seqera 0.5.0 → 0.7.0-dev.2f7b343
This diff represents the content of publicly available package versions that have been released to one of the supported registries. The information contained in this diff is provided for informational purposes only and reflects changes between package versions as they appear in their respective public registries.
- package/README.md +24 -29
- package/package.json +4 -4
package/README.md
CHANGED
|
@@ -2,16 +2,12 @@
|
|
|
2
2
|
|
|
3
3
|
AI-powered assistant for bioinformatics workflows and [Seqera Platform](https://seqera.io).
|
|
4
4
|
|
|
5
|
-
> **Beta:** Seqera AI CLI is currently in beta. Features and commands may change as we continue to improve the product.
|
|
6
|
-
|
|
7
|
-
> **Note:** Seqera Cloud users receive $20 in free credits to get started with Seqera AI. [Contact us](https://seqera.io/platform/seqera-ai/request-credits/) for additional credits.
|
|
8
|
-
|
|
9
5
|
## Get started
|
|
10
6
|
|
|
11
7
|
1. Install the Seqera AI CLI:
|
|
12
8
|
|
|
13
9
|
```bash
|
|
14
|
-
|
|
10
|
+
npm install -g seqera
|
|
15
11
|
```
|
|
16
12
|
|
|
17
13
|
See [Installation](https://docs.seqera.io/platform-cloud/seqera-ai/installation) for a comprehensive installation guide.
|
|
@@ -33,7 +29,7 @@ AI-powered assistant for bioinformatics workflows and [Seqera Platform](https://
|
|
|
