0
- package/gdb/base/1/4170 +0 -0
- package/gdb/base/1/4171 +0 -0
- package/gdb/base/1/4172 +0 -0
- package/gdb/base/1/4173 +0 -0
- package/gdb/base/1/4174 +0 -0
- package/gdb/base/1/5002 +0 -0
- package/gdb/base/1/548 +0 -0
- package/gdb/base/1/549 +0 -0
- package/gdb/base/1/6102 +0 -0
- package/gdb/base/1/6104 +0 -0
- package/gdb/base/1/6106 +0 -0
- package/gdb/base/1/6110 +0 -0
- package/gdb/base/1/6111 +0 -0
- package/gdb/base/1/6112 +0 -0
- package/gdb/base/1/6113 +0 -0
- package/gdb/base/1/6117 +0 -0
- package/gdb/base/1/6175 +0 -0
- package/gdb/base/1/6176 +0 -0
- package/gdb/base/1/826 +0 -0
- package/gdb/base/1/827 +0 -0
- package/gdb/base/1/828 +0 -0
- package/gdb/base/1/PG_VERSION +0 -1
- package/gdb/base/1/pg_filenode.map +0 -0
- package/gdb/base/14033/112 +0 -0
- package/gdb/base/14033/113 +0 -0
- package/gdb/base/14033/1247 +0 -0
- package/gdb/base/14033/1247_fsm +0 -0
- package/gdb/base/14033/1247_vm +0 -0
- package/gdb/base/14033/1249 +0 -0
- package/gdb/base/14033/1249_fsm +0 -0
- package/gdb/base/14033/1249_vm +0 -0
- package/gdb/base/14033/1255 +0 -0
- package/gdb/base/14033/1255_fsm +0 -0
- package/gdb/base/14033/1255_vm +0 -0
- package/gdb/base/14033/1259 +0 -0
- package/gdb/base/14033/1259_fsm +0 -0
- package/gdb/base/14033/1259_vm +0 -0
- package/gdb/base/14033/13852 +0 -0
- package/gdb/base/14033/13852_fsm +0 -0
- package/gdb/base/14033/13852_vm +0 -0
- package/gdb/base/14033/13855 +0 -0
- package/gdb/base/14033/13856 +0 -0
- package/gdb/base/14033/13857 +0 -0
- package/gdb/base/14033/13857_fsm +0 -0
- package/gdb/base/14033/13857_vm +0 -0
- package/gdb/base/14033/13860 +0 -0
- package/gdb/base/14033/13861 +0 -0
- package/gdb/base/14033/13862 +0 -0
- package/gdb/base/14033/13862_fsm +0 -0
- package/gdb/base/14033/13862_vm +0 -0
- package/gdb/base/14033/13865 +0 -0
- package/gdb/base/14033/13866 +0 -0
- package/gdb/base/14033/13867 +0 -0
- package/gdb/base/14033/13867_fsm +0 -0
- package/gdb/base/14033/13867_vm +0 -0
- package/gdb/base/14033/13870 +0 -0
- package/gdb/base/14033/13871 +0 -0
- package/gdb/base/14033/1417 +0 -0
- package/gdb/base/14033/1418 +0 -0
- package/gdb/base/14033/174 +0 -0
- package/gdb/base/14033/175 +0 -0
- package/gdb/base/14033/2187 +0 -0
- package/gdb/base/14033/2224 +0 -0
- package/gdb/base/14033/2228 +0 -0
- package/gdb/base/14033/2328 +0 -0
- package/gdb/base/14033/2336 +0 -0
- package/gdb/base/14033/2337 +0 -0
- package/gdb/base/14033/2579 +0 -0
- package/gdb/base/14033/2600 +0 -0
- package/gdb/base/14033/2600_fsm +0 -0
- package/gdb/base/14033/2600_vm +0 -0
- package/gdb/base/14033/2601 +0 -0
- package/gdb/base/14033/2601_fsm +0 -0
- package/gdb/base/14033/2601_vm +0 -0
- package/gdb/base/14033/2602 +0 -0
- package/gdb/base/14033/2602_fsm +0 -0
- package/gdb/base/14033/2602_vm +0 -0
- package/gdb/base/14033/2603 +0 -0
- package/gdb/base/14033/2603_fsm +0 -0
- package/gdb/base/14033/2603_vm +0 -0
- package/gdb/base/14033/2604 +0 -0
- package/gdb/base/14033/2605 +0 -0
- package/gdb/base/14033/2605_fsm +0 -0
- package/gdb/base/14033/2605_vm +0 -0
- package/gdb/base/14033/2606 +0 -0
- package/gdb/base/14033/2606_fsm +0 -0
- package/gdb/base/14033/2606_vm +0 -0
- package/gdb/base/14033/2607 +0 -0
- package/gdb/base/14033/2607_fsm +0 -0
- package/gdb/base/14033/2607_vm +0 -0
- package/gdb/base/14033/2608 +0 -0
- package/gdb/base/14033/2608_fsm +0 -0
- package/gdb/base/14033/2608_vm +0 -0
- package/gdb/base/14033/2609 +0 -0
- package/gdb/base/14033/2609_fsm +0 -0
- package/gdb/base/14033/2609_vm +0 -0
- package/gdb/base/14033/2610 +0 -0
- package/gdb/base/14033/2610_fsm +0 -0
- package/gdb/base/14033/2610_vm +0 -0
- package/gdb/base/14033/2611 +0 -0
- package/gdb/base/14033/2612 +0 -0
- package/gdb/base/14033/2612_fsm +0 -0
- package/gdb/base/14033/2612_vm +0 -0
- package/gdb/base/14033/2613 +0 -0
- package/gdb/base/14033/2615 +0 -0
- package/gdb/base/14033/2615_fsm +0 -0
- package/gdb/base/14033/2615_vm +0 -0
- package/gdb/base/14033/2616 +0 -0
- package/gdb/base/14033/2616_fsm +0 -0
- package/gdb/base/14033/2616_vm +0 -0
- package/gdb/base/14033/2617 +0 -0
- package/gdb/base/14033/2617_fsm +0 -0
- package/gdb/base/14033/2617_vm +0 -0
- package/gdb/base/14033/2618 +0 -0
- package/gdb/base/14033/2618_fsm +0 -0
- package/gdb/base/14033/2618_vm +0 -0
- package/gdb/base/14033/2619 +0 -0
- package/gdb/base/14033/2619_fsm +0 -0
- package/gdb/base/14033/2619_vm +0 -0
- package/gdb/base/14033/2620 +0 -0
- package/gdb/base/14033/2650 +0 -0
- package/gdb/base/14033/2651 +0 -0
- package/gdb/base/14033/2652 +0 -0
- package/gdb/base/14033/2653 +0 -0
- package/gdb/base/14033/2654 +0 -0
- package/gdb/base/14033/2655 +0 -0
- package/gdb/base/14033/2656 +0 -0
- package/gdb/base/14033/2657 +0 -0
- package/gdb/base/14033/2658 +0 -0
- package/gdb/base/14033/2659 +0 -0
- package/gdb/base/14033/2660 +0 -0
- package/gdb/base/14033/2661 +0 -0
- package/gdb/base/14033/2662 +0 -0
- package/gdb/base/14033/2663 +0 -0
- package/gdb/base/14033/2664 +0 -0
- package/gdb/base/14033/2665 +0 -0
- package/gdb/base/14033/2666 +0 -0
- package/gdb/base/14033/2667 +0 -0
- package/gdb/base/14033/2668 +0 -0
- package/gdb/base/14033/2669 +0 -0
- package/gdb/base/14033/2670 +0 -0
- package/gdb/base/14033/2673 +0 -0
- package/gdb/base/14033/2673_fsm +0 -0
- package/gdb/base/14033/2674 +0 -0
- package/gdb/base/14033/2674_fsm +0 -0
- package/gdb/base/14033/2675 +0 -0
- package/gdb/base/14033/2678 +0 -0
- package/gdb/base/14033/2679 +0 -0
- package/gdb/base/14033/2680 +0 -0
- package/gdb/base/14033/2681 +0 -0
- package/gdb/base/14033/2682 +0 -0
- package/gdb/base/14033/2683 +0 -0
- package/gdb/base/14033/2684 +0 -0
- package/gdb/base/14033/2685 +0 -0
- package/gdb/base/14033/2686 +0 -0
- package/gdb/base/14033/2687 +0 -0
- package/gdb/base/14033/2688 +0 -0
- package/gdb/base/14033/2689 +0 -0
- package/gdb/base/14033/2690 +0 -0
- package/gdb/base/14033/2691 +0 -0
- package/gdb/base/14033/2692 +0 -0
- package/gdb/base/14033/2693 +0 -0
- package/gdb/base/14033/2696 +0 -0
- package/gdb/base/14033/2699 +0 -0
- package/gdb/base/14033/2701 +0 -0
- package/gdb/base/14033/2702 +0 -0
- package/gdb/base/14033/2703 +0 -0
- package/gdb/base/14033/2704 +0 -0
- package/gdb/base/14033/2753 +0 -0
- package/gdb/base/14033/2753_fsm +0 -0
- package/gdb/base/14033/2753_vm +0 -0
- package/gdb/base/14033/2754 +0 -0
- package/gdb/base/14033/2755 +0 -0
- package/gdb/base/14033/2756 +0 -0
- package/gdb/base/14033/2757 +0 -0
- package/gdb/base/14033/2830 +0 -0
- package/gdb/base/14033/2831 +0 -0
- package/gdb/base/14033/2832 +0 -0
- package/gdb/base/14033/2833 +0 -0
- package/gdb/base/14033/2834 +0 -0
- package/gdb/base/14033/2835 +0 -0
- package/gdb/base/14033/2836 +0 -0
- package/gdb/base/14033/2836_fsm +0 -0
- package/gdb/base/14033/2836_vm +0 -0
- package/gdb/base/14033/2837 +0 -0
- package/gdb/base/14033/2838 +0 -0
- package/gdb/base/14033/2838_fsm +0 -0
- package/gdb/base/14033/2838_vm +0 -0
- package/gdb/base/14033/2839 +0 -0
- package/gdb/base/14033/2840 +0 -0
- package/gdb/base/14033/2840_fsm +0 -0
- package/gdb/base/14033/2840_vm +0 -0
- package/gdb/base/14033/2841 +0 -0
- package/gdb/base/14033/2995 +0 -0
- package/gdb/base/14033/2996 +0 -0
- package/gdb/base/14033/3079 +0 -0
