graphreduce 0.1__py3.9.egg → 1.2__py3.9.egg
This diff represents the content of publicly available package versions that have been released to one of the supported registries. The information contained in this diff is provided for informational purposes only and reflects changes between package versions as they appear in their respective public registries.
- EGG-INFO/PKG-INFO +101 -10
- EGG-INFO/requires.txt +5 -4
- graphreduce/__pycache__/enum.cpython-39.pyc +0 -0
- graphreduce/__pycache__/graph_reduce.cpython-39.pyc +0 -0
- graphreduce/__pycache__/node.cpython-39.pyc +0 -0
- graphreduce/graph_reduce.py +76 -17
- graphreduce/node.py +210 -28
EGG-INFO/PKG-INFO
CHANGED
|
@@ -1,6 +1,6 @@
|
|
|
1
1
|
Metadata-Version: 2.1
|
|
2
2
|
Name: graphreduce
|
|
3
|
-
Version:
|
|
3
|
+
Version: 1.2
|
|
4
4
|
Summary: Leveraging graph data structures for complex feature engineering pipelines.
|
|
5
5
|
Home-page: https://github.com/wesmadrigal/graphreduce
|
|
6
6
|
Author: Wes Madrigal
|
|
@@ -23,9 +23,9 @@ Description-Content-Type: text/markdown
|
|
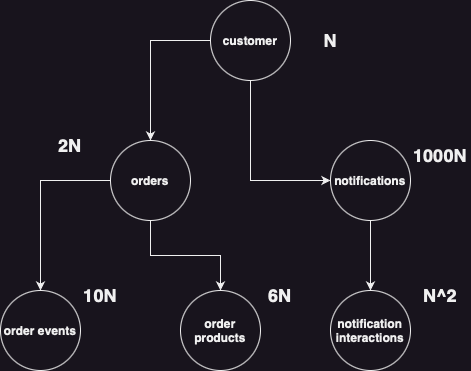
|
23
23
|
# GraphReduce
|
|
24
24
|
|
|
25
25
|
|
|
26
|
-
##
|
|
26
|
+
## Description
|
|
27
27
|
GraphReduce is an abstraction for building machine learning feature
|
|
28
|
-
engineering pipelines
|
|
28
|
+
engineering pipelines that involve many tables in a composable way.
|
|
29
29
|
The library is intended to help bridge the gap between research feature
|
|
30
30
|
definitions and production deployment without the overhead of a full
|
|
31
31
|
feature store. Underneath the hood, GraphReduce uses graph data
|
|
@@ -35,17 +35,108 @@ as edges.
|
|
|
35
35
|
GraphReduce allows for a unified feature engineering interface
|
|
36
36
|
to plug & play with multiple backends: `dask`, `pandas`, and `spark` are currently supported
|
|
37
37
|
|
|
38
|
-
## Motivation
|
|
39
|
-
As the number of features in an ML experiment grows so does the likelihood
|
|
40
|
-
for duplicate, one off implementations of the same code. This is further
|
|
41
|
-
exacerbated if there isn't seamless integration between R&D and deployment.
|
|
42
|
-
Feature stores are a good solution, but they are quite complicated to setup
|
|
43
|
-
and manage. GraphReduce is a lighter weight design pattern to production ready
|
|
44
|
-
feature engineering pipelines.
|
|
45
38
|
|
|
46
39
|
### Installation
|
|
47
40
|
```
|
|
41
|
+
# from pypi
|
|
42
|
+
pip install graphreduce
|
|
43
|
+
|
|
44
|
+
# from github
|
|
48
45
|
pip install 'graphreduce@git+https://github.com/wesmadrigal/graphreduce.git'
|
|
46
|
+
|
|
47
|
+
# install from source
|
|
48
|
+
git clone https://github.com/wesmadrigal/graphreduce && cd graphreduce && python setup.py install
|
|
49
|
+
```
|
|
50
|
+
|
|
51
|
+
|
|
52
|
+
|
|
53
|
+
## Motivation
|
|
54
|
+
Machine learning requires [vectors of data](https://arxiv.org/pdf/1212.4569.pdf), but our tabular datasets
|
|
55
|
+
are disconnected. They can be represented as a graph, where tables
|
|
56
|
+
are nodes and join keys are edges. In many model building scenarios
|
|
57
|
+
there isn't a nice ML-ready vector waiting for us, so we must curate
|
|
58
|
+
the data by joining many tables together to flatten them into a vector.
|
|
59
|
+
This is the problem `graphreduce` sets out to solve.
|
|
60
|
+
|
|
61
|
+
An example dataset might look like the following:
|
|
62
|
+
|
|
63
|
+
|
|
64
|
+
|
|
65
|
+
## data granularity and time travel
|
|
66
|
+
But we need to flatten this to a specific [granularity](https://en.wikipedia.org/wiki/Granularity#Data_granularity).
