seqera 0.4.0 → 0.5.0
This diff represents the content of publicly available package versions that have been released to one of the supported registries. The information contained in this diff is provided for informational purposes only and reflects changes between package versions as they appear in their respective public registries.
- package/README.md +326 -0
- package/package.json +7 -6
package/README.md
ADDED
|
@@ -0,0 +1,326 @@
|
|
|
1
|
+
# Seqera AI CLI
|
|
2
|
+
|
|
3
|
+
AI-powered assistant for bioinformatics workflows and [Seqera Platform](https://seqera.io).
|
|
4
|
+
|
|
5
|
+
> **Beta:** Seqera AI CLI is currently in beta. Features and commands may change as we continue to improve the product.
|
|
6
|
+
|
|
7
|
+
> **Note:** Seqera Cloud users receive $20 in free credits to get started with Seqera AI. [Contact us](https://seqera.io/platform/seqera-ai/request-credits/) for additional credits.
|
|
8
|
+
|
|
9
|
+
## Get started
|
|
10
|
+
|
|
11
|
+
1. Install the Seqera AI CLI:
|
|
12
|
+
|
|
13
|
+
```bash
|
|
14
|
+
pip install seqera
|
|
15
|
+
```
|
|
16
|
+
|
|
17
|
+
See [Installation](https://docs.seqera.io/platform-cloud/seqera-ai/installation) for a comprehensive installation guide.
|
|
18
|
+
|
|
19
|
+
2. Log in to your Seqera account:
|
|
20
|
+
|
|
21
|
+
```bash
|
|
22
|
+
seqera login
|
|
23
|
+
```
|
|
24
|
+
|
|
25
|
+
See [Authentication](https://docs.seqera.io/platform-cloud/seqera-ai/authentication) for a comprehensive authentication guide.
|
|
26
|
+
|
|
27
|
+
3. Start Seqera AI:
|
|
28
|
+
|
|
29
|
+
```bash
|
|
30
|
+
seqera ai
|
|
31
|
+
```
|
|
32
|
+
|
|
33
|
+
4. Run your first prompt:
|
|
34
|
+
|
|
35
|
+
```
|
|
36
|
+
/debug
|
|
37
|
+
```
|
|
38
|
+
|
|
39
|
+
See [Use cases](https://docs.seqera.io/platform-cloud/seqera-ai/use-cases) for a comprehensive list of use cases.
|
|
40