- package/gdb/base/14033/3079_fsm +0 -0
- package/gdb/base/14033/3079_vm +0 -0
- package/gdb/base/14033/3080 +0 -0
- package/gdb/base/14033/3081 +0 -0
- package/gdb/base/14033/3085 +0 -0
- package/gdb/base/14033/3118 +0 -0
- package/gdb/base/14033/3119 +0 -0
- package/gdb/base/14033/3164 +0 -0
- package/gdb/base/14033/3256 +0 -0
- package/gdb/base/14033/3257 +0 -0
- package/gdb/base/14033/3258 +0 -0
- package/gdb/base/14033/3350 +0 -0
- package/gdb/base/14033/3351 +0 -0
- package/gdb/base/14033/3379 +0 -0
- package/gdb/base/14033/3380 +0 -0
- package/gdb/base/14033/3381 +0 -0
- package/gdb/base/14033/3394 +0 -0
- package/gdb/base/14033/3394_fsm +0 -0
- package/gdb/base/14033/3394_vm +0 -0
- package/gdb/base/14033/3395 +0 -0
- package/gdb/base/14033/3429 +0 -0
- package/gdb/base/14033/3430 +0 -0
- package/gdb/base/14033/3431 +0 -0
- package/gdb/base/14033/3433 +0 -0
- package/gdb/base/14033/3439 +0 -0
- package/gdb/base/14033/3440 +0 -0
- package/gdb/base/14033/3455 +0 -0
- package/gdb/base/14033/3456 +0 -0
- package/gdb/base/14033/3456_fsm +0 -0
- package/gdb/base/14033/3456_vm +0 -0
- package/gdb/base/14033/3466 +0 -0
- package/gdb/base/14033/3467 +0 -0
- package/gdb/base/14033/3468 +0 -0
- package/gdb/base/14033/3501 +0 -0
- package/gdb/base/14033/3502 +0 -0
- package/gdb/base/14033/3503 +0 -0
- package/gdb/base/14033/3534 +0 -0
- package/gdb/base/14033/3541 +0 -0
- package/gdb/base/14033/3541_fsm +0 -0
- package/gdb/base/14033/3541_vm +0 -0
- package/gdb/base/14033/3542 +0 -0
- package/gdb/base/14033/3574 +0 -0
- package/gdb/base/14033/3575 +0 -0
- package/gdb/base/14033/3576 +0 -0
- package/gdb/base/14033/3596 +0 -0
- package/gdb/base/14033/3597 +0 -0
- package/gdb/base/14033/3598 +0 -0
- package/gdb/base/14033/3599 +0 -0
- package/gdb/base/14033/3600 +0 -0
- package/gdb/base/14033/3600_fsm +0 -0
- package/gdb/base/14033/3600_vm +0 -0
- package/gdb/base/14033/3601 +0 -0
- package/gdb/base/14033/3601_fsm +0 -0
- package/gdb/base/14033/3601_vm +0 -0
- package/gdb/base/14033/3602 +0 -0
- package/gdb/base/14033/3602_fsm +0 -0
- package/gdb/base/14033/3602_vm +0 -0
- package/gdb/base/14033/3603 +0 -0
- package/gdb/base/14033/3603_fsm +0 -0
- package/gdb/base/14033/3603_vm +0 -0
- package/gdb/base/14033/3604 +0 -0
- package/gdb/base/14033/3605 +0 -0
- package/gdb/base/14033/3606 +0 -0
- package/gdb/base/14033/3607 +0 -0
- package/gdb/base/14033/3608 +0 -0
- package/gdb/base/14033/3609 +0 -0
- package/gdb/base/14033/3712 +0 -0
- package/gdb/base/14033/3764 +0 -0
- package/gdb/base/14033/3764_fsm +0 -0
- package/gdb/base/14033/3764_vm +0 -0
- package/gdb/base/14033/3766 +0 -0
- package/gdb/base/14033/3767 +0 -0
- package/gdb/base/14033/3997 +0 -0
- package/gdb/base/14033/4143 +0 -0
- package/gdb/base/14033/4144 +0 -0
- package/gdb/base/14033/4145 +0 -0
- package/gdb/base/14033/4146 +0 -0
- package/gdb/base/14033/4147 +0 -0
- package/gdb/base/14033/4148 +0 -0
- package/gdb/base/14033/4149 +0 -0
- package/gdb/base/14033/4150 +0 -0
- package/gdb/base/14033/4151 +0 -0
- package/gdb/base/14033/4152 +0 -0
- package/gdb/base/14033/4153 +0 -0
- package/gdb/base/14033/4154 +0 -0
- package/gdb/base/14033/4155 +0 -0
- package/gdb/base/14033/4156 +0 -0
- package/gdb/base/14033/4157 +0 -0
- package/gdb/base/14033/4158 +0 -0
- package/gdb/base/14033/4159 +0 -0
- package/gdb/base/14033/4160 +0 -0
- package/gdb/base/14033/4163 +0 -0
- package/gdb/base/14033/4164 +0 -0
- package/gdb/base/14033/4165 +0 -0
- package/gdb/base/14033/4166 +0 -0
- package/gdb/base/14033/4167 +0 -0
- package/gdb/base/14033/4168 +0 -0
- package/gdb/base/14033/4169 +0 -0
- package/gdb/base/14033/4170 +0 -0
- package/gdb/base/14033/4171 +0 -0
- package/gdb/base/14033/4172 +0 -0
- package/gdb/base/14033/4173 +0 -0
- package/gdb/base/14033/4174 +0 -0
- package/gdb/base/14033/5002 +0 -0
- package/gdb/base/14033/548 +0 -0
- package/gdb/base/14033/549 +0 -0
- package/gdb/base/14033/6102 +0 -0
- package/gdb/base/14033/6104 +0 -0
- package/gdb/base/14033/6106 +0 -0
- package/gdb/base/14033/6110 +0 -0
- package/gdb/base/14033/6111 +0 -0
- package/gdb/base/14033/6112 +0 -0
- package/gdb/base/14033/6113 +0 -0
- package/gdb/base/14033/6117 +0 -0
- package/gdb/base/14033/6175 +0 -0
- package/gdb/base/14033/6176 +0 -0
- package/gdb/base/14033/826 +0 -0
- package/gdb/base/14033/827 +0 -0
- package/gdb/base/14033/828 +0 -0
- package/gdb/base/14033/PG_VERSION +0 -1
- package/gdb/base/14033/pg_filenode.map +0 -0
- package/gdb/base/14034/112 +0 -0
- package/gdb/base/14034/113 +0 -0
- package/gdb/base/14034/1247 +0 -0
- package/gdb/base/14034/1247_fsm +0 -0
- package/gdb/base/14034/1247_vm +0 -0
- package/gdb/base/14034/1249 +0 -0
- package/gdb/base/14034/1249_fsm +0 -0
- package/gdb/base/14034/1249_vm +0 -0
- package/gdb/base/14034/1255 +0 -0
- package/gdb/base/14034/1255_fsm +0 -0
- package/gdb/base/14034/1255_vm +0 -0
- package/gdb/base/14034/1259 +0 -0
- package/gdb/base/14034/1259_fsm +0 -0
- package/gdb/base/14034/1259_vm +0 -0
- package/gdb/base/14034/13852 +0 -0
- package/gdb/base/14034/13852_fsm +0 -0
- package/gdb/base/14034/13852_vm +0 -0
- package/gdb/base/14034/13855 +0 -0
- package/gdb/base/14034/13856 +0 -0
- package/gdb/base/14034/13857 +0 -0
- package/gdb/base/14034/13857_fsm +0 -0
- package/gdb/base/14034/13857_vm +0 -0
- package/gdb/base/14034/13860 +0 -0
- package/gdb/base/14034/13861 +0 -0
- package/gdb/base/14034/13862 +0 -0
- package/gdb/base/14034/13862_fsm +0 -0
- package/gdb/base/14034/13862_vm +0 -0
- package/gdb/base/14034/13865 +0 -0
- package/gdb/base/14034/13866 +0 -0
- package/gdb/base/14034/13867 +0 -0
- package/gdb/base/14034/13867_fsm +0 -0
- package/gdb/base/14034/13867_vm +0 -0
- package/gdb/base/14034/13870 +0 -0
- package/gdb/base/14034/13871 +0 -0
- package/gdb/base/14034/1417 +0 -0
- package/gdb/base/14034/1418 +0 -0
- package/gdb/base/14034/16386 +0 -0
- package/gdb/base/14034/16389 +0 -0
- package/gdb/base/14034/16390 +0 -0
- package/gdb/base/14034/16391 +0 -0
- package/gdb/base/14034/16393 +0 -0
- package/gdb/base/14034/16396 +0 -0
- package/gdb/base/14034/16397 +0 -0
- package/gdb/base/14034/16398 +0 -0
- package/gdb/base/14034/16400 +0 -0
- package/gdb/base/14034/16403 +0 -0
- package/gdb/base/14034/16404 +0 -0
- package/gdb/base/14034/16420 +0 -0
- package/gdb/base/14034/16423 +0 -0
- package/gdb/base/14034/16424 +0 -0
- package/gdb/base/14034/16425 +0 -0
- package/gdb/base/14034/16427 +0 -0
- package/gdb/base/14034/16430 +0 -0
- package/gdb/base/14034/16431 +0 -0
- package/gdb/base/14034/16432 +0 -0
- package/gdb/base/14034/16434 +0 -0
- package/gdb/base/14034/16437 +0 -0
- package/gdb/base/14034/16438 +0 -0
- package/gdb/base/14034/16439 +0 -0
- package/gdb/base/14034/16446 +0 -0
- package/gdb/base/14034/16449 +0 -0
- package/gdb/base/14034/16450 +0 -0
- package/gdb/base/14034/16451 +0 -0
- package/gdb/base/14034/16458 +0 -0
- package/gdb/base/14034/16461 +0 -0
- package/gdb/base/14034/16462 +0 -0
- package/gdb/base/14034/16463 +0 -0
- package/gdb/base/14034/16475 +0 -0
- package/gdb/base/14034/16478 +0 -0
- package/gdb/base/14034/16479 +0 -0
- package/gdb/base/14034/16480 +0 -0
- package/gdb/base/14034/174 +0 -0
- package/gdb/base/14034/175 +0 -0