|
|
67
|
+
To further complicate things we need to handle orientation in time to prevent
|
|
68
|
+
[data leakage](https://en.wikipedia.org/wiki/Leakage_(machine_learning)) and properly frame our train/test datasets. All of this
|
|
69
|
+
is controlled in `graphreduce` from top-level parameters.
|
|
70
|
+
|
|
71
|
+
### example of granularity and time travel parameters
|
|
72
|
+
|
|
73
|
+
* `cut_date` controls the date around which we orient the data in the graph
|
|
74
|
+
* `compute_period_val` controls the amount of time back in history we consider during compute over the graph
|
|
75
|
+
* `compute_period_unit` tells us what unit of time we're using
|
|
76
|
+
* `parent_node` specifies the parent-most node in the graph and, typically, the granularity to which to reduce the data
|
|
77
|
+
```python
|
|
78
|
+
from graphreduce.graph_reduce import GraphReduce
|
|
79
|
+
from graphreduce.enums import PeriodUnit
|
|
80
|
+
|
|
81
|
+
gr = GraphReduce(
|
|
82
|
+
cut_date=datetime.datetime(2023, 2, 1),
|
|
83
|
+
compute_period_val=365,
|
|
84
|
+
compute_period_unit=PeriodUnit.day,
|
|
85
|
+
parent_node=customer
|
|
86
|
+
)
|
|
87
|
+
```
|
|
88
|
+
|
|
89
|
+
### Node definition and parameterization
|
|
90
|
+
GraphReduce takes convention over configuration, so the user
|
|
91
|
+
is required to define a number of methods on each node class:
|
|
92
|
+
* `do_annotate` annotation definitions (e.g., split a string column into a new column)
|
|
93
|
+
* `do_filters` filter the data on column(s)
|
|
94
|
+
* `do_clip_cols` clip anomalies like exceedingly large values and do normalization
|
|
95
|
+
* `post_join_annotate` annotations on current node after relations are merged in and we have access to their columns, too
|
|
96
|
+
* `do_reduce` the most import node function, reduction operations: group bys, sum, min, max, etc.
|
|
97
|
+
* `do_labels` label definitions if any
|
|
98
|
+
At the instance level we need to parameterize a few things, such as where the
|
|
99
|
+
data is coming from, the date key, the primary key, prefixes for
|
|
100
|
+
preserving where the data originated after compute, and a few
|
|
101
|
+
other optional parameters.
|
|
102
|
+
|
|
103
|
+
```python
|
|
104
|
+
from graphreduce.node import GraphReduceNode
|
|
105
|
+
|
|
106
|
+
# define the customer node
|
|
107
|
+
class CustomerNode(GraphReduceNode):
|
|
108
|
+
def do_annotate(self):
|
|
109
|
+
# use the `self.colabbr` function to use prefixes
|
|
110
|
+
self.df[self.colabbr('is_big_spender')] = self.df[self.colabbr('total_revenue')].apply(
|
|
111
|
+
lambda x: x > 1000.00 then 1 else 0
|
|
112
|
+
)
|
|
113
|
+
|
|
114
|
+
|
|
115
|
+
def do_filters(self):
|
|
116
|
+
self.df = self.df[self.df[self.colabbr('some_bool_col')] == 0]
|
|
117
|
+
|
|
118
|
+
def do_clip_cols(self):
|
|
119
|
+
self.df[self.colabbr('high_variance_column')] = self.df[self.colabbr('high_variance_column')].apply(
|
|
120
|
+
lambda col: 1000 if col > 1000 else col
|
|
121
|
+
)
|
|
122
|
+
|
|
123
|
+
def post_join_annotate(self):
|
|
124
|
+
# filters after children are joined
|
|
125
|
+
pass
|
|
126
|
+
|
|
127
|
+
def do_reduce(self, reduce_key):
|
|
128
|
+
pass
|
|
129
|
+
|
|
130
|
+
def do_labels(self, reduce_key):
|
|
131
|
+
pass
|
|
132
|
+
|
|
133
|
+
cust = CustomerNode(
|
|
134
|
+
fpath='s3://somebucket/some/path/customer.parquet',
|
|
135
|
+
fmt='parquet',
|
|
136
|
+
prefix='cust',
|
|
137
|
+
date_key='last_login',
|
|
138
|
+
pk='customer_id'
|
|
139
|
+
)
|
|
49
140
|
```
|
|
50
141
|
|
|
51
142
|
## Usage
|
EGG-INFO/requires.txt
CHANGED
|
Binary file
|
|
Binary file
|
|
Binary file
|
graphreduce/graph_reduce.py
CHANGED
|