33
29
|
4. Run your first prompt:
|
|
34
30
|
|
|
35
31
|
```
|
|
36
|
-
|
|
32
|
+
List my recent workflows
|
|
37
33
|
```
|
|
38
34
|
|
|
39
35
|
See [Use cases](https://docs.seqera.io/platform-cloud/seqera-ai/use-cases) for a comprehensive list of use cases.
|
|
@@ -46,8 +42,6 @@ Seqera AI is an intelligent command-line assistant that helps you build, run, an
|
|
|
46
42
|
|
|
47
43
|
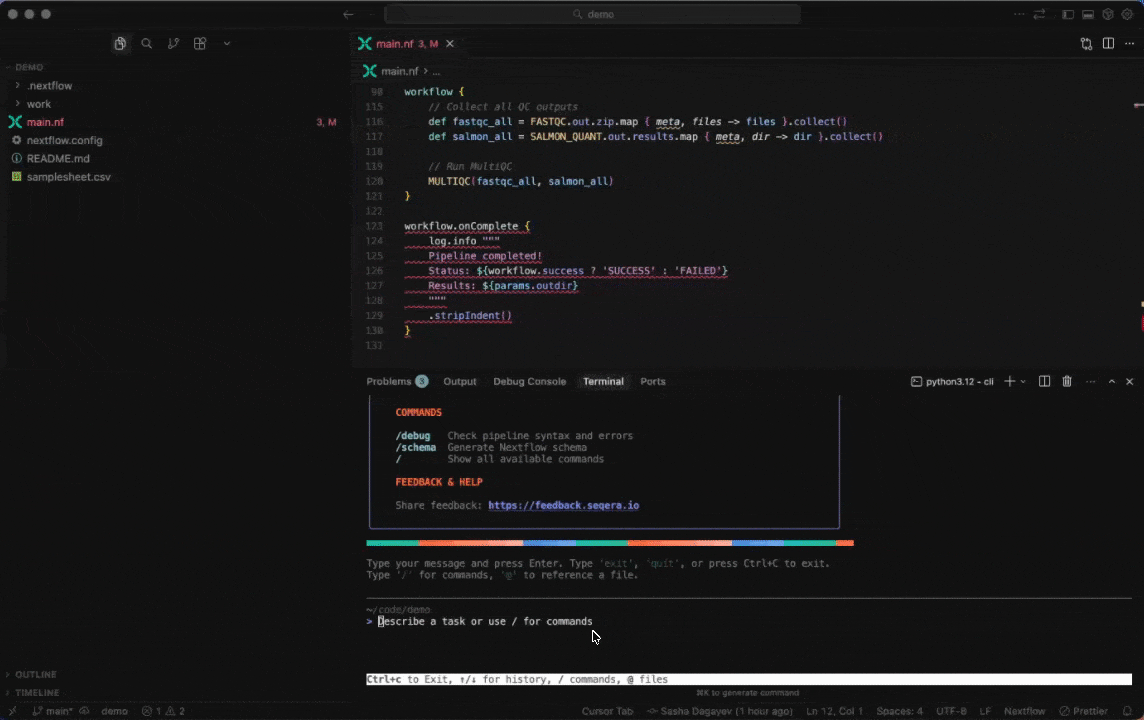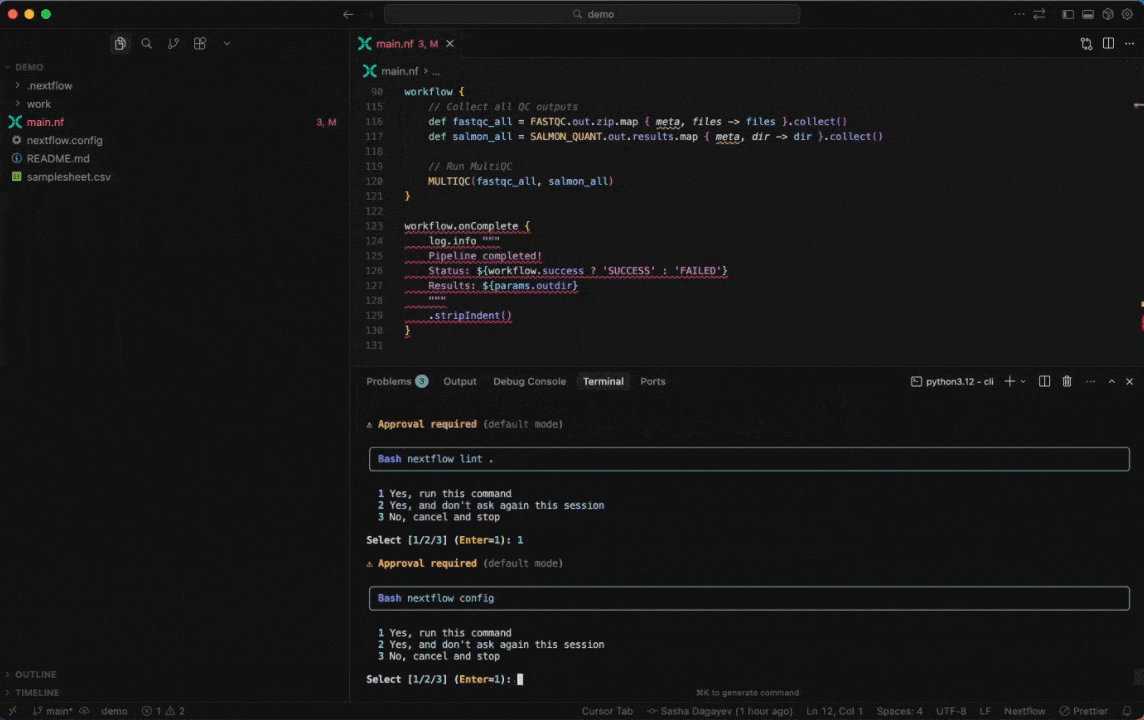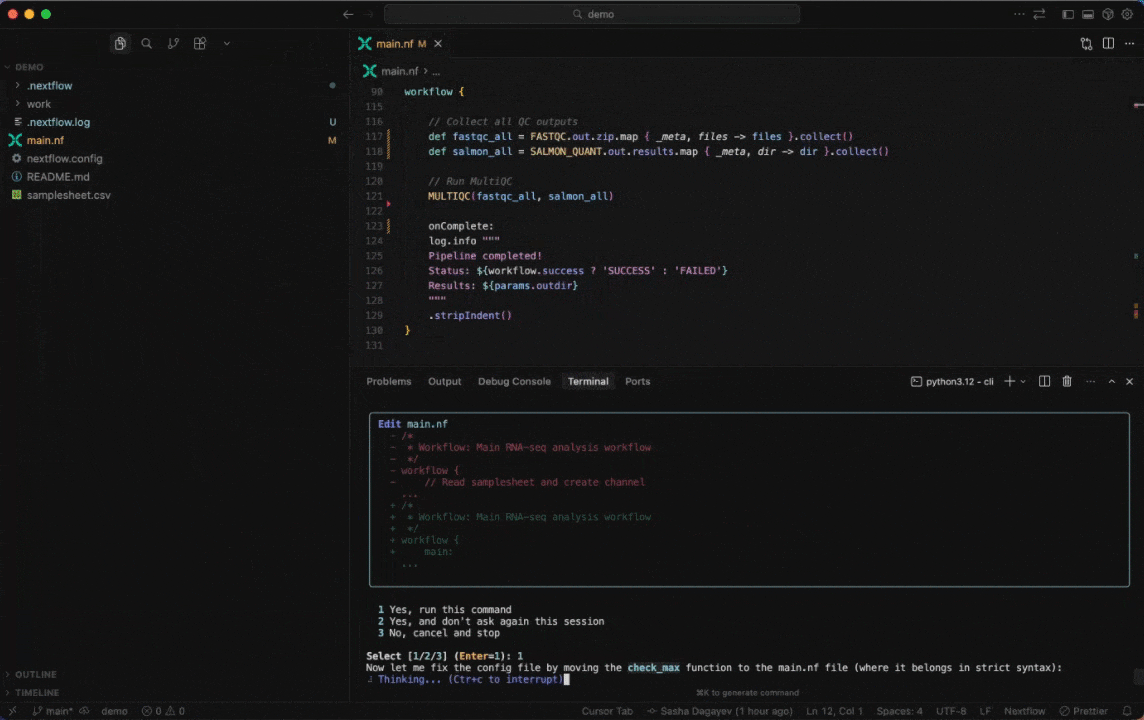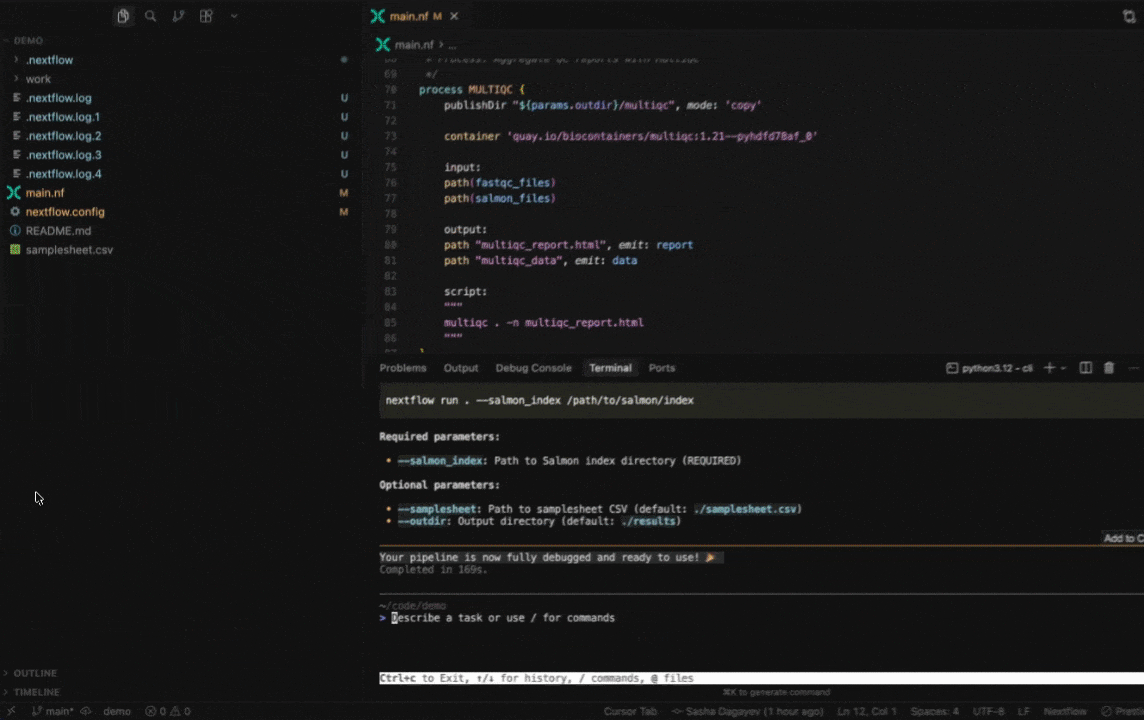
Seqera AI helps you develop, debug, and understand Nextflow pipelines with AI-powered analysis and code generation.
