@lsbjordao/type-taxon-script 1.1.2 → 1.1.3
This diff represents the content of publicly available package versions that have been released to one of the supported registries. The information contained in this diff is provided for informational purposes only and reflects changes between package versions as they appear in their respective public registries.
- package/package.json +1 -1
- package/readme.md +3 -9
package/package.json
CHANGED
|
@@ -1,6 +1,6 @@
|
|
|
1
1
|
{
|
|
2
2
|
"name": "@lsbjordao/type-taxon-script",
|
|
3
|
-
"version": "1.1.
|
|
3
|
+
"version": "1.1.3",
|
|
4
4
|
"author": "Lucas Jordão <tucarj@gmail.com> & André Eppinghaus <andreeppinghaus@gmail.com> & Vicente Calfo <vicentecalfo@gmail.com>",
|
|
5
5
|
"license": "ISC",
|
|
6
6
|
"description": "TypeTaxonScript",
|
package/readme.md
CHANGED
|
@@ -16,7 +16,7 @@ Visual Studio Code (VS Code) is a versatile code editor that provides a user-fri
|
|
|
16
16
|
|
|
17
17
|
## Clone the repository from GitHub in VS Code
|
|
18
18
|
|
|
19
|
-
To clone the *Mimosa* project repository for TTS from GitHub, follow these steps:
|
|
19
|
+
To clone the *Mimosa* project repository for TTS from GitHub (https://github.com/lsbjordao/TTS-Mimosa), follow these steps:
|
|
20
20
|
|
|
21
21
|
1. In VS Code, access the Command Palette by pressing `Ctrl + Shift + P` (Windows/Linux) or `Cmd + Shift + P` (macOS).
|
|
22
22
|
2. Type `Git: Clone` and select the option that appears.
|
|
@@ -93,14 +93,6 @@ tts new --genus Mimosa --species epithet
|
|
|
93
93
|
|
|
94
94
|
After the process, a new file named `Mimosa_epithet.ts` will be created in the `./output` directory. To access this script file, simply hold down the `Ctrl` key and click on the file path displayed in the console. However, before you begin editing the script, it is important to relocate this file to the `./taxon` directory, as the script specifically functions within that directory. Outside this directory, the script will not works properly. Opening the script outside of this directory will trigger multiple dependency errors.
|
|
95
95
|
|
|
96
|
-
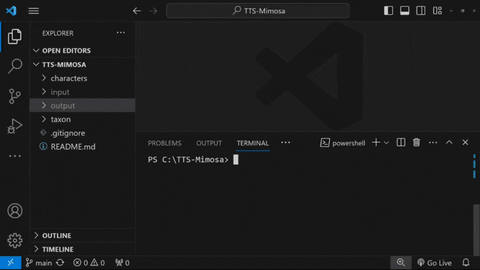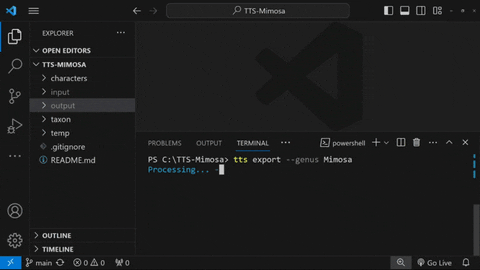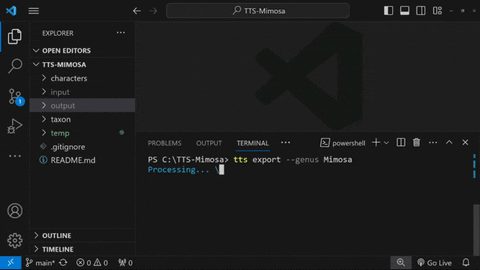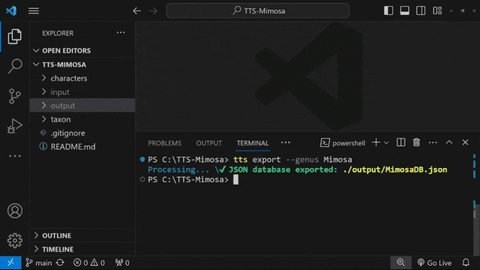
## Exporting JSON database
|
|
97
|
-
|
|
98
|
-
```bash
|
|
99
|
-
tts export --genus Mimosa
|
|
100
|
-
```
|
|
101
|
-
|
|
102
|
-
|
|
103
|
-
|
|
104
96
|
## Importing from `.csv` file
|
|
105
97
|
|
|
106
98
|
It is also possible to import data of multiple taxa from a `.csv` file with a header in the following manner:
|
|
@@ -163,6 +155,8 @@ To export all taxa inside `./taxon/Mimosa`, type:
|
|
|
163
155
|
tts export --genus Mimosa
|
|
164
156
|
```
|
|
165
157
|
|
|
158
|
+

|
|
159
|
+
|
|
166
160
|
If you intend to generate a database containing a specific list of taxa from the directory `./taxon/Mimosa`, edit the `./input/taxonToImport.csv` file accordingly. After making the necessary edits, execute the following command:
|
|
167
161
|
|
|
168
162
|
```bash
|