@@ -39,6 +39,15 @@ class GraphReduce(nx.DiGraph):
|
|
|
39
39
|
label_period_unit : typing.Optional[PeriodUnit] = None,
|
|
40
40
|
spark_sqlCtx : pyspark.sql.SQLContext = None,
|
|
41
41
|
feature_function : typing.Optional[str] = None,
|
|
42
|
+
dynamic_propagation : bool = False,
|
|
43
|
+
type_func_map : typing.Dict[str, typing.List[str]] = {
|
|
44
|
+
'int64' : ['min', 'max', 'sum'],
|
|
45
|
+
'str' : ['first'],
|
|
46
|
+
'object' : ['first'],
|
|
47
|
+
'float64' : ['min', 'max', 'sum'],
|
|
48
|
+
'bool' : ['first'],
|
|
49
|
+
'datetime64' : ['first']
|
|
50
|
+
},
|
|
42
51
|
*args,
|
|
43
52
|
**kwargs
|
|
44
53
|
):
|
|
@@ -58,6 +67,8 @@ Args:
|
|
|
58
67
|
label_period_unit : the unit for the label period value (e.g., day)
|
|
59
68
|
spark_sqlCtx : if compute layer is spark this must be passed
|
|
60
69
|
feature_function : optional custom feature function
|
|
70
|
+
dynamic_propagation : optional to dynamically propagate children data upward, useful for very large compute graphs
|
|
71
|
+
type_func_match : optional mapping from type to a list of functions (e.g., {'int' : ['min', 'max', 'sum'], 'str' : ['first']})
|
|
61
72
|
"""
|
|
62
73
|
super(GraphReduce, self).__init__(*args, **kwargs)
|
|
63
74
|
|
|
@@ -72,6 +83,8 @@ Args:
|
|
|
72
83
|
self.label_period_unit = label_period_unit
|
|
73
84
|
self.compute_layer = compute_layer
|
|
74
85
|
self.feature_function = feature_function
|
|
86
|
+
self.dynamic_propagation = dynamic_propagation
|
|
87
|
+
self.type_func_map = type_func_map
|
|
75
88
|
|
|
76
89
|
# if using Spark
|
|
77
90
|
self._sqlCtx = spark_sqlCtx
|
|
@@ -165,7 +178,7 @@ Add an entity relation
|
|
|
165
178
|
}
|
|
166
179
|
)
|
|
167
180
|
|
|
168
|
-
|
|
181
|
+
|
|
169
182
|
def join (
|
|
170
183
|
self,
|
|
171
184
|
parent_node : GraphReduceNode,
|
|
@@ -218,9 +231,16 @@ Add an entity relation
|
|
|
218
231
|
return joined
|
|
219
232
|
|
|
220
233
|
elif self.compute_layer == ComputeLayerEnum.spark:
|
|
221
|
-
if isinstance(
|
|
234
|
+
if isinstance(relation_df, pyspark.sql.dataframe.DataFrame) and isinstance(parent_node.df, pyspark.sql.dataframe.DataFrame):
|
|
235
|
+
joined = parent_node.df.join(
|
|
236
|
+
relation_df,
|
|
237
|
+
on=parent_node.df[f"{parent_node.prefix}_{parent_pk}"] == relation_df[f"{relation_node.prefix}_{relation_fk}"],
|
|
238
|
+
how="left"
|
|
239
|
+
)
|
|
240
|
+
return joined
|
|
241
|
+
elif isinstance(parent_node.df, pyspark.sql.dataframe.DataFrame) and isinstance(relation_node.df, pyspark.sql.dataframe.DataFrame):
|
|
222
242
|
joined = parent_node.df.join(
|
|
223
|
-
|
|
243
|
+
relation_node.df,
|
|
224
244
|
on=parent_node.df[f"{parent_node.prefix}_{parent_pk}"] == relation_node.df[f"{relation_node.prefix}_{relation_fk}"],
|
|
225
245
|
how="left"
|
|
226
246
|
)
|
|
@@ -258,8 +278,6 @@ Get the children of a given node
|
|
|
258
278
|
def plot_graph (
|
|
259
279
|
self,
|
|
260
280
|
fname : str = 'graph.html',
|
|
261
|
-
notebook : bool = False,
|
|
262
|
-
cdn_resources : str = 'in_line',
|
|
263
281
|
):
|
|
264
282
|
"""
|
|
265
283
|
Plot the graph
|
|
@@ -267,7 +285,6 @@ Plot the graph
|
|
|
267
285
|
Args
|
|
268
286
|
fname : file name to save the graph to - should be .html
|
|
269
287
|
notebook : whether or not to render in notebook
|
|
270
|
-
cdn_resources : pyvis parameter https://pyvis.readthedocs.io/en/latest/tutorial.html
|
|
271
288
|
"""
|
|
272
289
|
# need to populate a new graph
|
|
273
290
|
# with string representations
|
|
@@ -286,25 +303,41 @@ Args
|
|
|
286
303
|
edge[1].__class__.__name__,
|
|
287
304
|
title=edge_title)
|
|
288
305
|
|
|
289
|
-
nt = pyvis.network.Network(
|
|
306
|
+
nt = pyvis.network.Network()
|
|
290
307
|