- package/gdb/base/14034/2187 +0 -0
- package/gdb/base/14034/2224 +0 -0
- package/gdb/base/14034/2228 +0 -0
- package/gdb/base/14034/2328 +0 -0
- package/gdb/base/14034/2336 +0 -0
- package/gdb/base/14034/2337 +0 -0
- package/gdb/base/14034/2579 +0 -0
- package/gdb/base/14034/2600 +0 -0
- package/gdb/base/14034/2600_fsm +0 -0
- package/gdb/base/14034/2600_vm +0 -0
- package/gdb/base/14034/2601 +0 -0
- package/gdb/base/14034/2601_fsm +0 -0
- package/gdb/base/14034/2601_vm +0 -0
- package/gdb/base/14034/2602 +0 -0
- package/gdb/base/14034/2602_fsm +0 -0
- package/gdb/base/14034/2602_vm +0 -0
- package/gdb/base/14034/2603 +0 -0
- package/gdb/base/14034/2603_fsm +0 -0
- package/gdb/base/14034/2603_vm +0 -0
- package/gdb/base/14034/2604 +0 -0
- package/gdb/base/14034/2605 +0 -0
- package/gdb/base/14034/2605_fsm +0 -0
- package/gdb/base/14034/2605_vm +0 -0
- package/gdb/base/14034/2606 +0 -0
- package/gdb/base/14034/2606_fsm +0 -0
- package/gdb/base/14034/2606_vm +0 -0
- package/gdb/base/14034/2607 +0 -0
- package/gdb/base/14034/2607_fsm +0 -0
- package/gdb/base/14034/2607_vm +0 -0
- package/gdb/base/14034/2608 +0 -0
- package/gdb/base/14034/2608_fsm +0 -0
- package/gdb/base/14034/2608_vm +0 -0
- package/gdb/base/14034/2609 +0 -0
- package/gdb/base/14034/2609_fsm +0 -0
- package/gdb/base/14034/2609_vm +0 -0
- package/gdb/base/14034/2610 +0 -0
- package/gdb/base/14034/2610_fsm +0 -0
- package/gdb/base/14034/2610_vm +0 -0
- package/gdb/base/14034/2611 +0 -0
- package/gdb/base/14034/2612 +0 -0
- package/gdb/base/14034/2612_fsm +0 -0
- package/gdb/base/14034/2612_vm +0 -0
- package/gdb/base/14034/2613 +0 -0
- package/gdb/base/14034/2615 +0 -0
- package/gdb/base/14034/2615_fsm +0 -0
- package/gdb/base/14034/2615_vm +0 -0
- package/gdb/base/14034/2616 +0 -0
- package/gdb/base/14034/2616_fsm +0 -0
- package/gdb/base/14034/2616_vm +0 -0
- package/gdb/base/14034/2617 +0 -0
- package/gdb/base/14034/2617_fsm +0 -0
- package/gdb/base/14034/2617_vm +0 -0
- package/gdb/base/14034/2618 +0 -0
- package/gdb/base/14034/2618_fsm +0 -0
- package/gdb/base/14034/2618_vm +0 -0
- package/gdb/base/14034/2619 +0 -0
- package/gdb/base/14034/2619_fsm +0 -0
- package/gdb/base/14034/2619_vm +0 -0
- package/gdb/base/14034/2620 +0 -0
- package/gdb/base/14034/2650 +0 -0
- package/gdb/base/14034/2651 +0 -0
- package/gdb/base/14034/2652 +0 -0
- package/gdb/base/14034/2653 +0 -0
- package/gdb/base/14034/2654 +0 -0
- package/gdb/base/14034/2655 +0 -0
- package/gdb/base/14034/2656 +0 -0
- package/gdb/base/14034/2657 +0 -0
- package/gdb/base/14034/2658 +0 -0
- package/gdb/base/14034/2659 +0 -0
- package/gdb/base/14034/2660 +0 -0
- package/gdb/base/14034/2661 +0 -0
- package/gdb/base/14034/2662 +0 -0
- package/gdb/base/14034/2663 +0 -0
- package/gdb/base/14034/2664 +0 -0
- package/gdb/base/14034/2665 +0 -0
- package/gdb/base/14034/2666 +0 -0
- package/gdb/base/14034/2667 +0 -0
- package/gdb/base/14034/2668 +0 -0
- package/gdb/base/14034/2669 +0 -0
- package/gdb/base/14034/2670 +0 -0
- package/gdb/base/14034/2673 +0 -0
- package/gdb/base/14034/2673_fsm +0 -0
- package/gdb/base/14034/2674 +0 -0
- package/gdb/base/14034/2674_fsm +0 -0
- package/gdb/base/14034/2675 +0 -0
- package/gdb/base/14034/2678 +0 -0
- package/gdb/base/14034/2679 +0 -0
- package/gdb/base/14034/2680 +0 -0
- package/gdb/base/14034/2681 +0 -0
- package/gdb/base/14034/2682 +0 -0
- package/gdb/base/14034/2683 +0 -0
- package/gdb/base/14034/2684 +0 -0
- package/gdb/base/14034/2685 +0 -0
- package/gdb/base/14034/2686 +0 -0
- package/gdb/base/14034/2687 +0 -0
- package/gdb/base/14034/2688 +0 -0
- package/gdb/base/14034/2689 +0 -0
- package/gdb/base/14034/2690 +0 -0
- package/gdb/base/14034/2691 +0 -0
- package/gdb/base/14034/2692 +0 -0
- package/gdb/base/14034/2693 +0 -0
- package/gdb/base/14034/2696 +0 -0
- package/gdb/base/14034/2699 +0 -0
- package/gdb/base/14034/2701 +0 -0
- package/gdb/base/14034/2702 +0 -0
- package/gdb/base/14034/2703 +0 -0
- package/gdb/base/14034/2704 +0 -0
- package/gdb/base/14034/2753 +0 -0
- package/gdb/base/14034/2753_fsm +0 -0
- package/gdb/base/14034/2753_vm +0 -0
- package/gdb/base/14034/2754 +0 -0
- package/gdb/base/14034/2755 +0 -0
- package/gdb/base/14034/2756 +0 -0
- package/gdb/base/14034/2757 +0 -0
- package/gdb/base/14034/2830 +0 -0
- package/gdb/base/14034/2831 +0 -0
- package/gdb/base/14034/2832 +0 -0
- package/gdb/base/14034/2833 +0 -0
- package/gdb/base/14034/2834 +0 -0
- package/gdb/base/14034/2835 +0 -0
- package/gdb/base/14034/2836 +0 -0
- package/gdb/base/14034/2836_fsm +0 -0
- package/gdb/base/14034/2836_vm +0 -0
- package/gdb/base/14034/2837 +0 -0
- package/gdb/base/14034/2838 +0 -0
- package/gdb/base/14034/2838_fsm +0 -0
- package/gdb/base/14034/2838_vm +0 -0
- package/gdb/base/14034/2839 +0 -0
- package/gdb/base/14034/2840 +0 -0
- package/gdb/base/14034/2840_fsm +0 -0
- package/gdb/base/14034/2840_vm +0 -0
- package/gdb/base/14034/2841 +0 -0
- package/gdb/base/14034/2995 +0 -0
- package/gdb/base/14034/2996 +0 -0
- package/gdb/base/14034/3079 +0 -0
- package/gdb/base/14034/3079_fsm +0 -0
- package/gdb/base/14034/3079_vm +0 -0
- package/gdb/base/14034/3080 +0 -0
- package/gdb/base/14034/3081 +0 -0
- package/gdb/base/14034/3085 +0 -0
- package/gdb/base/14034/3118 +0 -0
- package/gdb/base/14034/3119 +0 -0
- package/gdb/base/14034/3164 +0 -0
- package/gdb/base/14034/3256 +0 -0
- package/gdb/base/14034/3257 +0 -0
- package/gdb/base/14034/3258 +0 -0
- package/gdb/base/14034/3350 +0 -0
- package/gdb/base/14034/3351 +0 -0
- package/gdb/base/14034/3379 +0 -0
- package/gdb/base/14034/3380 +0 -0
- package/gdb/base/14034/3381 +0 -0
- package/gdb/base/14034/3394 +0 -0
- package/gdb/base/14034/3394_fsm +0 -0
- package/gdb/base/14034/3394_vm +0 -0
- package/gdb/base/14034/3395 +0 -0
- package/gdb/base/14034/3429 +0 -0
- package/gdb/base/14034/3430 +0 -0
- package/gdb/base/14034/3431 +0 -0
- package/gdb/base/14034/3433 +0 -0
- package/gdb/base/14034/3439 +0 -0
- package/gdb/base/14034/3440 +0 -0
- package/gdb/base/14034/3455 +0 -0
- package/gdb/base/14034/3456 +0 -0
- package/gdb/base/14034/3456_fsm +0 -0
- package/gdb/base/14034/3456_vm +0 -0
- package/gdb/base/14034/3466 +0 -0
- package/gdb/base/14034/3467 +0 -0
- package/gdb/base/14034/3468 +0 -0
- package/gdb/base/14034/3501 +0 -0
- package/gdb/base/14034/3502 +0 -0
- package/gdb/base/14034/3503 +0 -0
- package/gdb/base/14034/3534 +0 -0
- package/gdb/base/14034/3541 +0 -0
- package/gdb/base/14034/3541_fsm +0 -0
- package/gdb/base/14034/3541_vm +0 -0
- package/gdb/base/14034/3542 +0 -0
- package/gdb/base/14034/3574 +0 -0
- package/gdb/base/14034/3575 +0 -0
- package/gdb/base/14034/3576 +0 -0
- package/gdb/base/14034/3596 +0 -0
- package/gdb/base/14034/3597 +0 -0
- package/gdb/base/14034/3598 +0 -0
- package/gdb/base/14034/3599 +0 -0
- package/gdb/base/14034/3600 +0 -0
- package/gdb/base/14034/3600_fsm +0 -0
- package/gdb/base/14034/3600_vm +0 -0
- package/gdb/base/14034/3601 +0 -0
- package/gdb/base/14034/3601_fsm +0 -0
- package/gdb/base/14034/3601_vm +0 -0
- package/gdb/base/14034/3602 +0 -0
- package/gdb/base/14034/3602_fsm +0 -0
- package/gdb/base/14034/3602_vm +0 -0
- package/gdb/base/14034/3603 +0 -0
- package/gdb/base/14034/3603_fsm +0 -0