|
|
48
44
|
|
|
49
|
-
|
|
50
|
-
|
|
51
45
|
<details open>
|
|
52
46
|
<summary><strong>Working with Nextflow</strong></summary>
|
|
53
47
|
|
|
@@ -64,29 +58,29 @@ Seqera AI helps you develop, debug, and understand Nextflow pipelines with AI-po
|
|
|
64
58
|
**Generate a `nextflow.config` file**:
|
|
65
59
|
|
|
66
60
|
```
|
|
67
|
-
>
|
|
61
|
+
> Generate a nextflow.config for my pipeline
|
|
68
62
|
```
|
|
69
63
|
|
|
70
64
|
**Debug your pipeline**:
|
|
71
65
|
|
|
72
66
|
```
|
|
73
|
-
>
|
|
67
|
+
> Why is my pipeline failing?
|
|
74
68
|
```
|
|
75
69
|
|
|
76
70
|
```
|
|
77
|
-
>
|
|
71
|
+
> Run nextflow lint on my pipeline
|
|
78
72
|
```
|
|
79
73
|
|
|
80
74
|
**Generate a schema (`nextflow_schema.json`) file**:
|
|
81
75
|
|
|
82
76
|
```
|
|
83
|
-
>
|
|
77
|
+
> Generate a nextflow_schema.json for my pipeline
|
|
84
78
|
```
|
|
85
79
|
|
|
86
80
|
**Convert scripts to Nextflow**:
|
|
87
81
|
|
|
88
82
|
```
|
|
89
|
-
>
|
|
83
|
+