nt.from_nx(stringG)
|
|
291
308
|
logger.info(f"plotted graph at {fname}")
|
|
292
309
|
nt.show(fname)
|
|
293
|
-
|
|
294
|
-
|
|
310
|
+
|
|
311
|
+
|
|
312
|
+
def prefix_uniqueness(self):
|
|
313
|
+
"""
|
|
314
|
+
Identify children with duplicate prefixes, if any
|
|
315
|
+
"""
|
|
316
|
+
prefixes = {}
|
|
317
|
+
dupes = []
|
|
318
|
+
for node in self.nodes():
|
|
319
|
+
if not prefixes.get(node.prefix):
|
|
320
|
+
prefixes[node.prefix] = node
|
|
321
|
+
else:
|
|
322
|
+
dupes.append(node)
|
|
323
|
+
dupes.append(prefixes[node.prefix])
|
|
324
|
+
if len(dupes):
|
|
325
|
+
raise Exception(f"duplicate prefix on the following nodes: {dupes}")
|
|
326
|
+
|
|
295
327
|
|
|
296
328
|
def do_transformations(self):
|
|
297
329
|
"""
|
|
298
330
|
Perform all graph transformations
|
|
299
331
|
1) hydrate graph
|
|
300
|
-
2)
|
|
301
|
-
3)
|
|
302
|
-
4)
|
|
303
|
-
5)
|
|
304
|
-
|
|
305
|
-
|
|
306
|
-
|
|
307
|
-
|
|
332
|
+
2) check for duplicate prefixes
|
|
333
|
+
3) filter data
|
|
334
|
+
4) clip anomalies
|
|
335
|
+
5) annotate data
|
|
336
|
+
6) depth-first edge traversal to: aggregate / reduce features and labels
|
|
337
|
+
6a) optional alternative feature_function mapping
|
|
338
|
+
6b) join back to parent edge
|
|
339
|
+
6c) post-join annotations if any
|
|
340
|
+
7) repeat step 6 on all edges up the hierarchy
|
|
308
341
|
"""
|
|
309
342
|
|
|
310
343
|
# get data, filter data, clip columns, and annotate
|
|
@@ -312,6 +345,9 @@ Perform all graph transformations
|
|
|
312
345
|
self.hydrate_graph_attrs()
|
|
313
346
|
logger.info("hydrating graph data")
|
|
314
347
|
self.hydrate_graph_data()
|
|
348
|
+
|
|
349
|
+
logger.info("checking for prefix uniqueness")
|
|
350
|
+
self.prefix_uniqueness()
|
|
315
351
|
|
|
316
352
|
for node in self.nodes():
|
|
317
353
|
logger.info(f"running filters, clip cols, and annotations for {node.__class__.__name__}")
|
|
@@ -330,6 +366,29 @@ Perform all graph transformations
|
|
|
330
366
|
if edge_data['reduce']:
|
|
331
367
|
logger.info(f"reducing relation {relation_node.__class__.__name__}")
|
|
332
368
|
join_df = relation_node.do_reduce(edge_data['relation_key'])
|
|
369
|
+
# only relevant when reducing
|
|
370
|
+
if self.dynamic_propagation:
|
|
371
|
+
logger.info(f"doing dynamic propagation on node {relation_node.__class__.__name__}")
|
|
372
|
+
child_df = relation_node.dynamic_propagation(
|
|
373
|
+
reduce_key=edge_data['relation_key'],
|
|
374
|
+
type_func_map=self.type_func_map,
|
|
375
|
+
compute_layer=self.compute_layer
|
|
376
|
+
)
|
|
377
|
+
# NOTE: this is pandas specific and will break
|
|
378
|
+
# on other compute layers for now
|
|
379
|
+
if self.compute_layer in [ComputeLayerEnum.pandas, ComputeLayerEnum.dask]:
|
|
380
|
+
join_df = join_df.merge(
|
|
381
|
+
child_df,
|
|
382
|
+
on=relation_node.colabbr(edge_data['relation_key']),
|
|
383
|
+
suffixes=('', '_dupe')
|
|
384
|
+
)
|
|
385
|
+
elif self.compute_layer == ComputeLayerEnum.spark:
|
|
386
|
+
join_df = join_df.join(
|
|
387
|
+
child_df,
|
|
388
|
+
on=join_df[relation_node.colabbr(edge_data['relation_key'])] == child_df[relation_node.colabbr(edge_data['relation_key'])],
|
|
389
|
+
how="left"
|
|
390
|
+
)
|
|
391
|
+
|
|
333
392
|
elif not edge_data['reduce'] and self.feature_function:
|
|
334
393
|
logger.info(f"not reducing relation {relation_node.__class__.__name__}")
|
|
335
394
|
join_df = getattr(relation_node, self.feature_function)()
|
graphreduce/node.py
CHANGED
|
@@ -70,11 +70,17 @@ Args
|
|
|
70
70
|
feature_function : optional feature function, usually used when reduce is false
|
|
71
71
|
columns : optional list of columns to include
|
|
72
72
|
"""