|
+
|
|
41
|
+
## Use cases
|
|
42
|
+
|
|
43
|
+
Seqera AI is an intelligent command-line assistant that helps you build, run, and manage bioinformatics workflows. The following sections describe several common use cases.
|
|
44
|
+
|
|
45
|
+
### Work with Nextflow
|
|
46
|
+
|
|
47
|
+
Seqera AI helps you develop, debug, and understand Nextflow pipelines with AI-powered analysis and code generation.
|
|
48
|
+
|
|
49
|
+
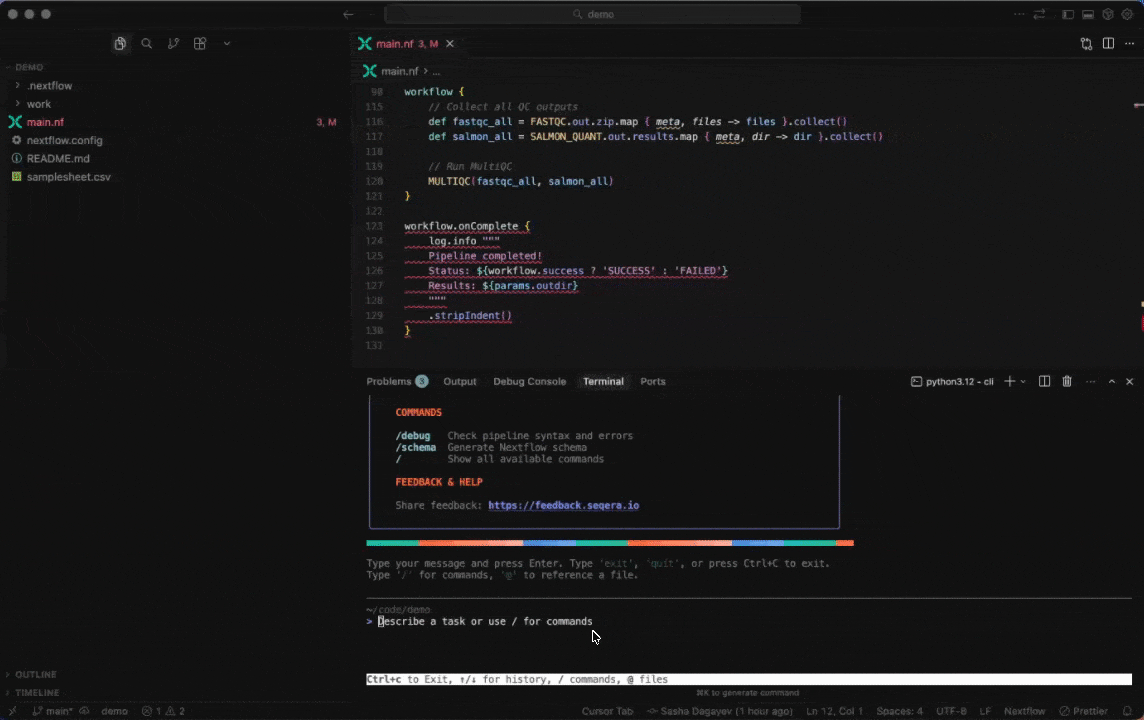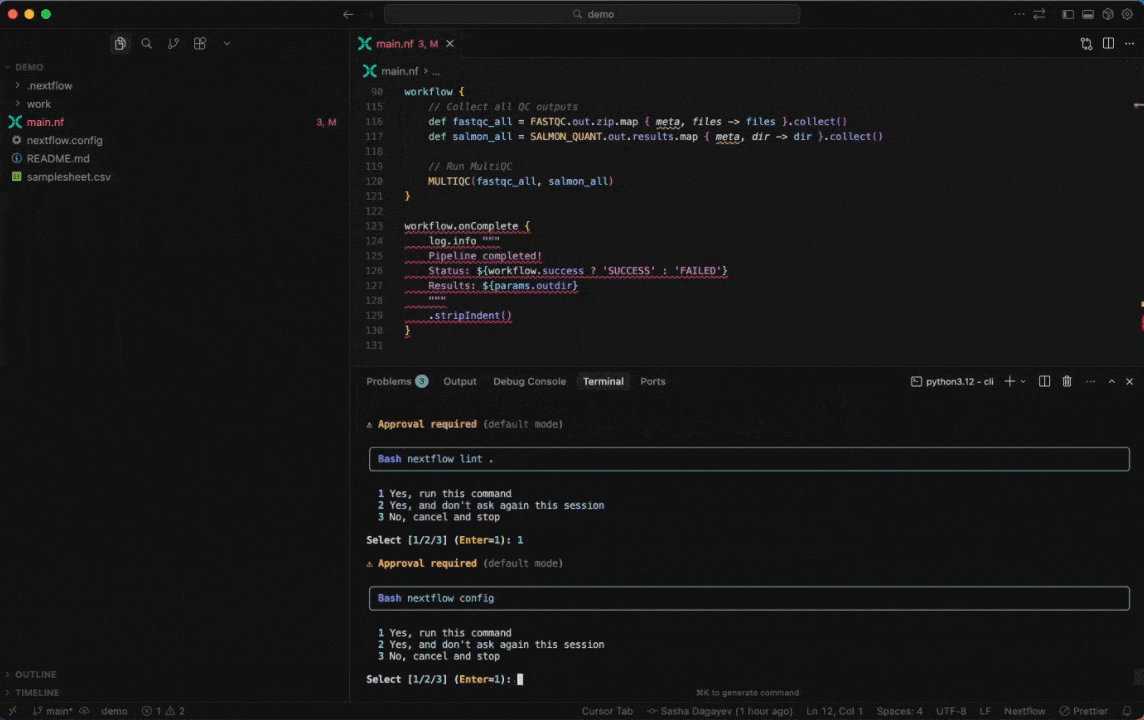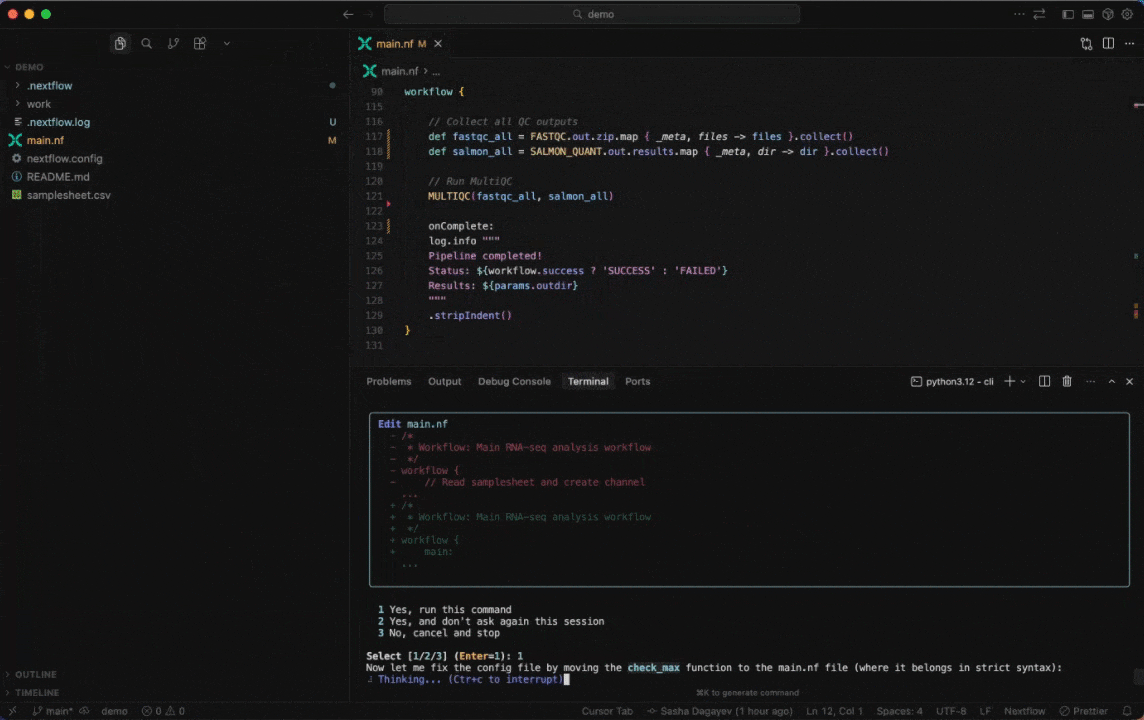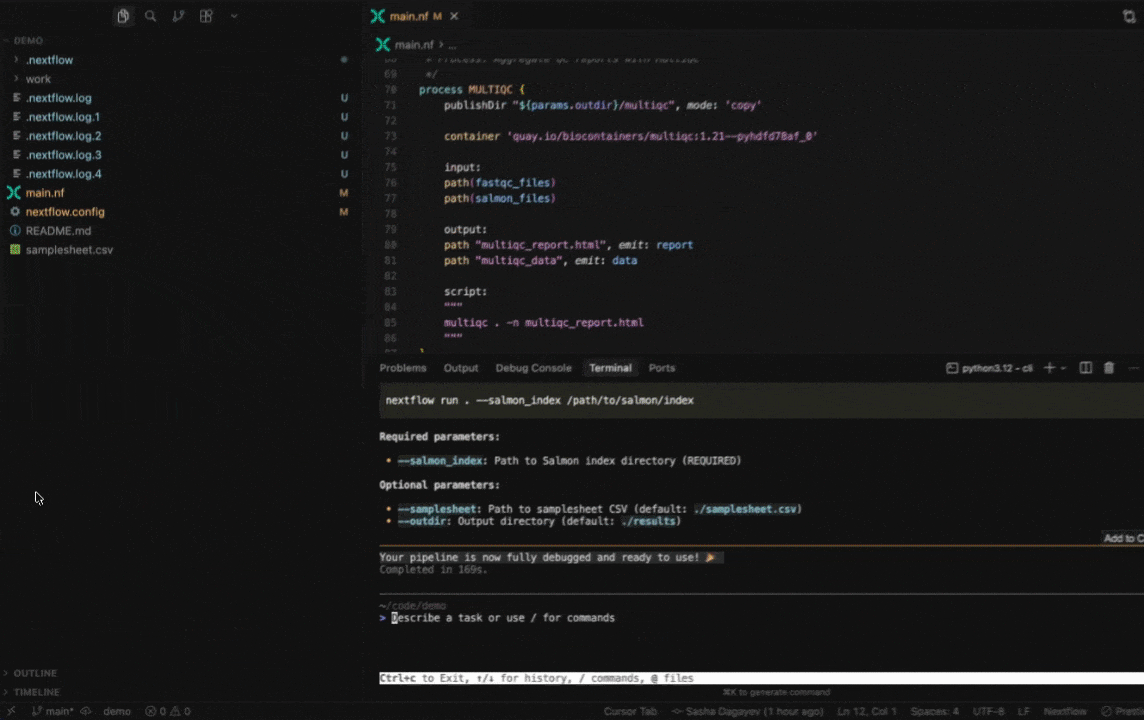
|
|
50
|
+
|
|
51
|
+
<details open>
|
|
52
|
+
<summary><strong>Working with Nextflow</strong></summary>
|
|
53
|
+
|
|
54
|
+
**Understand your pipeline structure**:
|
|
55
|
+
|
|
56
|
+
```
|
|
57
|
+
> Show me the structure of main.nf
|
|
58
|
+
```
|
|
59
|
+
|
|
60
|
+
```
|
|
61
|
+
> What processes are defined in this pipeline?