> Convert my Python script to a Nextflow process
|
|
90
84
|
```
|
|
91
85
|
|
|
92
86
|
</details>
|
|
@@ -95,8 +89,6 @@ Seqera AI helps you develop, debug, and understand Nextflow pipelines with AI-po
|
|
|
95
89
|
|
|
96
90
|
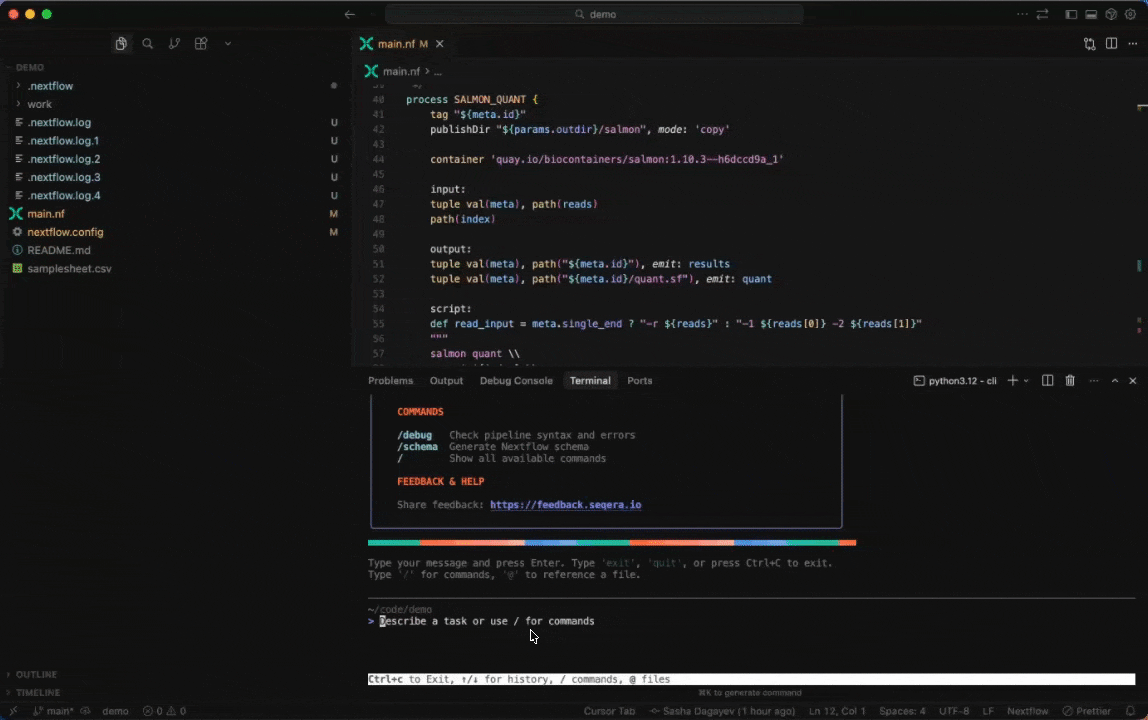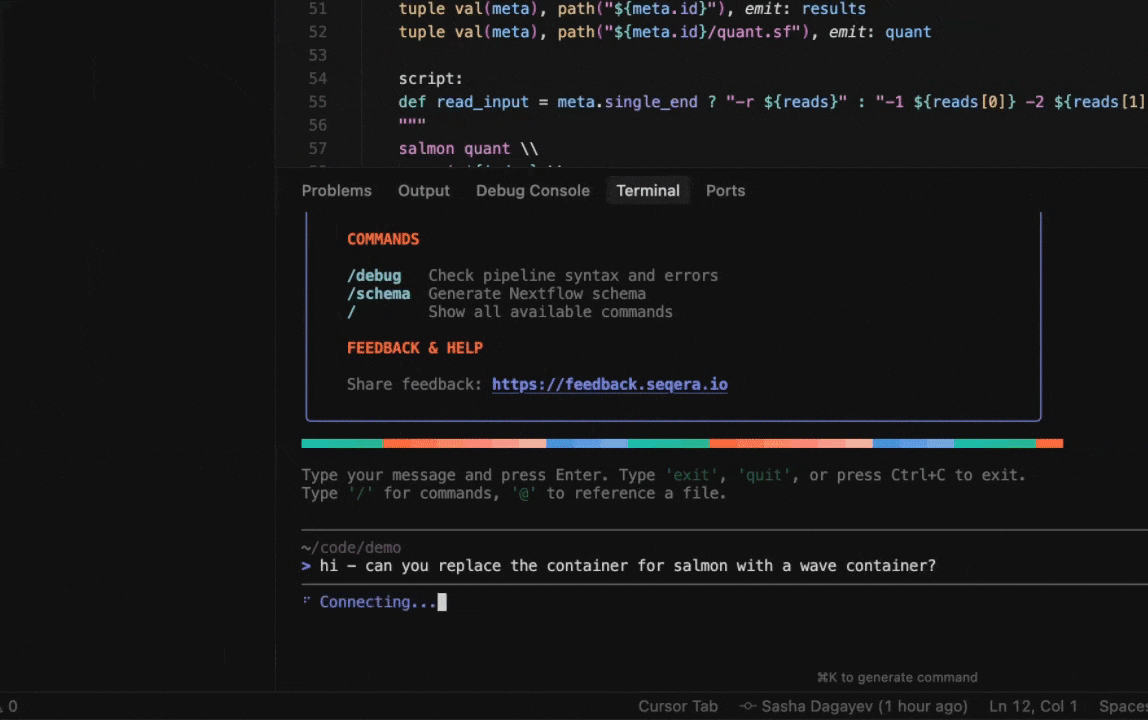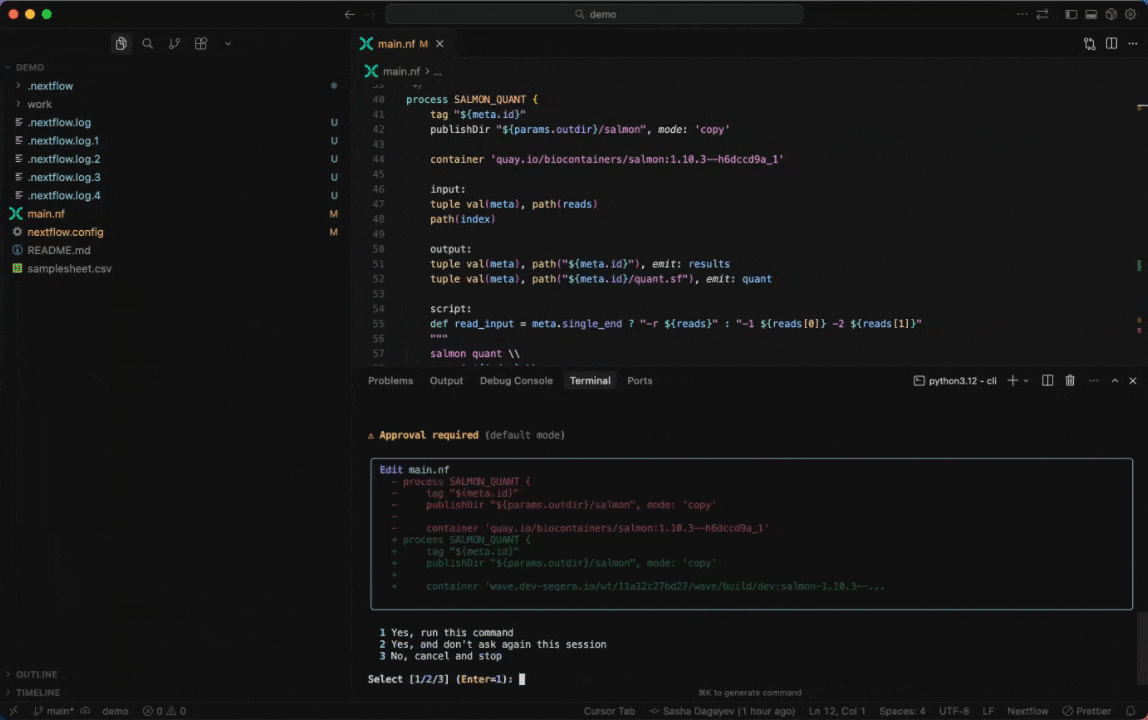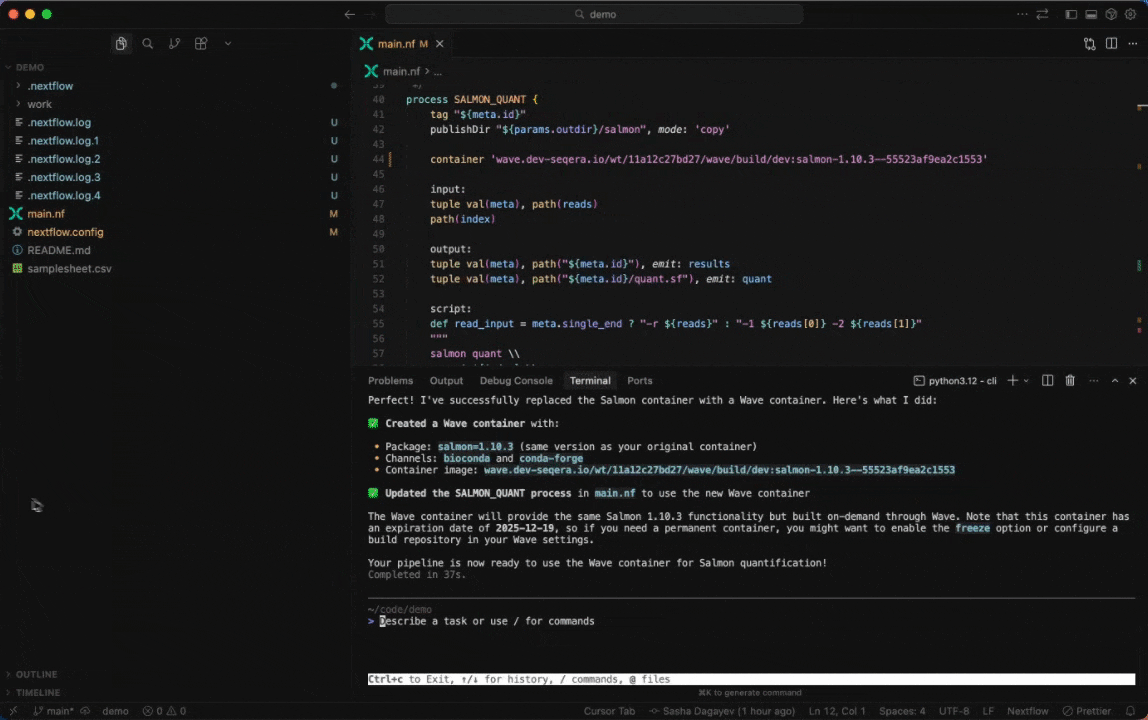
Seqera AI can create containerized environments using Wave, without requiring you to write Dockerfiles.