|
|
73
|
+
# For when this is already set on the class definition.
|
|
74
|
+
if not hasattr(self, 'pk'):
|
|
75
|
+
self.pk = pk
|
|
76
|
+
# For when this is already set on the class definition.
|
|
77
|
+
if not hasattr(self, 'prefix'):
|
|
78
|
+
self.prefix = prefix
|
|
79
|
+
# For when this is already set on the class definition.
|
|
80
|
+
if not hasattr(self, 'date_key'):
|
|
81
|
+
self.date_key = date_key
|
|
73
82
|
self.fpath = fpath
|
|
74
83
|
self.fmt = fmt
|
|
75
|
-
self.pk = pk
|
|
76
|
-
self.prefix = prefix
|
|
77
|
-
self.date_key = date_key
|
|
78
84
|
self.compute_layer = compute_layer
|
|
79
85
|
self.cut_date = cut_date
|
|
80
86
|
self.compute_period_val = compute_period_val
|
|
@@ -86,7 +92,7 @@ Args
|
|
|
86
92
|
self.feature_function = feature_function
|
|
87
93
|
self.spark_sqlctx = spark_sqlctx
|
|
88
94
|
|
|
89
|
-
self.columns =
|
|
95
|
+
self.columns = columns
|
|
90
96
|
|
|
91
97
|
|
|
92
98
|
|
|
@@ -117,10 +123,26 @@ Get some data
|
|
|
117
123
|
self.df.columns = [f"{self.prefix}_{c}" for c in self.df.columns]
|
|
118
124
|
elif self.compute_layer.value == 'spark':
|
|
119
125
|
if not hasattr(self, 'df') or (hasattr(self, 'df') and not isinstance(self.df, pyspark.sql.DataFrame)):
|
|
120
|
-
self.df = getattr(self.spark_sqlctx.read, {self.fmt})(self.fpath)
|
|
126
|
+
self.df = getattr(self.spark_sqlctx.read, f"{self.fmt}")(self.fpath)
|
|
127
|
+
if self.columns:
|
|
128
|
+
self.df = self.df.select(self.columns)
|
|
121
129
|
for c in self.df.columns:
|
|
122
130
|
self.df = self.df.withColumnRenamed(c, f"{self.prefix}_{c}")
|
|
123
131
|
|
|
132
|
+
# at this point of connectors we may want to try integrating
|
|
133
|
+
# with something like fugue: https://github.com/fugue-project/fugue
|
|
134
|
+
elif self.compute_layer.value == 'ray':
|
|
135
|
+
pass
|
|
136
|
+
|
|
137
|
+
elif self.compute_layer.value == 'snowflake':
|
|
138
|
+
pass
|
|
139
|
+
|
|
140
|
+
elif self.compute_layer.value == 'postgres':
|
|
141
|
+
pass
|
|
142
|
+
|
|
143
|
+
elif self.compute_layer.value == 'redshift':
|
|
144
|
+
pass
|
|
145
|
+
|
|
124
146
|
|
|
125
147
|
@abc.abstractmethod
|
|
126
148
|
def do_filters (
|
|
@@ -134,30 +156,110 @@ do some filters on the data
|
|
|
134
156
|
|
|
135
157
|
@abc.abstractmethod
|
|
136
158
|
def do_annotate(self):
|
|
137
|
-
|
|
138
|
-
|
|
139
|
-
|
|
140
|
-
|
|
159
|
+
"""
|
|
160
|
+
Implement custom annotation functionality
|
|
161
|
+
for annotating this particular data
|
|
162
|
+
"""
|
|
141
163
|
return
|
|
142
164
|
|
|
143
165
|
|
|
144
166
|
@abc.abstractmethod
|
|
145
167
|
def do_post_join_annotate(self):
|
|
146
|
-
|
|
147
|
-
|
|
148
|
-
|
|
149
|
-
|
|
150
|
-
|
|
168
|
+
"""
|
|
169
|
+
Implement custom annotation functionality
|
|
170
|
+
for annotating data after joining with
|
|
171
|
+
child data
|
|
172
|
+
"""
|
|
151
173
|
pass
|
|
152
174
|
|
|
153
175
|
|
|
154
176
|
@abc.abstractmethod
|
|
155
177
|
def do_clip_cols(self):
|
|
156
178
|
return
|
|
157
|
-
|
|
179
|
+
|
|
180
|
+
|
|
181
|
+
def dynamic_propagation (
|
|
182
|
+
self,
|
|
183
|
+
reduce_key : str,
|
|
184
|
+
type_func_map : dict = {},
|
|
185
|
+
compute_layer : ComputeLayerEnum = ComputeLayerEnum.pandas,
|
|
186
|
+
):
|
|
187
|
+
"""
|
|
188
|
+