|
|
62
|
+
```
|
|
63
|
+
|
|
64
|
+
**Generate a `nextflow.config` file**:
|
|
65
|
+
|
|
66
|
+
```
|
|
67
|
+
> /config
|
|
68
|
+
```
|
|
69
|
+
|
|
70
|
+
**Debug your pipeline**:
|
|
71
|
+
|
|
72
|
+
```
|
|
73
|
+
> /debug
|
|
74
|
+
```
|
|
75
|
+
|
|
76
|
+
```
|
|
77
|
+
> Why is my pipeline failing?
|
|
78
|
+
```
|
|
79
|
+
|
|
80
|
+
**Generate a schema (`nextflow_schema.json`) file**:
|
|
81
|
+
|
|
82
|
+
```
|
|
83
|
+
> /schema
|
|
84
|
+
```
|
|
85
|
+
|
|
86
|
+
**Convert scripts to Nextflow**:
|
|
87
|
+
|
|
88
|
+
```
|
|
89
|
+
> /convert-python-script
|
|
90
|
+
```
|
|
91
|
+
|
|
92
|
+
</details>
|
|
93
|
+
|
|
94
|
+
### Build containers with Wave
|
|
95
|
+
|
|
96
|
+
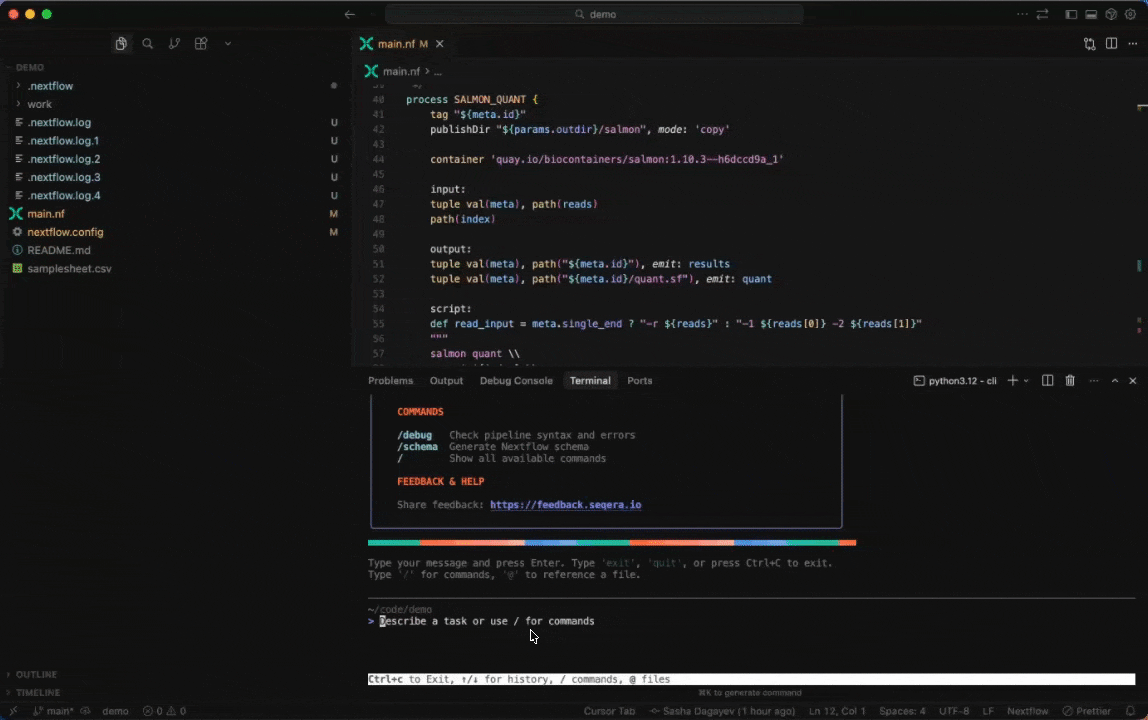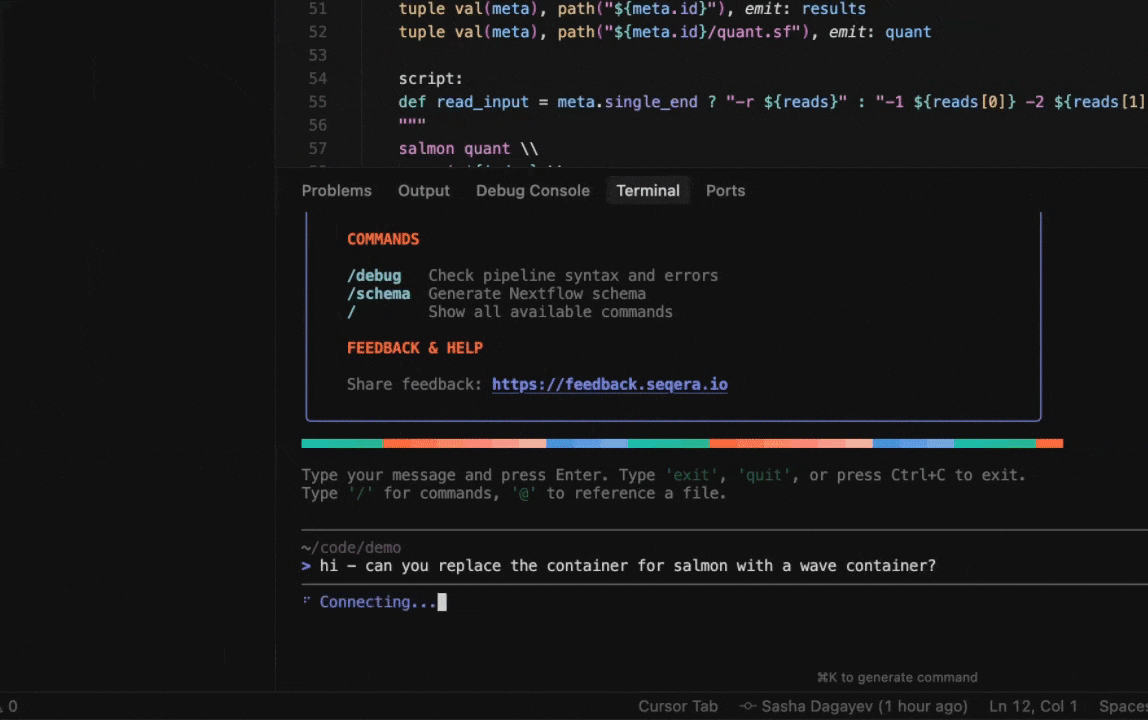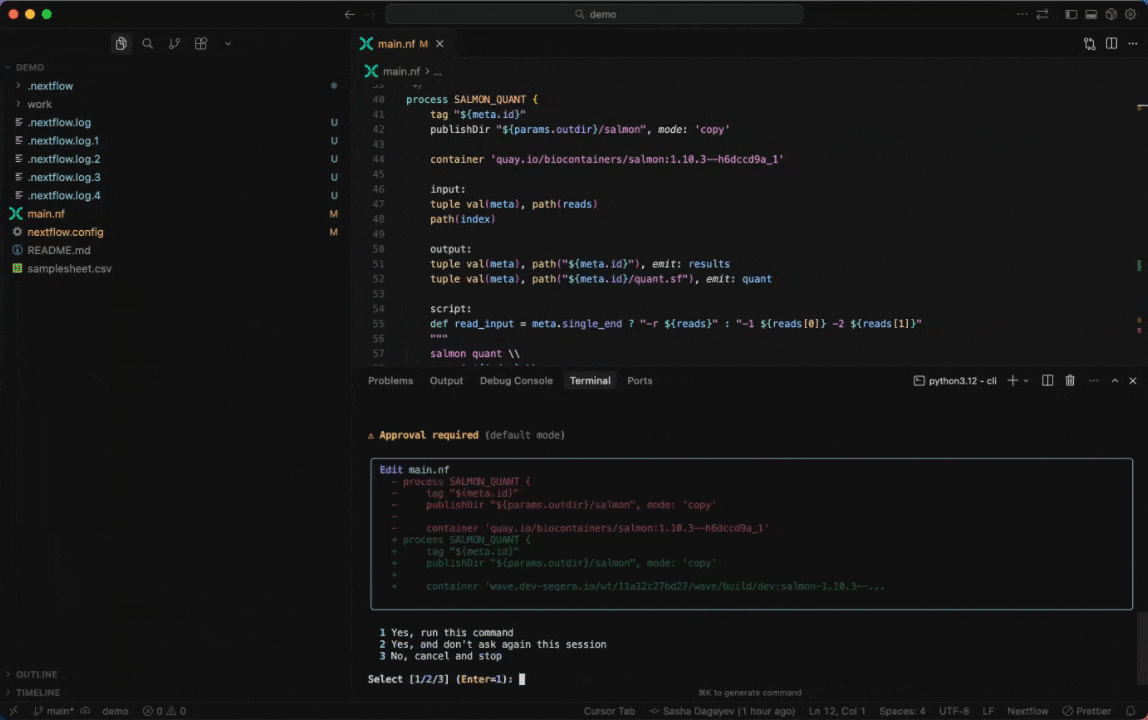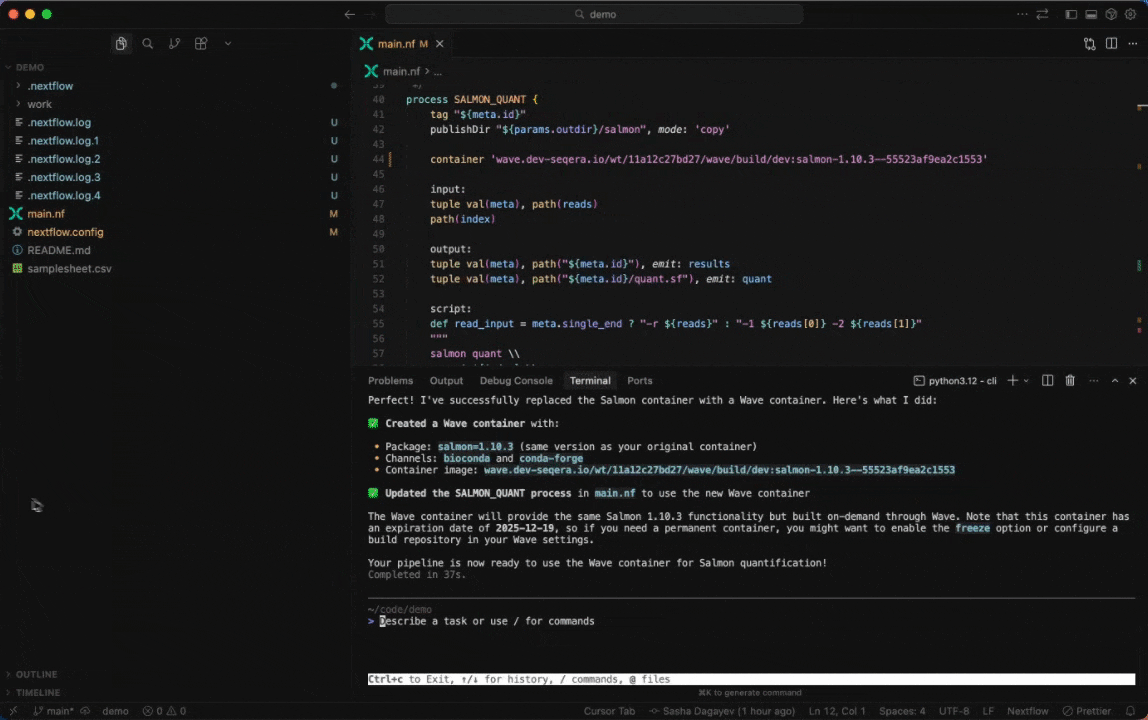
Seqera AI can create containerized environments using Wave, without requiring you to write Dockerfiles.