- package/gdb/base/14034/3603_vm +0 -0
- package/gdb/base/14034/3604 +0 -0
- package/gdb/base/14034/3605 +0 -0
- package/gdb/base/14034/3606 +0 -0
- package/gdb/base/14034/3607 +0 -0
- package/gdb/base/14034/3608 +0 -0
- package/gdb/base/14034/3609 +0 -0
- package/gdb/base/14034/3712 +0 -0
- package/gdb/base/14034/3764 +0 -0
- package/gdb/base/14034/3764_fsm +0 -0
- package/gdb/base/14034/3764_vm +0 -0
- package/gdb/base/14034/3766 +0 -0
- package/gdb/base/14034/3767 +0 -0
- package/gdb/base/14034/3997 +0 -0
- package/gdb/base/14034/4143 +0 -0
- package/gdb/base/14034/4144 +0 -0
- package/gdb/base/14034/4145 +0 -0
- package/gdb/base/14034/4146 +0 -0
- package/gdb/base/14034/4147 +0 -0
- package/gdb/base/14034/4148 +0 -0
- package/gdb/base/14034/4149 +0 -0
- package/gdb/base/14034/4150 +0 -0
- package/gdb/base/14034/4151 +0 -0
- package/gdb/base/14034/4152 +0 -0
- package/gdb/base/14034/4153 +0 -0
- package/gdb/base/14034/4154 +0 -0
- package/gdb/base/14034/4155 +0 -0
- package/gdb/base/14034/4156 +0 -0
- package/gdb/base/14034/4157 +0 -0
- package/gdb/base/14034/4158 +0 -0
- package/gdb/base/14034/4159 +0 -0
- package/gdb/base/14034/4160 +0 -0
- package/gdb/base/14034/4163 +0 -0
- package/gdb/base/14034/4164 +0 -0
- package/gdb/base/14034/4165 +0 -0
- package/gdb/base/14034/4166 +0 -0
- package/gdb/base/14034/4167 +0 -0
- package/gdb/base/14034/4168 +0 -0
- package/gdb/base/14034/4169 +0 -0
- package/gdb/base/14034/4170 +0 -0
- package/gdb/base/14034/4171 +0 -0
- package/gdb/base/14034/4172 +0 -0
- package/gdb/base/14034/4173 +0 -0
- package/gdb/base/14034/4174 +0 -0
- package/gdb/base/14034/5002 +0 -0
- package/gdb/base/14034/548 +0 -0
- package/gdb/base/14034/549 +0 -0
- package/gdb/base/14034/6102 +0 -0
- package/gdb/base/14034/6104 +0 -0
- package/gdb/base/14034/6106 +0 -0
- package/gdb/base/14034/6110 +0 -0
- package/gdb/base/14034/6111 +0 -0
- package/gdb/base/14034/6112 +0 -0
- package/gdb/base/14034/6113 +0 -0
- package/gdb/base/14034/6117 +0 -0
- package/gdb/base/14034/6175 +0 -0
- package/gdb/base/14034/6176 +0 -0
- package/gdb/base/14034/826 +0 -0
- package/gdb/base/14034/827 +0 -0
- package/gdb/base/14034/828 +0 -0
- package/gdb/base/14034/PG_VERSION +0 -1
- package/gdb/base/14034/pg_filenode.map +0 -0
- package/gdb/base/14034/pg_internal.init +0 -0
- package/gdb/global/1213 +0 -0
- package/gdb/global/1213_fsm +0 -0
- package/gdb/global/1213_vm +0 -0
- package/gdb/global/1214 +0 -0
- package/gdb/global/1214_fsm +0 -0
- package/gdb/global/1214_vm +0 -0
- package/gdb/global/1232 +0 -0
- package/gdb/global/1233 +0 -0
- package/gdb/global/1260 +0 -0
- package/gdb/global/1260_fsm +0 -0
- package/gdb/global/1260_vm +0 -0
- package/gdb/global/1261 +0 -0
- package/gdb/global/1261_fsm +0 -0
- package/gdb/global/1261_vm +0 -0
- package/gdb/global/1262 +0 -0
- package/gdb/global/1262_fsm +0 -0
- package/gdb/global/1262_vm +0 -0
- package/gdb/global/2396 +0 -0
- package/gdb/global/2396_fsm +0 -0
- package/gdb/global/2396_vm +0 -0
- package/gdb/global/2397 +0 -0
- package/gdb/global/2671 +0 -0
- package/gdb/global/2672 +0 -0
- package/gdb/global/2676 +0 -0
- package/gdb/global/2677 +0 -0
- package/gdb/global/2694 +0 -0
- package/gdb/global/2695 +0 -0
- package/gdb/global/2697 +0 -0
- package/gdb/global/2698 +0 -0
- package/gdb/global/2846 +0 -0
- package/gdb/global/2847 +0 -0
- package/gdb/global/2964 +0 -0
- package/gdb/global/2965 +0 -0
- package/gdb/global/2966 +0 -0
- package/gdb/global/2967 +0 -0
- package/gdb/global/3592 +0 -0
- package/gdb/global/3593 +0 -0
- package/gdb/global/4060 +0 -0
- package/gdb/global/4061 +0 -0
- package/gdb/global/4175 +0 -0
- package/gdb/global/4176 +0 -0
- package/gdb/global/4177 +0 -0
- package/gdb/global/4178 +0 -0
- package/gdb/global/4181 +0 -0
- package/gdb/global/4182 +0 -0
- package/gdb/global/4183 +0 -0
- package/gdb/global/4184 +0 -0
- package/gdb/global/4185 +0 -0
- package/gdb/global/4186 +0 -0
- package/gdb/global/6000 +0 -0
- package/gdb/global/6001 +0 -0
- package/gdb/global/6002 +0 -0
- package/gdb/global/6100 +0 -0
- package/gdb/global/6114 +0 -0
- package/gdb/global/6115 +0 -0
- package/gdb/global/pg_control +0 -0
- package/gdb/global/pg_filenode.map +0 -0
- package/gdb/global/pg_internal.init +0 -0
- package/gdb/pg_hba.conf +0 -98
- package/gdb/pg_ident.conf +0 -42
- package/gdb/pg_logical/replorigin_checkpoint +0 -0
- package/gdb/pg_multixact/members/0000 +0 -0
- package/gdb/pg_multixact/offsets/0000 +0 -0
- package/gdb/pg_stat/db_0.stat +0 -0
- package/gdb/pg_stat/db_14034.stat +0 -0
- package/gdb/pg_stat/global.stat +0 -0
- package/gdb/pg_subtrans/0000 +0 -0
- package/gdb/pg_wal/000000010000000000000001 +0 -0
- package/gdb/pg_xact/0000 +0 -0
- package/gdb/postgresql.auto.conf +0 -2
- package/gdb/postgresql.conf +0 -796
- package/gdb/postmaster.opts +0 -1
- package/server/API Project-7d45e9f24a96.p12 +0 -0
- package/server/grnmap.pem +0 -31
- package/server/grnsight.pem +0 -19
- package/test-files/import-samples/attributes.graphml +0 -40
- package/test-files/import-samples/port.graphml +0 -32
- package/test-files/import-samples/simple.graphml +0 -31
- package/web-client/public/js/grnsight.min.js +0 -2371
package/database/README.md
CHANGED
|
@@ -2,10 +2,18 @@
|
|
|
2
2
|
Here are the files pertaining to both the network and expression databases. Look within the README.md files of both folders for information pertinent to the schema that you intend to be using.
|
|
3
3
|
## Setting up a local postgres GRNsight Database
|
|
4
4
|
1. Installing PostgreSQL on your computer
|
|
5
|
-
- MacOS and Windows can follow these
|
|
6
|
-
-
|
|
7
|
-
-
|
|
8
|
-
|
|
5
|
+
- MacOS and Windows can follow these instructions on how to install postgreSQL.
|
|
6
|
+
- Install the software at this [link](https://www.postgresql.org/download/)
|
|
7
|
+
- Initialize the database
|
|
8
|
+
- If your terminal emits a message that looks like `initdb --locale=C -E UTF-8 location-of-cluster` from Step 1B, then your installer has initialized a database for you.
|
|
9
|
+
- Open the terminal and type the command `initdb --locale=C -E UTF-8 location-of-cluster`
|
|
10
|
+
- "Cluster" is the PostgreSQL term for the file structure of a PostgreSQL database instance
|
|
11
|
+
- You will have to modify location-of-cluster to the folder name you want to store the database (you don't need to create a folder, the command will create the folder for you, just create the name)
|
|
12
|
+
- Start and stop the server
|
|
13
|
+
- Additionally, your installer may start the server for you upon installation (You can save this command for further reuse).
|
|
14
|
+
- To start the server yourself run `pg_ctl start -D location-of-cluster` (You can save this command for further reuse).
|
|
15
|
+
- To stop the server run `pg_ctl stop -D location-of-cluster`.
|
|
16
|
+
|
|
9
17
|
- Linux users
|
|
10
18
|
- The MacOS and Windows instructions will _probably_ not work for you. You can try at your own risk to check.
|
|
11
19
|
- Linux users can try these [instructions](https://www.geeksforgeeks.org/install-postgresql-on-linux/) and that should work for you (...maybe...). If it doesn't try googling instructions with your specific operating system. Sorry!