If we're doing dynamic propagation
|
|
189
|
+
this function will run a series of
|
|
190
|
+
automatic aggregations
|
|
191
|
+
"""
|
|
192
|
+
if compute_layer == ComputeLayerEnum.pandas:
|
|
193
|
+
return self.pandas_dynamic_propagation(reduce_key=reduce_key, type_func_map=type_func_map)
|
|
194
|
+
elif compute_layer == ComputeLayerEnum.dask:
|
|
195
|
+
return self.dask_dynamic_propagation(reduce_key=reduce_key, type_func_map=type_func_map)
|
|
196
|
+
elif compute_layer == ComputeLayerEnum.spark:
|
|
197
|
+
return self.spark_dynamic_propagation(reduce_key=reduce_key, type_func_map=type_func_map)
|
|
198
|
+
|
|
199
|
+
|
|
200
|
+
def pandas_dynamic_propagation (
|
|
201
|
+
self,
|
|
202
|
+
reduce_key : str,
|
|
203
|
+
type_func_map : dict = {}
|
|
204
|
+
) -> pd.DataFrame:
|
|
205
|
+
"""
|
|
206
|
+
Pandas implementation of dynamic propagation of features
|
|
207
|
+
This could be extended slightly to perform automated feature
|
|
208
|
+
aggregation on dynamic nodes
|
|
209
|
+
"""
|
|
210
|
+
agg_funcs = {}
|
|
211
|
+
for col, _type in dict(self.df.dtypes).items():
|
|
212
|
+
_type = str(_type)
|
|
213
|
+
if type_func_map.get(_type):
|
|
214
|
+
for func in type_func_map[_type]:
|
|
215
|
+
col_new = f"{col}_{func}"
|
|
216
|
+
agg_funcs[col_new] = pd.NamedAgg(column=col, aggfunc=func)
|
|
217
|
+
return self.prep_for_features().groupby(self.colabbr(reduce_key)).agg(
|
|
218
|
+
**agg_funcs
|
|
219
|
+
).reset_index()
|
|
220
|
+
|
|
221
|
+
|
|
222
|
+
def dask_dynamic_propagation (
|
|
223
|
+
self,
|
|
224
|
+
reduce_key : str,
|
|
225
|
+
type_func_map : dict = {},
|
|
226
|
+
) -> dd.DataFrame:
|
|
227
|
+
"""
|
|
228
|
+
Dask implementation of dynamic propagation of features
|
|
229
|
+
This could be extended slightly to perform automated
|
|
230
|
+
feature aggregation on dynamic nodes
|
|
231
|
+
"""
|
|
232
|
+
agg_funcs = {}
|
|
233
|
+
for col, _type in dict(self.df.dtypes).items():
|
|
234
|
+
_type = str(_type)
|
|
235
|
+
if type_func_map.get(_type):
|
|
236
|
+
for func in type_func_map[_type]:
|
|
237
|
+
col_new = f"{col}_{func}"
|
|
238
|
+
agg_funcs[col_new] = pd.NamedAgg(column=col, aggfunc=func)
|
|
239
|
+
return self.prep_for_features().groupby(self.colabbr(reduce_key)).agg(
|
|
240
|
+
**agg_funcs
|
|
241
|
+
).reset_index()
|
|
242
|
+
|
|
243
|
+
|
|
244
|
+
def spark_dynamic_propagation (
|
|
245
|
+
self,
|
|
246
|
+
reduce_key : str,
|
|
247
|
+
type_func_map : dict = {},
|
|
248
|
+
) -> pyspark.sql.DataFrame:
|
|
249
|
+
"""
|
|
250
|
+
Spark implementation of dynamic propagation of features
|
|
251
|
+
This could be extended slightly to perform automated
|
|
252
|
+
feature aggregation on dynamic nodes
|
|
253
|
+
"""
|
|
254
|
+
agg_funcs = {}
|
|
255
|
+
pass
|
|
256
|
+
|
|
158
257
|
|
|
159
258
|
@abc.abstractmethod
|
|
160
|
-
def do_reduce(
|
|
259
|
+
def do_reduce (
|
|
260
|
+
self,
|
|
261
|
+
reduce_key
|
|
262
|
+
):
|
|
161
263
|
"""
|
|
162
264
|
Reduce operation or the node
|
|
163
265
|
|
|
@@ -175,36 +277,76 @@ Args
|
|
|
175
277
|
|
|
176
278
|
def colabbr(self, col: str) -> str:
|
|
177
279
|
return f"{self.prefix}_{col}"
|
|
280
|
+
|
|
281
|
+
|
|
282
|
+
def compute_period_minutes (
|
|
283
|
+
self,
|
|
284
|
+
) -> int:
|
|
285
|
+
"""
|
|
286
|
+
Convert the compute period to minutes