|
|
97
91
|
|
|
98
|
-
|
|
99
|
-
|
|
100
92
|
<details open>
|
|
101
93
|
<summary><strong>Building containers with Wave</strong></summary>
|
|
102
94
|
|
|
@@ -152,7 +144,7 @@ End your Seqera AI session when done.
|
|
|
152
144
|
|
|
153
145
|
**To end your session**:
|
|
154
146
|
|
|
155
|
-
- Type
|
|
147
|
+
- Type `/exit` or `/quit`
|
|
156
148
|
- Press `Ctrl+C`
|
|
157
149
|
|
|
158
150
|
> **Note:** Your conversation history is preserved for the session but not stored permanently.
|
|
@@ -170,17 +162,22 @@ Seqera AI includes built-in slash commands for common workflows.
|
|
|
170
162
|
|
|
171
163
|
| Command | Description |
|
|
172
164
|
|---------|-------------|
|
|
173
|
-
| `/
|
|
174
|
-
| `/
|
|
175
|
-
| `/
|
|
176
|
-
| `/
|
|
177
|
-
| `/
|
|
178
|
-
| `/
|
|
179
|
-
| `/
|
|
180
|
-
| `/
|
|
181
|
-
| `/
|
|
182
|
-
| `/
|
|
183
|
-
| `/
|
|
165
|
+
| `/help` (or `?`) | Show available commands |
|
|
166
|
+
| `/exit` (`/quit`, `/q`) | Exit the application |
|
|
167
|
+
| `/clear` | Clear conversation history |
|
|
168
|
+
| `/thinking` | Toggle thinking display |
|
|
169
|
+
| `/scroll` | Toggle auto-scroll |
|
|
170
|
+
| `/org` | Show current organization |
|
|
171
|
+
| `/session` | Show current session ID |
|
|
172
|
+
| `/sessions` | Browse and switch sessions |
|
|
173
|
+
| `/lsp` | Show LSP server status |
|
|
174
|
+
| `/status` | Show system status |
|
|
175
|
+
| `/credits` | Show credit balance and usage |
|
|
176
|
+
| `/approval [mode]` | Show or set approval mode (`always`/`default`/`suggest`) |
|
|
177
|
+
| `/update` | Check CLI updates and next steps |
|
|
178
|
+
| `/feedback` | Open feedback form |
|
|
179
|
+
| `/help-community` | Open community help |
|
|
180
|
+
| `/stickers` | Get Seqera stickers |
|
|
184
181
|
|
|
185
182
|
</details>
|
|
186
183
|
|
|
@@ -288,8 +285,6 @@ Seqera AI provides access to over 1,000 nf-core modules for common bioinformatic
|
|
|
288
285
|
|
|
289
286
|
Use Seqera Platform capabilities to run and manage workflows at scale with AI assistance.
|
|
290
287
|
|
|
291
|
-
|
|
292
|
-
|
|
293
288
|
<details open>
|
|
294
289
|
<summary><strong>Working with Seqera Platform</strong></summary>
|
|
295
290
|
|
package/package.json
CHANGED
|
@@ -1,6 +1,6 @@
|
|
|
1
1
|
{
|
|
2
2
|
"name": "seqera",
|
|
3
|
-
"version": "0.
|
|
3
|
+
"version": "0.7.0-dev.2f7b343",
|
|
4
4
|
"description": "AI-powered CLI for Seqera Platform - bioinformatics workflow management",
|
|
5
5
|
"keywords": [
|
|
6
6
|
"seqera",
|
|
@@ -20,9 +20,9 @@
|
|
|
20
20
|
"README.md"
|
|
21
21
|
],
|
|
22
22
|
"optionalDependencies": {
|
|
23
|
-
"@seqera/cli-darwin-arm64": "0.
|
|
24
|
-
"@seqera/cli-darwin-x64": "0.
|
|
25
|
-
"@seqera/cli-linux-x64": "0.
|
|
23
|
+
"@seqera/cli-darwin-arm64": "0.7.0-dev.2f7b343",
|
|
24
|
+
"@seqera/cli-darwin-x64": "0.7.0-dev.2f7b343",
|
|
25
|
+
"@seqera/cli-linux-x64": "0.7.0-dev.2f7b343"
|
|
26
26
|
},
|
|
27
27
|
"engines": {
|
|
28
28
|
"node": ">=18"
|