|
|
@@ -20,11 +28,11 @@ Here are the files pertaining to both the network and expression databases. Look
|
|
|
20
28
|
From there, create the schemas using the following commands:
|
|
21
29
|
|
|
22
30
|
```
|
|
23
|
-
CREATE SCHEMA
|
|
31
|
+
CREATE SCHEMA gene_regulatory_network;
|
|
24
32
|
```
|
|
25
33
|
|
|
26
34
|
```
|
|
27
|
-
CREATE SCHEMA
|
|
35
|
+
CREATE SCHEMA gene_expression;
|
|
28
36
|
```
|
|
29
37
|
|
|
30
38
|
Once they are created you can exit your database using the command `\q`.
|
|
@@ -47,18 +55,32 @@ Here are the files pertaining to both the network and expression databases. Look
|
|
|
47
55
|
pip3 install pandas requests intermine tzlocal
|
|
48
56
|
```
|
|
49
57
|
|
|
50
|
-
Once the dependencies have been installed, you can run
|
|
51
|
-
|
|
58
|
+
Once the dependencies have been installed, you can run
|
|
52
59
|
```
|
|
53
|
-
|
|
60
|
+
cd <path to GRNsight/database/network-database/scripts>
|
|
61
|
+
python3 generate_network.py
|
|
54
62
|
```
|
|
55
|
-
|
|
56
63
|
This will take a while to get all of the network data and generate all of the files. This will create a folder full of the processed files in `database/network-database/script-results`.
|
|
64
|
+
|
|
65
|
+
*** Note: *** If you get an error similar to the following image where it references the in then you are one of the unlucky few who has to edit the intermine.py file directly.
|
|
66
|
+
|
|
67
|
+
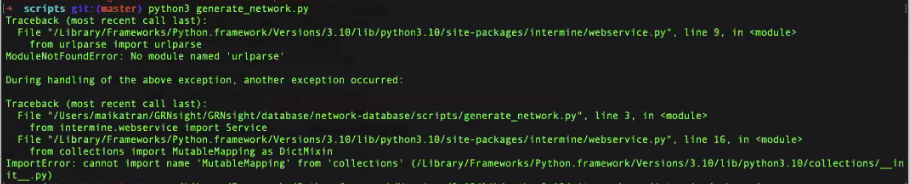
|
|
68
|
+
|
|
69
|
+
- Navigate the referenced file ( \<path specific to your machine>/intermine/webservice.py )
|
|
70
|
+
|
|
71
|
+
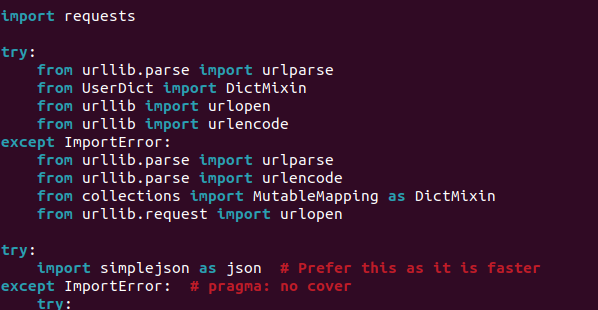
- The try-catch block should look like this:
|
|
72
|
+
|
|
73
|
+
- 
|
|
74
|
+
|
|
75
|
+
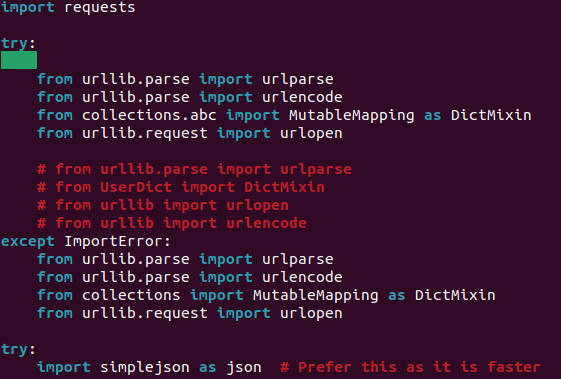
- Change it to the following, rerun the `generate_network.py` command and it should work! If it doesn't you may need to troubleshoot a bit further (´◕ ᵔ ◕`✿)*ᶜʳᶦᵉˢ*.
|
|
76
|
+
|
|
77
|
+
- 
|
|
57
78
|
|
|
58
|
-
2. Load the processed files into your database.
|
|
79
|
+
2. Load the processed files into your database.
|
|
59
80
|
|
|
60
81
|
```
|
|
61
|
-
|
|
82
|
+
cd <path to GRNsight/database/network-database/scripts>
|
|
83
|
+
python3 loader.py | psql postgresql://localhost/postgres
|
|
62
84
|
```
|
|
63
85
|
|
|
64
86
|
This should output a bunch of COPY print statements to your terminal. Once complete your database is now loaded with the network data.
|
|
@@ -70,18 +92,20 @@ Here are the files pertaining to both the network and expression databases. Look
|
|
|
70
92
|
mkdir <path to GRNsight/database/expression-database>/source-files
|
|
71
93
|
```
|
|
72
94
|
|
|
73
|
-
2. Download the _"Expression 2020"_ folder from Box located in `GRNsight > GRNsight Expression > Expression 2020` to your newly created `source-files` folder
|
|
95
|
+
2. Download the _"Expression 2020"_ folder from Box located in `GRNsight > GRNsight Expression > Expression 2020` to your newly created `source-files` folder. Your the path should look like this: GRNsight > database > expression-database > source-files > Expression 2020 > [the actual csv and xlsx files are here!]
|
|
74
96
|
3. Run the pre-processing script on the data. This will create a folder full of the processed files in `database/expression-database/script-results`.
|
|
75
97
|
|
|
76
98
|
```
|
|
77
|
-
|
|
99
|
+
cd <path to GRNsight/database/expression-database/scripts>
|
|
100
|
+
python3 preprocessing.py
|
|
78
101
|
```
|
|
79
102
|
|
|
80
103
|
4. Load the processed files into your database.
|
|
81
104
|
|
|
82
105
|
```
|
|
83
|
-
|
|
106
|
+
cd <path to GRNsight/database/expression-database/scripts>
|
|
107
|
+
python3 loader.py | psql postgresql://localhost/postgres
|
|
84
108
|
```
|
|
85
109
|
|
|
86
110
|
This should output a bunch of COPY print statements to your terminal. Once complete your database is now loaded with the expression data.
|
|
87
|
-
|
|
111
|
+
3. Continue setting up in the [Initial Setup Wiki page](https://github.com/dondi/GRNsight/wiki/Initial-Setup)
|
|
@@ -6,9 +6,9 @@ All files pertaining the expression database live within this directory.
|
|
|
6
6
|
|
|
7
7
|
#### Schema
|
|
8
8
|
|
|
9
|
-
All network data is stored within the
|
|
9
|
+
All network data is stored within the gene_expression schema on our Postgres database.
|
|
10
10
|
|
|
11
|
-
The schema is located within this directory at the top level in the file `schema.sql`. It defines the tables located within the
|
|
11
|
+
The schema is located within this directory at the top level in the file `schema.sql`. It defines the tables located within the gene_expression schema.
|
|
12
12
|
|
|
13
13
|
Usage:
|
|
14
14
|
To load to local database
|
|
@@ -1,4 +1,4 @@
|
|
|
1
|
-
CREATE TABLE
|
|
1
|
+
CREATE TABLE gene_expression.ref (
|
|
2
2
|
pubmed_id VARCHAR,
|
|
3
3
|
authors VARCHAR,
|
|
4
4
|
publication_year VARCHAR,
|
|
@@ -8,7 +8,7 @@ CREATE TABLE fall2021.ref (
|
|
|
8
8
|
PRIMARY KEY(ncbi_geo_id, pubmed_id)
|
|
9
9
|
);
|
|
10
10
|
|
|
11
|
-
CREATE TABLE
|
|
11
|
+
CREATE TABLE gene_expression.gene (
|
|
12
12
|
gene_id VARCHAR, -- systematic like name
|
|
13
13
|
display_gene_id VARCHAR, -- standard like name
|
|
14
14
|
species VARCHAR,
|
|
@@ -16,10 +16,10 @@ CREATE TABLE fall2021.gene (
|
|
|
16
16
|
PRIMARY KEY(gene_id, taxon_id)
|
|
17
17
|
);
|
|
18
18
|
|
|
19
|
-
CREATE TABLE
|
|
19
|
+
CREATE TABLE gene_expression.expression_metadata (
|
|
20
20
|
ncbi_geo_id VARCHAR,
|
|
21
21
|
pubmed_id VARCHAR,
|
|
22
|
-
FOREIGN KEY (ncbi_geo_id, pubmed_id) REFERENCES
|
|
22
|
+
FOREIGN KEY (ncbi_geo_id, pubmed_id) REFERENCES gene_expression.ref(ncbi_geo_id, pubmed_id),
|
|
23
23
|
control_yeast_strain VARCHAR,
|
|
24
24
|
treatment_yeast_strain VARCHAR,
|
|
25
25
|
control VARCHAR,
|
|
@@ -33,10 +33,10 @@ CREATE TABLE fall2021.expression_metadata (
|
|
|
33
33
|
display_expression_table VARCHAR,
|
|
34
34
|
PRIMARY KEY(ncbi_geo_id, pubmed_id, time_value)
|
|
35
35
|
);
|
|
36
|
-
CREATE TABLE
|
|
36
|
+
CREATE TABLE gene_expression.expression (
|
|
37
37
|
gene_id VARCHAR,
|
|
38
38
|
taxon_id VARCHAR,
|
|
39
|
-
FOREIGN KEY (gene_id, taxon_id) REFERENCES
|
|
39
|
+
FOREIGN KEY (gene_id, taxon_id) REFERENCES gene_expression.gene(gene_id, taxon_id),
|
|
40
40
|
-- ncbi_geo_id VARCHAR,
|
|
41
41
|
-- pubmed_id VARCHAR,
|
|
42
42
|
sort_index INT,
|
|
@@ -45,27 +45,27 @@ CREATE TABLE fall2021.expression (
|
|
|
45
45
|
time_point FLOAT,
|
|
46
46
|
dataset VARCHAR,
|
|
47
47
|
PRIMARY KEY(gene_id, sample_id)
|
|
48
|
-
-- FOREIGN KEY (ncbi_geo_id, pubmed_id, time_point) REFERENCES
|
|
48
|
+
-- FOREIGN KEY (ncbi_geo_id, pubmed_id, time_point) REFERENCES gene_expression.expression_metadata(ncbi_geo_id, pubmed_id, time_value)
|
|
49
49
|
);
|
|
50
|
-
CREATE TABLE
|
|
50
|
+
CREATE TABLE gene_expression.degradation_rate (
|
|
51
51
|
gene_id VARCHAR,
|
|
52
52
|
taxon_id VARCHAR,
|
|
53
|
-
FOREIGN KEY (gene_id, taxon_id) REFERENCES
|
|
53
|
+
FOREIGN KEY (gene_id, taxon_id) REFERENCES gene_expression.gene(gene_id, taxon_id),
|
|
54
54
|
ncbi_geo_id VARCHAR,
|
|
55
55
|
pubmed_id VARCHAR,
|
|
56
|
-
FOREIGN KEY (ncbi_geo_id, pubmed_id) REFERENCES
|
|
56
|
+
FOREIGN KEY (ncbi_geo_id, pubmed_id) REFERENCES gene_expression.ref(ncbi_geo_id, pubmed_id),
|
|
57
57
|