|
|
287
|
+
"""
|
|
288
|
+
if self.compute_period_unit == PeriodUnit.second:
|
|
289
|
+
return self.compute_period_val / 60
|
|
290
|
+
elif self.compute_period_unit == PeriodUnit.minute:
|
|
291
|
+
return self.compute_period_val
|
|
292
|
+
elif self.compute_period_unit == PeriodUnit.hour:
|
|
293
|
+
return self.compute_period_val * 60
|
|
294
|
+
elif self.compute_period_unit == PeriodUnit.day:
|
|
295
|
+
return self.compute_period_val * 1440
|
|
296
|
+
elif self.compute_period_unit == PeriodUnit.week:
|
|
297
|
+
return (self.compute_period_val * 7)*1440
|
|
298
|
+
elif self.compute_period_unit == PeriodUnit.month:
|
|
299
|
+
return (self.compute_period_val * 30.417)*1440
|
|
300
|
+
|
|
301
|
+
|
|
302
|
+
def label_period_minutes (
|
|
303
|
+
self,
|
|
304
|
+
) -> int:
|
|
305
|
+
"""
|
|
306
|
+
Convert the label period to minutes
|
|
307
|
+
"""
|
|
308
|
+
if self.label_period_unit == PeriodUnit.second:
|
|
309
|
+
return self.label_period_val / 60
|
|
310
|
+
elif self.label_period_unit == PeriodUnit.minute:
|
|
311
|
+
return self.label_period_val
|
|
312
|
+
elif self.label_period_unit == PeriodUnit.hour:
|
|
313
|
+
return self.label_period_val * 60
|
|
314
|
+
elif self.label_period_unit == PeriodUnit.day:
|
|
315
|
+
return self.label_period_val * 1440
|
|
316
|
+
elif self.label_period_unit == PeriodUnit.week:
|
|
317
|
+
return (self.label_period_val * 7)*1440
|
|
318
|
+
elif self.label_period_unit == PeriodUnit.month:
|
|
319
|
+
return (self.label_period_val * 30.417)*1440
|
|
178
320
|
|
|
179
321
|
|
|
180
322
|
def prep_for_features(self):
|
|
181
323
|
"""
|
|
182
324
|
Prepare the dataset for feature aggregations / reduce
|
|
183
325
|
"""
|
|
184
|
-
if self.date_key:
|
|
326
|
+
if self.date_key:
|
|
185
327
|
if self.cut_date and isinstance(self.cut_date, str) or isinstance(self.cut_date, datetime.datetime):
|
|
186
328
|
if isinstance(self.df, pd.DataFrame) or isinstance(self.df, dd.DataFrame):
|
|
187
329
|
return self.df[
|
|
188
330
|
(self.df[self.colabbr(self.date_key)] < self.cut_date)
|
|
189
331
|
&
|
|
190
|
-
(self.df[self.colabbr(self.date_key)] > (self.cut_date - datetime.timedelta(
|
|
332
|
+
(self.df[self.colabbr(self.date_key)] > (self.cut_date - datetime.timedelta(minutes=self.compute_period_minutes())))
|
|
191
333
|
]
|
|
192
334
|
elif isinstance(self.df, pyspark.sql.dataframe.DataFrame):
|
|
193
335
|
return self.df.filter(
|
|
194
336
|
(self.df[self.colabbr(self.date_key)] < self.cut_date)
|
|
195
337
|
&
|
|
196
|
-
(self.df[self.colabbr(self.date_key)] > (self.cut_date - datetime.timedelta(
|
|
338
|
+
(self.df[self.colabbr(self.date_key)] > (self.cut_date - datetime.timedelta(minutes=self.compute_period_minutes())))
|
|
197
339
|
)
|
|
198
340
|
else:
|
|
199
341
|
if isinstance(self.df, pd.DataFrame) or isinstance(self.df, dd.DataFrame):
|
|
200
342
|
return self.df[
|
|
201
343
|
(self.df[self.colabbr(self.date_key)] < datetime.datetime.now())
|
|
202
344
|
&
|
|
203
|
-
(self.df[self.colabbr(self.date_key)] > (datetime.datetime.now() - datetime.timedelta(
|
|
345
|
+
(self.df[self.colabbr(self.date_key)] > (datetime.datetime.now() - datetime.timedelta(minutes=self.compute_period_minutes())))
|
|
204
346
|
]
|
|
205
347
|