|
|
97
|
+
|
|
98
|
+
|
|
99
|
+
|
|
100
|
+
<details open>
|
|
101
|
+
<summary><strong>Building containers with Wave</strong></summary>
|
|
102
|
+
|
|
103
|
+
**Create a container with conda packages**:
|
|
104
|
+
|
|
105
|
+
```
|
|
106
|
+
> Create a container with samtools and bwa from bioconda
|
|
107
|
+
```
|
|
108
|
+
|
|
109
|
+
**Create a container with pip packages**:
|
|
110
|
+
|
|
111
|
+
```
|
|
112
|
+
> Build a container with pandas, numpy, and scikit-learn
|
|
113
|
+
```
|
|
114
|
+
|
|
115
|
+
**Get a container for a specific tool**:
|
|
116
|
+
|
|
117
|
+
```
|
|
118
|
+
> I need a container with FastQC version 0.12.1
|
|
119
|
+
```
|
|
120
|
+
|
|
121
|
+
> **Note:** The assistant will generate a Wave container URL that you can use directly in your Nextflow pipelines or pull with Docker.
|
|
122
|
+
|
|
123
|
+
</details>
|
|
124
|
+
|
|
125
|
+
### Customize your session
|
|
126
|
+
|
|
127
|
+
Customize your session with command-line options.
|
|
128
|
+
|
|
129
|
+
<details open>
|
|
130
|
+
<summary><strong>Customize your session</strong></summary>
|
|
131
|
+
|
|
132
|
+
**Start in a specific directory**:
|
|
133
|
+
|
|
134
|
+
```bash
|
|
135
|
+
seqera ai -w /path/to/project
|
|
136
|
+
```
|
|
137
|
+
|
|
138
|
+
**Set approval mode for local commands**:
|
|
139
|
+
|
|
140
|
+
```bash
|
|
141
|
+
seqera ai -a full
|
|
142
|
+
```
|
|
143
|
+
|
|
144
|
+
</details>
|
|
145
|
+
|
|
146
|
+
### Exit the assistant
|
|
147
|
+
|
|
148
|
+
End your Seqera AI session when done.
|
|
149
|
+
|
|
150
|
+
<details open>
|
|
151
|
+
<summary><strong>Exit the assistant</strong></summary>
|
|
152
|
+
|
|
153
|
+
**To end your session**:
|
|
154
|
+
|
|
155
|
+
- Type `exit` or `quit`
|
|
156
|
+
- Press `Ctrl+C`
|
|
157
|
+
|
|
158
|
+
> **Note:** Your conversation history is preserved for the session but not stored permanently.