PRIMARY KEY(gene_id, ncbi_geo_id, pubmed_id),
|
|
58
58
|
degradation_rate FLOAT
|
|
59
59
|
);
|
|
60
60
|
|
|
61
|
-
CREATE TABLE
|
|
61
|
+
CREATE TABLE gene_expression.production_rate (
|
|
62
62
|
gene_id VARCHAR,
|
|
63
63
|
taxon_id VARCHAR,
|
|
64
|
-
FOREIGN KEY (gene_id, taxon_id) REFERENCES
|
|
64
|
+
FOREIGN KEY (gene_id, taxon_id) REFERENCES gene_expression.gene(gene_id, taxon_id),
|
|
65
65
|
ncbi_geo_id VARCHAR,
|
|
66
66
|
pubmed_id VARCHAR,
|
|
67
|
-
FOREIGN KEY (ncbi_geo_id, pubmed_id) REFERENCES
|
|
67
|
+
FOREIGN KEY (ncbi_geo_id, pubmed_id) REFERENCES gene_expression.ref(ncbi_geo_id, pubmed_id),
|
|
68
68
|
PRIMARY KEY(gene_id, ncbi_geo_id, pubmed_id),
|
|
69
69
|
production_rate FLOAT
|
|
70
|
-
-- FOREIGN KEY (gene_id, ncbi_geo_id, pubmed_id) REFERENCES
|
|
70
|
+
-- FOREIGN KEY (gene_id, ncbi_geo_id, pubmed_id) REFERENCES gene_expression.degradation_rate(gene_id, ncbi_geo_id, pubmed_id) -- not sure if we want to link the generated production rate to it's original degradation rate
|
|
71
71
|
);
|
|
@@ -45,7 +45,7 @@ def convert_int(potential_int):
|
|
|
45
45
|
This program Loads Refs into the database
|
|
46
46
|
"""
|
|
47
47
|
def LOAD_REFS():
|
|
48
|
-
print('COPY
|
|
48
|
+
print('COPY gene_expression.ref (pubmed_id, authors, publication_year, title, doi, ncbi_geo_id) FROM stdin;')
|
|
49
49
|
REFS_SOURCE = '../script-results/processed-expression/refs.csv'
|
|
50
50
|
with open(REFS_SOURCE, 'r+') as f:
|
|
51
51
|
reader = csv.reader(f)
|
|
@@ -67,7 +67,7 @@ def LOAD_REFS():
|
|
|
67
67
|
This program Loads ID Mapping into the database
|
|
68
68
|
"""
|
|
69
69
|
def LOAD_GENES():
|
|
70
|
-
print('COPY
|
|
70
|
+
print('COPY gene_expression.gene (gene_id, display_gene_id, species, taxon_id) FROM stdin;')
|
|
71
71
|
GENE_SOURCE = '../script-results/processed-expression/genes.csv'
|
|
72
72
|
with open(GENE_SOURCE, 'r+') as f:
|
|
73
73
|
reader = csv.reader(f)
|
|
@@ -87,7 +87,7 @@ def LOAD_GENES():
|
|
|
87
87
|
This program Loads Expression Metadata into the database
|
|
88
88
|
"""
|
|
89
89
|
def LOAD_EXPRESSION_METADATA():
|
|
90
|
-
print('COPY
|
|
90
|
+
print('COPY gene_expression.expression_metadata (ncbi_geo_id, pubmed_id, control_yeast_strain, treatment_yeast_strain, control, treatment, concentration_value, concentration_unit, time_value, time_unit, number_of_replicates, expression_table) FROM stdin;')
|
|
91
91
|
EXPRESSION_METADATA_SOURCE = '../script-results/processed-expression/expression-metadata.csv'
|
|
92
92
|
with open(EXPRESSION_METADATA_SOURCE, 'r+') as f:
|
|
93
93
|
reader = csv.reader(f)
|
|
@@ -116,7 +116,7 @@ def LOAD_EXPRESSION_METADATA():
|
|
|
116
116
|
This program Loads Expression Data into the database
|
|
117
117
|
"""
|
|
118
118
|
def LOAD_EXPRESSION_DATA():
|
|
119
|
-
print('COPY
|
|
119
|
+
print('COPY gene_expression.expression (gene_id, taxon_id, sort_index, sample_id, expression, time_point, dataset) FROM stdin;')
|
|
120
120
|
EXPRESSION_DATA_SOURCE = '../script-results/processed-expression/expression-data.csv'
|
|
121
121
|
with open(EXPRESSION_DATA_SOURCE, 'r+') as f:
|
|
122
122
|
reader = csv.reader(f)
|
|
@@ -140,7 +140,7 @@ def LOAD_EXPRESSION_DATA():
|
|
|
140
140
|
This program Loads Production Rates into the database
|
|
141
141
|
"""
|
|
142
142
|
def LOAD_PRODUCTION_RATES():
|
|
143
|
-
print('COPY
|
|
143
|
+
print('COPY gene_expression.production_rate (gene_id, taxon_id, ncbi_geo_id, pubmed_id, production_rate) FROM stdin;')
|
|
144
144
|
PRODUCTION_RATES_SOURCE = '../script-results/processed-expression/production-rates.csv'
|
|
145
145
|
with open(PRODUCTION_RATES_SOURCE, 'r+') as f:
|
|
146
146
|
reader = csv.reader(f)
|
|
@@ -161,7 +161,7 @@ def LOAD_PRODUCTION_RATES():
|
|
|
161
161
|
This program Loads Degradation Rates into the database
|
|
162
162
|
"""
|
|
163
163
|
def LOAD_DEGRADATION_RATES():
|
|
164
|
-
print('COPY
|
|
164
|
+
print('COPY gene_expression.degradation_rate (gene_id, taxon_id, ncbi_geo_id, pubmed_id, degradation_rate) FROM stdin;')
|
|
165
165
|
DEGRADATION_RATES_SOURCE = '../script-results/processed-expression/degradation-rates.csv'
|
|
166
166
|
with open(DEGRADATION_RATES_SOURCE, 'r+') as f:
|
|
167
167
|
reader = csv.reader(f)
|
|
@@ -6,9 +6,9 @@ All files pertaining the network database live within this directory.
|
|
|
6
6
|
|
|
7
7
|
### Schema
|
|
8
8
|
|
|
9
|
-
All network data is stored within the
|
|
9
|
+
All network data is stored within the gene_regulatory_network schema on our Postgres database.
|
|
10
10
|
|
|
11
|
-
The schema is located within this directory at the top level in the file `schema.sql`. It defines the tables located within the
|
|
11
|
+
The schema is located within this directory at the top level in the file `schema.sql`. It defines the tables located within the gene_regulatory_network schema.
|
|
12
12
|
|
|
13
13
|
Usage:
|
|
14
14
|
To load to local database
|
|
@@ -32,10 +32,13 @@ Within the scripts directory, there are the following files:
|
|
|
32
32
|
|
|
33
33
|
- `generate_network.py`
|
|
34
34
|
- `loader.py`
|
|
35
|
+
- `generate_new_network_verion.py`
|
|
36
|
+
- `loader_updates.py`
|
|
35
37
|
- `filter_genes.py`
|
|
36
38
|
- `generate_sgd_network_from_yeastract_network.py`
|
|
37
39
|
|
|
38
|
-
|
|
40
|
+
|
|
41
|
+
#### Network Generator (and data preprocessor) (FOR FRESH DATABASE INSTALLS ONLY)
|
|
39
42
|
|
|
40
43
|
This script (`generate_network.py`) is a two-for-one. It first uses the yeastmine service from the SGD database to query for all regulator genes relating to Saccharomyces cerevisiae. From there it gets all all of the targets for each regulator gene. We then construct two networks from these connections (a regulator by regulator matrix as well as a regulator by target matrix). We also construct the processed loader files, so that they are ready to load using `loader.py`.
|
|
41
44
|
|
|
@@ -47,7 +50,7 @@ Usage:
|
|
|
47
50
|
```
|
|
48
51
|
python3 generate_network.py
|
|
49
52
|
```
|
|
50
|
-
#### Database Loader
|
|
53
|
+
#### Database Loader (FOR FRESH DATABASE INSTALLS ONLY)
|
|
51
54
|
|
|
52
55
|
This script (`loader.py`) is to be used to load your preprocessed genes into the database.
|
|
53
56
|
|
|
@@ -62,6 +65,35 @@ To load to production database
|
|
|
62
65
|
```
|
|
63
66
|
python3 loader.py | psql <address to database>
|
|
64
67
|
```
|
|
68
|
+
#### Network Generator (and data preprocessor) (FOR UPDATES TO EXISTING DATABASE ONLY)
|
|
69
|
+
|
|
70
|
+
This script (`generate_new_network_verion.py`) is similar to its counterpart `generate_network.py`. It gets all existing genes in the database using the environment variable 'DB_URL'. You can set this environment variable on the terminal right before the command. It uses the yeastmine service from the SGD database to query for all regulator genes relating to Saccharomyces cerevisiae. From there it gets all all of the targets for each regulator gene. We then construct two networks from these connections (a regulator by regulator matrix as well as a regulator by target matrix). We then see if the genes in the newly constructed network have any updates (i.e a gene's standard name was set or a new gene was added to the database). We also construct the processed loader files, so that they are ready to load using `loader_updates.py`.
