sude 0.1.0__tar.gz
This diff represents the content of publicly available package versions that have been released to one of the supported registries. The information contained in this diff is provided for informational purposes only and reflects changes between package versions as they appear in their respective public registries.
- sude-0.1.0/.gitignore +11 -0
- sude-0.1.0/.python-version +1 -0
- sude-0.1.0/PKG-INFO +128 -0
- sude-0.1.0/README.md +105 -0
- sude-0.1.0/benchmarks/breast.csv +683 -0
- sude-0.1.0/benchmarks/dermat.csv +358 -0
- sude-0.1.0/benchmarks/drybean.csv +13611 -0
- sude-0.1.0/benchmarks/mfeat.csv +2000 -0
- sude-0.1.0/benchmarks/rice.csv +3810 -0
- sude-0.1.0/benchmarks/shuttle.csv +14500 -0
- sude-0.1.0/benchmarks/spambase.csv +4601 -0
- sude-0.1.0/benchmarks/wine.csv +178 -0
- sude-0.1.0/image/sude.jpg +0 -0
- sude-0.1.0/pyproject.toml +41 -0
- sude-0.1.0/sude/__init__.py +109 -0
- sude-0.1.0/sude/clle.py +29 -0
- sude-0.1.0/sude/init_pca.py +33 -0
- sude-0.1.0/sude/learning_l.py +128 -0
- sude-0.1.0/sude/learning_s.py +107 -0
- sude-0.1.0/sude/mds.py +26 -0
- sude-0.1.0/sude/opt_scale.py +25 -0
- sude-0.1.0/sude/pca.py +20 -0
- sude-0.1.0/sude/plotcluster2.py +39 -0
- sude-0.1.0/sude/pps.py +27 -0
- sude-0.1.0/sude/scml.py +107 -0
- sude-0.1.0/uv.lock +659 -0
sude-0.1.0/.gitignore
ADDED
|
@@ -0,0 +1 @@
|
|
|
1
|
+
3.8
|
sude-0.1.0/PKG-INFO
ADDED
|
@@ -0,0 +1,128 @@
|
|
|
1
|
+
Metadata-Version: 2.4
|
|
2
|
+
Name: sude
|
|
3
|
+
Version: 0.1.0
|
|
4
|
+
Summary: A scalable manifold learning (SUDE) method that can cope with large-scale and high-dimensional data in an efficient manner.
|
|
5
|
+
Project-URL: Homepage, https://github.com/ZPGuiGroupWhu/SUDE-pkg
|
|
6
|
+
Project-URL: Repository, https://github.com/ZPGuiGroupWhu/SUDE-pkg.git
|
|
7
|
+
Project-URL: Issues, https://github.com/ZPGuiGroupWhu/SUDE-pkg/issues
|
|
8
|
+
Author-email: pdh <pengdh@whu.edu.cn>
|
|
9
|
+
License: MIT
|
|
10
|
+