elif isinstance(self.df, pyspark.sql.dataframe.DataFrame):
|
|
206
348
|
return self.df.filter(
|
|
207
|
-
self.df[self.colabbr(self.date_key)] > (datetime.datetime.now() - datetime.timedelta(
|
|
349
|
+
self.df[self.colabbr(self.date_key)] > (datetime.datetime.now() - datetime.timedelta(minutes=self.compute_period_minutes()))
|
|
208
350
|
)
|
|
209
351
|
# no-op
|
|
210
352
|
return self.df
|
|
@@ -212,29 +354,69 @@ Prepare the dataset for feature aggregations / reduce
|
|
|
212
354
|
|
|
213
355
|
def prep_for_labels(self):
|
|
214
356
|
"""
|
|
215
|
-
|
|
357
|
+
Prepare the dataset for labels
|
|
216
358
|
"""
|
|
217
359
|
if self.date_key:
|
|
218
360
|
if self.cut_date and isinstance(self.cut_date, str) or isinstance(self.cut_date, datetime.datetime):
|
|
219
361
|
if isinstance(self.df, pd.DataFrame):
|
|
220
362
|
return self.df[
|
|
221
|
-
(self.df[self.colabbr(self.date_key)] >
|
|
363
|
+
(self.df[self.colabbr(self.date_key)] > self.cut_date)
|
|
222
364
|
&
|
|
223
|
-
(self.df[self.colabbr(self.date_key)] < (self.cut_date + datetime.timedelta(
|
|
365
|
+
(self.df[self.colabbr(self.date_key)] < (self.cut_date + datetime.timedelta(minutes=self.label_period_minutes())))
|
|
224
366
|
]
|
|
225
367
|
elif isinstance(self.df, pyspark.sql.dataframe.DataFrame):
|
|
226
368
|
return self.df.filter(
|
|
227
|
-
(self.df[self.colabbr(self.date_key)] >
|
|
369
|
+
(self.df[self.colabbr(self.date_key)] > self.cut_date)
|
|
228
370
|
&
|
|
229
|
-
(self.df[self.colabbr(self.date_key)] < (self.
|
|
371
|
+
(self.df[self.colabbr(self.date_key)] < (self.cut_date + datetime.timedelta(minutes=self.label_period_minutes())))
|
|
230
372
|
)
|
|
231
373
|
else:
|
|
232
374
|
if isinstance(self.df, pd.DataFrame):
|
|
233
375
|
return self.df[
|
|
234
|
-
self.df[self.colabbr(self.date_key)] > (datetime.datetime.now() - datetime.timedelta(
|
|
376
|
+
self.df[self.colabbr(self.date_key)] > (datetime.datetime.now() - datetime.timedelta(minutes=self.label_period_minutes()))
|
|
235
377
|
]
|
|
236
378
|
elif isinstance(self.df, pyspark.sql.dataframe.DataFrame):
|
|
237
379
|
return self.df.filter(
|
|
238
|
-
self.df[self.colabbr(self.date_key)] > (datetime.datetime.now() - datetime.timedelta(
|
|
380
|
+
self.df[self.colabbr(self.date_key)] > (datetime.datetime.now() - datetime.timedelta(minutes=self.label_period_minutes()))
|
|
239
381
|
)
|
|
240
382
|
return self.df
|
|
383
|
+
|
|
384
|
+
|
|
385
|
+
|
|
386
|
+
|
|
387
|
+
|
|
388
|
+
|
|
389
|
+
class DynamicNode(GraphReduceNode):
|
|
390
|
+
"""
|
|
391
|
+
A dynamic architecture for entities with no logic
|
|
392
|
+
needed in addition to the top-level GraphReduceNode
|
|
393
|
+
parameters
|
|
394
|
+
"""
|
|
395
|
+
def __init__ (
|
|
396
|
+
self,
|
|
397
|
+
*args,
|
|
398
|
+
**kwargs
|
|
399
|
+
):
|
|
400
|
+
"""
|
|
401
|
+
Constructor
|
|
402
|
+
"""
|
|
403
|
+
super().__init__(*args, **kwargs)
|
|
404
|
+
|
|
405
|
+
|
|
406
|
+
def do_filters(self):
|
|
407
|
+
pass
|
|
408
|
+
|
|
409
|
+
def do_annotate(self):
|
|
410
|
+
pass
|
|
411
|
+
|
|
412
|
+
def do_post_join_annotate(self):
|
|
413
|
+
pass
|
|
414
|
+
|
|
415
|
+
def do_clip_cols(self):
|
|
416
|
+
pass
|
|
417
|
+
|
|
418
|
+
def do_reduce(self):
|
|
419
|
+
pass
|
|
420
|
+
|
|
421
|
+
def do_labels(self):
|
|
422
|
+
pass
|