|
|
71
|
+
|
|
72
|
+
The resulting network matrices are located in `script-results/networks` and the resulting processed loader files are located within `script-results/processed-loader-files`
|
|
73
|
+
|
|
74
|
+
Make sure to have all dependencies installed beforehand or you will recieve errors. (pip3 install intermine, tzlocal, etc. [see file for all imports]
|
|
75
|
+
|
|
76
|
+
Usage:
|
|
77
|
+
```
|
|
78
|
+
DB_URL="postgresql://[<db_user>:<password>]@<address to database>/<database name>" python3 generate_new_network_version.py
|
|
79
|
+
```
|
|
80
|
+
#### Database Loader (FOR UPDATES TO EXISTING DATABASE ONLY)
|
|
81
|
+
|
|
82
|
+
This script (`loader_updates.py`) is to be used to load your preprocessed genes into the database.
|
|
83
|
+
|
|
84
|
+
This program generates direct SQL statements from the source files generated by the network generator in order to populate a relational database with those files’ data as well as make any needed updates to existing genes within the database. If necessary you will be prompted to enter a password.
|
|
85
|
+
|
|
86
|
+
Usage:
|
|
87
|
+
To load to local database
|
|
88
|
+
```
|
|
89
|
+
python3 loader_updates.py | psql postgresql://localhost/postgres
|
|
90
|
+
```
|
|
91
|
+
To load to production database
|
|
92
|
+
```
|
|
93
|
+
python3 loader_updates.py | psql -h <grnsight database link> -U <user> <database name>
|
|
94
|
+
|
|
95
|
+
```
|
|
96
|
+
|
|
65
97
|
|
|
66
98
|
#### Filter Genes (beta functionality, not tested)
|
|
67
99
|
|
|
@@ -1,11 +1,11 @@
|
|
|
1
|
-
CREATE TABLE
|
|
1
|
+
CREATE TABLE gene_regulatory_network.source (
|
|
2
2
|
time_stamp TIMESTAMP WITH TIME ZONE,
|
|
3
3
|
source VARCHAR,
|
|
4
|
-
|
|
4
|
+
display_name VARCHAR,
|
|
5
5
|
PRIMARY KEY(time_stamp, source)
|
|
6
6
|
);
|
|
7
7
|
|
|
8
|
-
CREATE TABLE
|
|
8
|
+
CREATE TABLE gene_regulatory_network.gene (
|
|
9
9
|
gene_id VARCHAR, -- systematic like name
|
|
10
10
|
display_gene_id VARCHAR, -- standard like name
|
|
11
11
|
species VARCHAR,
|
|
@@ -13,13 +13,13 @@ CREATE TABLE spring2022_network.gene (
|
|
|
13
13
|
regulator BOOLEAN,
|
|
14
14
|
PRIMARY KEY(gene_id, taxon_id)
|
|
15
15
|
);
|
|
16
|
-
CREATE TABLE
|
|
16
|
+
CREATE TABLE gene_regulatory_network.network (
|
|
17
17
|
regulator_gene_id VARCHAR,
|
|
18
18
|
target_gene_id VARCHAR,
|
|
19
19
|
taxon_id VARCHAR,
|
|
20
20
|
time_stamp TIMESTAMP WITH TIME ZONE,
|
|
21
21
|
source VARCHAR,
|
|
22
|
-
FOREIGN KEY (regulator_gene_id, taxon_id) REFERENCES
|
|
23
|
-
FOREIGN KEY (target_gene_id, taxon_id) REFERENCES
|
|
24
|
-
FOREIGN KEY (time_stamp, source) REFERENCES
|
|
22
|
+
FOREIGN KEY (regulator_gene_id, taxon_id) REFERENCES gene_regulatory_network.gene(gene_id, taxon_id),
|
|
23
|
+
FOREIGN KEY (target_gene_id, taxon_id) REFERENCES gene_regulatory_network.gene(gene_id, taxon_id),
|
|
24
|
+
FOREIGN KEY (time_stamp, source) REFERENCES gene_regulatory_network.source(time_stamp, source)
|
|
25
25
|
);
|
|
@@ -13,7 +13,7 @@ try:
|
|
|
13
13
|
port="5432",
|
|
14
14
|
database="postgres")
|
|
15
15
|
cursor = connection.cursor()
|
|
16
|
-
postgreSQL_select_Query = "select * from
|
|
16
|
+
postgreSQL_select_Query = "select * from gene_regulatory_network.gene"
|
|
17
17
|
|
|
18
18
|
cursor.execute(postgreSQL_select_Query)
|
|
19
19
|
print("Selecting rows from gene table using cursor.fetchall")
|
|
@@ -140,7 +140,7 @@ regulator_to_regulator_file.close()
|
|
|
140
140
|
# Source Table
|
|
141
141
|
|
|
142
142
|
SOURCE_DESTINATION = '../script-results/processed-loader-files/source.csv'
|
|
143
|
-
timestamp = datetime.datetime.now(datetime.timezone.utc)
|
|
143
|
+
timestamp = datetime.datetime.now(datetime.timezone.utc).replace(microsecond=0)
|
|
144
144
|
|
|
145
145
|
source = "YeastMine - Saccharomyces Genome Database"
|
|
146
146
|
display_name = "Yeastmine - SGD"
|
|
@@ -0,0 +1,223 @@
|
|
|
1
|
+
from __future__ import print_function
|
|
2
|
+
|
|
3
|
+
from intermine.webservice import Service
|
|
4
|
+
service = Service("https://yeastmine.yeastgenome.org/yeastmine/service")
|
|
5
|
+
from sqlalchemy import create_engine
|
|
6
|
+
|
|
7
|
+
import csv
|
|
8
|
+
import re
|
|
9
|
+
import sys
|
|
10
|
+
import os
|
|
11
|
+
import datetime
|
|
12
|
+
|
|
13
|
+
|
|
14
|
+
def get_all_genes():
|
|
15
|
+
db = create_engine(os.environ['DB_URL'])
|
|
16
|
+
|
|
17
|
+
with db.connect() as connection:
|
|
18
|
+
result_set = connection.execute(
|
|
19
|
+
f"""
|
|
20
|
+
SELECT display_gene_id, gene_id FROM gene_regulatory_network.gene;
|
|
21
|
+
""")
|
|
22
|
+
result = result_set.fetchall()
|
|
23
|
+
return list(result)
|
|
24
|
+
|
|
25
|
+
|
|
26
|
+
|
|
27
|
+
# Get Network Data from Yeastmine
|
|
28
|
+
|
|
29
|
+
query = service.new_query("Gene")
|
|
30
|
+
|
|
31
|
+
query.add_view(
|
|
32
|
+
"primaryIdentifier", "secondaryIdentifier", "symbol", "name", "sgdAlias",
|
|
33
|
+
"regulationSummary.summaryParagraph",
|
|
34
|
+
"regulationSummary.publications.pubMedId",
|
|
35
|
+
"regulationSummary.publications.citation"
|
|
36
|
+
)
|
|
37
|
+
query.outerjoin("regulationSummary.publications")
|
|
38
|
+
|
|
39
|
+
regulators = {}
|
|
40
|
+
all_genes = {}
|
|
41
|
+
print("COLLECTING REGULATORS\n")
|
|
42
|
+
for row in query.rows():
|
|
43
|
+
systematic_name = row["secondaryIdentifier"]
|
|
44
|
+
standard_name = row["symbol"]
|
|
45
|
+
if standard_name == None:
|
|
46
|
+
standard_name = systematic_name
|
|
47
|
+
|
|
48
|
+
regulators[standard_name] = systematic_name
|
|
49
|
+
all_genes[standard_name] = systematic_name
|
|
50
|
+
|
|
51
|
+
regulators_to_targets = {}
|
|
52
|
+
all_targets = {}
|
|
53
|
+
|
|
54
|
+
|
|
55
|
+
print("COLLECTING TARGETS\n")
|
|
56
|
+
for regulator in regulators:
|
|
57
|
+
query = service.new_query("Gene")
|
|
58
|
+
query.add_constraint("regulatoryRegions", "TFBindingSite")
|
|
59
|
+
query.add_view(
|
|
60
|
+
"regulatoryRegions.regulator.symbol",
|
|
61
|
+
"regulatoryRegions.regulator.secondaryIdentifier", "symbol",
|
|
62
|
+
"secondaryIdentifier", "regulatoryRegions.regEvidence.ontologyTerm.name",
|
|
63
|
+
"regulatoryRegions.regEvidence.ontologyTerm.identifier",
|
|
64
|
+
"regulatoryRegions.experimentCondition",
|
|
65
|
+
"regulatoryRegions.strainBackground",
|
|
66
|
+
"regulatoryRegions.regulationDirection",
|
|
67
|
+
"regulatoryRegions.publications.pubMedId", "regulatoryRegions.datasource",
|
|
68
|
+
"regulatoryRegions.annotationType"
|
|
69
|
+
)
|
|
70
|
+
query.add_sort_order("Gene.secondaryIdentifier", "ASC")
|
|
71
|
+
query.add_constraint("regulatoryRegions.regulator", "LOOKUP", regulator, "S. cerevisiae", code="A")
|
|
72
|
+
targets = {}
|
|
73
|
+
|
|
74
|
+
for row in query.rows():
|
|
75
|
+
target_systematic_name = row["secondaryIdentifier"]
|
|
76
|
+
target_standard_name = row["symbol"]
|
|
77
|
+
if target_standard_name == None:
|
|
78
|
+
target_standard_name = target_systematic_name
|
|
79
|
+
targets[target_standard_name] = target_systematic_name
|
|
80
|
+
all_targets[target_standard_name] = target_systematic_name
|
|
81
|
+
all_genes[target_standard_name] = target_systematic_name
|
|
82
|
+
|
|
83
|
+
regulators_to_targets[regulator] = { "systematic_name": regulators[regulator], "targets": targets}
|
|
84
|
+
|
|
85
|
+
|
|
86
|
+
|
|
87
|
+
def create_regulator_to_target_row(target, all_regulators):