|
|
159
|
+
|
|
160
|
+
</details>
|
|
161
|
+
|
|
162
|
+
### Use slash commands
|
|
163
|
+
|
|
164
|
+
Seqera AI includes built-in slash commands for common workflows.
|
|
165
|
+
|
|
166
|
+
<details open>
|
|
167
|
+
<summary><strong>Use slash commands</strong></summary>
|
|
168
|
+
|
|
169
|
+
**Type `/` to see all available commands**:
|
|
170
|
+
|
|
171
|
+
| Command | Description |
|
|
172
|
+
|---------|-------------|
|
|
173
|
+
| `/config` | Generate a nextflow.config file |
|
|
174
|
+
| `/schema` | Generate a Nextflow schema |
|
|
175
|
+
| `/debug` | Run nextflow lint and preview |
|
|
176
|
+
| `/debug-last-run` | Debug the last local run |
|
|
177
|
+
| `/debug-last-run-on-seqera` | Debug the last Platform run |
|
|
178
|
+
| `/migrate-from-wdl` | Convert WDL to Nextflow |
|
|
179
|
+
| `/migrate-from-snakemake` | Convert Snakemake to Nextflow |
|
|
180
|
+
| `/convert-python-script` | Convert Python script to Nextflow |
|
|
181
|
+
| `/convert-r-script` | Convert R script to Nextflow |
|
|
182
|
+
| `/convert-jupyter-notebook` | Convert Jupyter notebook to Nextflow |
|
|
183
|
+
| `/write-nf-test` | Write nf-tests for your pipeline |
|
|
184
|
+
|
|
185
|
+
</details>
|
|
186
|
+
|
|
187
|
+
### Work with data
|
|
188
|
+
|
|
189
|
+
Seqera AI helps you manage data through Platform data links and access reference datasets.
|
|
190
|
+
|
|
191
|
+
<details open>
|
|
192
|
+
<summary><strong>Working with data</strong></summary>
|
|
193
|
+
|
|
194
|
+
**Browse data links**:
|
|
195
|
+
|
|
196
|
+
```
|
|
197
|
+
> List my data links
|
|
198
|
+
```
|
|
199
|
+
|
|
200
|
+
```
|
|
201
|
+
> Show me the contents of my S3 data link
|
|
202
|
+
```
|
|
203
|
+
|
|
204
|
+
**Download and upload files**:
|
|
205
|
+
|
|
206
|
+
```
|
|
207
|
+
> Generate a download URL for results/final_report.html
|
|
208
|
+
```
|
|
209
|
+
|
|
210
|
+
```
|
|
211
|
+
> Upload my local results to the data link
|
|
212
|
+
```
|
|
213
|
+
|
|
214
|
+
**Access reference data**:
|
|
215
|
+
|
|
216
|
+
```
|
|
217
|
+
> Find the human reference genome GRCh38
|
|
218
|
+
```
|
|
219
|
+
|
|
220
|
+
```
|
|
221
|
+
> Search for RNA-Seq test data
|
|
222
|
+
```
|
|
223
|
+
|
|
224
|
+
</details>
|
|
225
|
+
|
|
226
|
+
### Work with local files
|
|
227
|
+
|
|
228
|
+
Seqera AI can interact with files in your current working directory.
|
|
229
|
+
|
|
230
|
+
<details open>
|
|
231
|
+
<summary><strong>Work with local files</strong></summary>
|
|
232
|
+
|
|
233
|
+
**Start the assistant from your project folder**:
|
|
234
|
+
|
|
235
|
+
```bash
|
|
236
|
+
cd /path/to/your/project
|
|
237
|
+
seqera ai
|
|
238
|
+
```
|
|
239
|
+
|
|
240
|
+
**Then, ask the assistant to help with local tasks**:
|
|
241
|
+
|
|
242
|
+
```
|
|
243
|
+
> Show me the structure of main.nf
|
|
244
|
+
```
|
|
245
|
+
|
|
246
|
+
```
|
|
247
|
+
> Add a new process to handle quality control
|
|
248
|
+
```
|
|
249
|
+
|
|
250
|
+
> **Note:** Local file operations are controlled by [approval modes](https://docs.seqera.io/platform-cloud/seqera-ai/command-approval). By default, the assistant will ask for your approval before making changes outside your working directory or running potentially dangerous commands.
|
|
251
|
+
|
|
252
|
+
</details>
|
|
253
|
+
|
|
254
|
+
### Work with nf-core modules
|
|
255
|
+
|
|
256
|
+
Seqera AI provides access to over 1,000 nf-core modules for common bioinformatics tasks.