|
|
88
|
+
result = "" + target
|
|
89
|
+
for regulator in all_regulators:
|
|
90
|
+
if target in all_regulators[regulator]["targets"]:
|
|
91
|
+
result += "\t" + "1"
|
|
92
|
+
else:
|
|
93
|
+
result += "\t" + "0"
|
|
94
|
+
return result
|
|
95
|
+
|
|
96
|
+
|
|
97
|
+
# Create files
|
|
98
|
+
|
|
99
|
+
# Create folder paths
|
|
100
|
+
if not os.path.exists('../script-results'):
|
|
101
|
+
os.makedirs('../script-results')
|
|
102
|
+
|
|
103
|
+
if not os.path.exists('../script-results/networks'):
|
|
104
|
+
os.makedirs('../script-results/networks')
|
|
105
|
+
|
|
106
|
+
if not os.path.exists('../script-results/processed-loader-files'):
|
|
107
|
+
os.makedirs('../script-results/processed-loader-files')
|
|
108
|
+
|
|
109
|
+
|
|
110
|
+
|
|
111
|
+
# Files to be generated
|
|
112
|
+
|
|
113
|
+
# Generate Networks
|
|
114
|
+
|
|
115
|
+
REGULATORS_TO_TARGETS_MATRIX = '../script-results/networks/regulators_to_targets.csv'
|
|
116
|
+
REGULATORS_TO_REGULATORS_MATRIX = '../script-results/networks/regulators_to_regulators.csv'
|
|
117
|
+
|
|
118
|
+
|
|
119
|
+
targets = []
|
|
120
|
+
for target in all_targets:
|
|
121
|
+
if target != None:
|
|
122
|
+
targets.append(target)
|
|
123
|
+
|
|
124
|
+
regulators_list = []
|
|
125
|
+
for regulator in regulators_to_targets:
|
|
126
|
+
if regulator != None:
|
|
127
|
+
regulators_list.append(regulator)
|
|
128
|
+
|
|
129
|
+
print(f'Creating REGULATORS TO TARGETS MATRIX\n')
|
|
130
|
+
regulator_to_target_file = open(REGULATORS_TO_TARGETS_MATRIX, 'w')
|
|
131
|
+
headers = "cols regulators/rows targets"
|
|
132
|
+
headers += '\t'.join(regulators_list)
|
|
133
|
+
regulator_to_target_file.write(f'{headers}\n')
|
|
134
|
+
for target in targets:
|
|
135
|
+
result = create_regulator_to_target_row(target, regulators_to_targets)
|
|
136
|
+
if result != False:
|
|
137
|
+
regulator_to_target_file.write(f'{result}\n')
|
|
138
|
+
regulator_to_target_file.close()
|
|
139
|
+
|
|
140
|
+
print(f'Creating REGULATORS TO TARGETS MATRIX\n')
|
|
141
|
+
regulator_to_regulator_file = open(REGULATORS_TO_REGULATORS_MATRIX, 'w')
|
|
142
|
+
headers = "cols regulators/rows targets"
|
|
143
|
+
headers += '\t'.join(regulators_list)
|
|
144
|
+
regulator_to_regulator_file.write(f'{headers}\n')
|
|
145
|
+
for target in targets:
|
|
146
|
+
result = create_regulator_to_target_row(target, regulators_to_targets)
|
|
147
|
+
if result != False:
|
|
148
|
+
regulator_to_regulator_file.write(f'{result}\n')
|
|
149
|
+
regulator_to_regulator_file.close()
|
|
150
|
+
|
|
151
|
+
|
|
152
|
+
|
|
153
|
+
# Create loader-files
|
|
154
|
+
|
|
155
|
+
# Source Table
|
|
156
|
+
|
|
157
|
+
SOURCE_DESTINATION = '../script-results/processed-loader-files/source.csv'
|
|
158
|
+
timestamp = datetime.datetime.now(datetime.timezone.utc).replace(microsecond=0)
|
|
159
|
+
|
|
160
|
+
source = "YeastMine - Saccharomyces Genome Database"
|
|
161
|
+
display_name = "Yeastmine - SGD"
|
|
162
|
+
|
|
163
|
+
source_file = open(SOURCE_DESTINATION, 'w')
|
|
164
|
+
headers = f'Timestamp\tSource\tDisplay Name\n{timestamp}\t{source}\t{display_name}'
|
|
165
|
+
source_file.write(f'{headers}\n')
|
|
166
|
+
source_file.close()
|
|
167
|
+
|
|
168
|
+
# Gene Table
|
|
169
|
+
|
|
170
|
+
GENE_DESTINATION = '../script-results/processed-loader-files/gene.csv'
|
|
171
|
+
|
|
172
|
+
database_genes = {g[1] : g[0] for g in get_all_genes()}
|
|
173
|
+
database_standard_names = [g[0] for g in get_all_genes()]
|
|
174
|
+
database_systematic_names = [g[1] for g in get_all_genes()]
|
|
175
|
+
species = "Saccharomyces cerevisiae"
|
|
176
|
+
taxon_id = "559292"
|
|
177
|
+
genes_to_update = {}
|
|
178
|
+
print(f'Creating gene.csv\n')
|
|
179
|
+
gene_file = open(GENE_DESTINATION, 'w')
|
|
180
|
+
headers = f'Gene ID\tDisplay Gene ID\tSpecies\tTaxon ID\tRegulator'
|
|
181
|
+
gene_file.write(f'{headers}\n')
|
|
182
|
+
for gene in all_genes:
|
|
183
|
+
if gene not in database_standard_names:
|
|
184
|
+
if all_genes[gene] not in database_systematic_names:
|
|
185
|
+
if gene in regulators:
|
|
186
|
+
gene_file.write(f'{all_genes[gene]}\t{gene}\t{species}\t{taxon_id}\ttrue\n')
|
|
187
|
+
else:
|
|
188
|
+
gene_file.write(f'{all_genes[gene]}\t{gene}\t{species}\t{taxon_id}\tfalse\n')
|
|
189
|
+
else:
|
|
190
|
+
# if systematic name is found, then we need to update the gene
|
|
191
|
+
genes_to_update[gene] = all_genes[gene]
|
|
192
|
+
gene_file.close()
|
|
193
|
+
|
|
194
|
+
# Gene Table Updates
|
|
195
|
+
|
|
196
|
+
GENE_UPDATES_DESTINATION = '../script-results/processed-loader-files/gene_update.csv'
|
|
197
|
+
|
|
198
|
+
|
|
199
|
+
print(f'Creating gene_update.csv\n')
|
|
200
|
+
gene_file = open(GENE_UPDATES_DESTINATION, 'w')
|
|
201
|
+
headers = f'Gene ID\tDisplay Gene ID\tRegulator'
|
|
202
|
+
gene_file.write(f'{headers}\n')
|
|
203
|
+
for gene in genes_to_update:
|
|
204
|
+
if gene in regulators:
|
|
205
|
+
gene_file.write(f'{all_genes[gene]}\t{gene}\ttrue\n')
|
|
206
|
+
else:
|
|
207
|
+
gene_file.write(f'{all_genes[gene]}\t{gene}\tfalse\n')
|
|
208
|
+
gene_file.close()
|
|
209
|
+
|
|
210
|
+
|
|
211
|
+
# Network Table
|
|
212
|
+
|
|
213
|
+
NETWORK_DESTINATION = '../script-results/processed-loader-files/network.csv'
|
|
214
|
+
|
|
215
|
+
|
|
216
|
+
print(f'Creating network.csv\n')
|
|
217
|
+
network_file = open(NETWORK_DESTINATION, 'w')
|
|
218
|
+
headers = f'Regulator Gene ID\tTarget Gene ID\tTaxon ID\tTimestamp\tSource'
|
|
219
|
+
network_file.write(f'{headers}\n')
|
|
220
|
+
for gene in regulators_to_targets:
|
|
221
|
+
for target_gene in regulators_to_targets[gene]["targets"]:
|
|
222
|
+
network_file.write(f'{regulators_to_targets[gene]["systematic_name"]}\t{regulators_to_targets[gene]["targets"][target_gene]}\t{taxon_id}\t{timestamp}\t{source}\n')
|
|
223
|
+
network_file.close()
|
|
@@ -15,7 +15,7 @@ directly into a database command line utility such as `psql`.
|
|
|
15
15
|
This function Loads Network Data Sources into the database
|
|
16
16
|
"""
|
|
17
17
|
def LOAD_SOURCES():
|
|
18
|
-
print('COPY
|
|
18
|
+
print('COPY gene_regulatory_network.source (time_stamp, source, display_name) FROM stdin;')
|
|
19
19
|
NETWORK_DATA_SOURCE = '../script-results/processed-loader-files/source.csv'
|
|
20
20
|
with open(NETWORK_DATA_SOURCE, 'r+') as f:
|
|
21
21
|
reader = csv.reader(f)
|
|
@@ -35,7 +35,7 @@ def LOAD_SOURCES():
|
|
|
35
35
|
This function Loads Gene ID Mapping into the database
|
|
36
36
|
"""
|
|
37
37
|
def LOAD_GENES():
|
|
38
|
-
print('COPY
|
|
38
|
+
print('COPY gene_regulatory_network.gene (gene_id, display_gene_id, species, taxon_id, regulator) FROM stdin;')
|
|
39
39
|
GENE_SOURCE = '../script-results/processed-loader-files/gene.csv'
|
|
40
40
|
with open(GENE_SOURCE, 'r+') as f:
|
|
41
41
|
reader = csv.reader(f)
|
|
@@ -57,7 +57,7 @@ def LOAD_GENES():
|
|
|
57
57
|
This function Loads the Network Matrix into the database
|
|
58
58
|
"""
|
|
59
59
|
def LOAD_NETWORK():
|
|
60
|
-
print('COPY
|
|
60
|
+
print('COPY gene_regulatory_network.network (regulator_gene_id, target_gene_id, taxon_id, time_stamp, source) FROM stdin;')
|
|
61
61
|
NETWORK_SOURCE = '../script-results/processed-loader-files/network.csv'
|
|
62
62
|
with open(NETWORK_SOURCE, 'r+') as f:
|
|
63
63
|
reader = csv.reader(f)
|