|
|
257
|
+
|
|
258
|
+
<details open>
|
|
259
|
+
<summary><strong>Working with nf-core modules</strong></summary>
|
|
260
|
+
|
|
261
|
+
**Search for modules**:
|
|
262
|
+
|
|
263
|
+
```
|
|
264
|
+
> Find nf-core modules for sequence alignment
|
|
265
|
+
```
|
|
266
|
+
|
|
267
|
+
```
|
|
268
|
+
> What modules are available for variant calling?
|
|
269
|
+
```
|
|
270
|
+
|
|
271
|
+
**Get module details**:
|
|
272
|
+
|
|
273
|
+
```
|
|
274
|
+
> Show me how to use the nf-core/bwa/mem module
|
|
275
|
+
```
|
|
276
|
+
|
|
277
|
+
**Run a module**:
|
|
278
|
+
|
|
279
|
+
```
|
|
280
|
+
> Run FastQC on my FASTQ files
|
|
281
|
+
```
|
|
282
|
+
|
|
283
|
+
> **Note:** The assistant can generate the exact Nextflow command with proper parameters for your data.
|
|
284
|
+
|
|
285
|
+
</details>
|
|
286
|
+
|
|
287
|
+
### Work with Seqera Platform
|
|
288
|
+
|
|
289
|
+
Use Seqera Platform capabilities to run and manage workflows at scale with AI assistance.
|
|
290
|
+
|
|
291
|
+
|
|
292
|
+
|
|
293
|
+
<details open>
|
|
294
|
+
<summary><strong>Working with Seqera Platform</strong></summary>
|
|
295
|
+
|
|
296
|
+
**List your workflows**:
|
|
297
|
+
|
|
298
|
+
```
|
|
299
|
+
> List my recent workflows
|
|
300
|
+
```
|
|
301
|
+
|
|
302
|
+
**Launch a pipeline**:
|
|
303
|
+
|
|
304
|
+
```
|
|
305
|
+
> Launch the nf-core/rnaseq pipeline with the test profile
|
|
306
|
+
```
|
|
307
|
+
|
|
308
|
+
**Debug failed runs**:
|
|
309
|
+
|
|
310
|
+
```
|
|
311
|
+
> Why did my last workflow fail?
|
|
312
|
+
```
|
|
313
|
+
|
|
314
|
+
```
|
|
315
|
+
> Get the logs for the failed task in my last run
|
|
316
|
+
```
|
|
317
|
+
|
|
318
|
+
</details>
|
|
319
|
+
|
|
320
|
+
## Learn more
|
|
321
|
+
|
|
322
|
+
- [Installation](https://docs.seqera.io/platform-cloud/seqera-ai/installation): Detailed installation instructions
|
|
323
|
+
- [Authentication](https://docs.seqera.io/platform-cloud/seqera-ai/authentication): Log in, log out, and session management
|
|
324
|
+
- [Command approval](https://docs.seqera.io/platform-cloud/seqera-ai/command-approval): Control which commands run automatically
|
|
325
|
+
- [Use cases](https://docs.seqera.io/platform-cloud/seqera-ai/use-cases): Seqera AI CLI use cases
|
|
326
|
+
- [Troubleshooting](https://docs.seqera.io/platform-cloud/troubleshooting_and_faqs/seqera-ai): Troubleshoot common errors
|
package/package.json
CHANGED
|
@@ -1,6 +1,6 @@
|
|
|
1
1
|
{
|
|
2
2
|
"name": "seqera",
|
|
3
|
-
"version": "0.
|
|
3
|
+
"version": "0.5.0",
|
|
4
4
|
"description": "AI-powered CLI for Seqera Platform - bioinformatics workflow management",
|
|
5
5
|
"keywords": [
|
|
6
6
|
"seqera",
|
|
@@ -11,17 +11,18 @@
|
|
|
11
11
|
"cli"
|
|
12
12
|
],
|
|
13
13
|
"author": "Seqera Labs",
|
|
14
|
-
"license": "
|
|
14
|
+
"license": "Custom",
|
|
15
15
|
"bin": {
|
|
16
16
|
"seqera": "bin/seqera"
|
|
17
17
|
},
|
|
18
18
|
"files": [
|
|
19
|
-
"bin/**"
|
|
19
|
+
"bin/**",
|
|
20
|
+
"README.md"
|
|
20
21
|
],
|
|
21
22
|
"optionalDependencies": {
|
|
22
|
-
"@seqera/cli-darwin-arm64": "0.
|
|
23
|
-
"@seqera/cli-darwin-x64": "0.
|
|
24
|
-
"@seqera/cli-linux-x64": "0.
|
|
23
|
+
"@seqera/cli-darwin-arm64": "0.5.0",
|
|
24
|
+
"@seqera/cli-darwin-x64": "0.5.0",
|
|
25
|
+
"@seqera/cli-linux-x64": "0.5.0"
|
|
25
26
|
},
|
|
26
27
|
"engines": {
|
|
27
28
|
"node": ">=18"
|