Keywords: dimension reduction,embedding,landmark sampling,manifold learning
|
|
11
|
+
Classifier: Development Status :: 3 - Alpha
|
|
12
|
+
Classifier: Intended Audience :: Science/Research
|
|
13
|
+
Classifier: License :: OSI Approved :: MIT License
|
|
14
|
+
Classifier: Programming Language :: Python :: 3
|
|
15
|
+
Classifier: Programming Language :: Python :: 3.8
|
|
16
|
+
Classifier: Programming Language :: Python :: 3.9
|
|
17
|
+
Classifier: Programming Language :: Python :: 3.10
|
|
18
|
+
Classifier: Programming Language :: Python :: 3.11
|
|
19
|
+
Classifier: Topic :: Scientific/Engineering :: Artificial Intelligence
|
|
20
|
+
Requires-Python: >=3.8
|
|
21
|
+
Requires-Dist: scikit-learn>=1.3.2
|
|
22
|
+
Description-Content-Type: text/markdown
|
|
23
|
+
|
|
24
|
+
# Sampling-enabled scalable manifold learning unveils the discriminative cluster structure of high-dimensional data (SUDE)
|
|
25
|
+
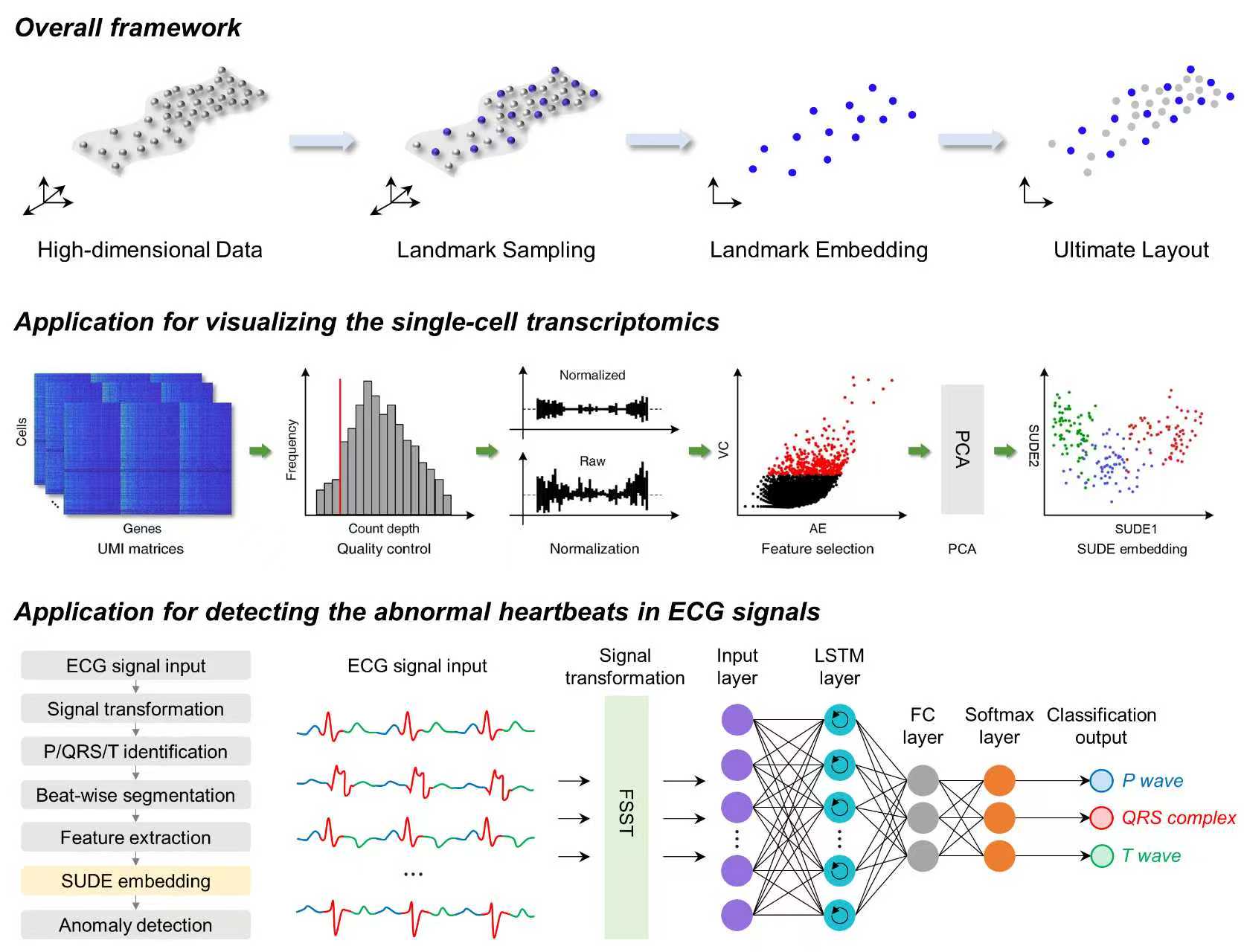
We propose a scalable manifold learning (SUDE) method that can cope with large-scale and high-dimensional data in an efficient manner. It starts by seeking a set of landmarks to construct the low-dimensional skeleton of the entire data, and then incorporates the non-landmarks into this skeleton based on the constrained locally linear embedding. This project provides the ***Python version of SUDE***, and the MATLAB version can be found at https://github.com/ZPGuiGroupWhu/sude. This paper has been published in ***Nature Machine Intelligence***, and more details can be seen https://www.nature.com/articles/s42256-025-01112-9.
|
|
26
|
+
|
|
27
|
+
|
|
28
|
+
|
|
29
|
+
# Installation
|
|
30
|
+
Supported `python` versions are `3.8` and above.
|
|
31
|
+
|
|
32
|
+
This project has been uploaded to [PyPI](https://pypi.org/project/sude/), supporting direct download and installation from pypi
|
|
33
|
+
|
|
34
|
+
```
|
|
35
|
+
pip install sude
|
|
36
|
+
```
|
|
37
|
+
|
|
38
|
+
## Manual Installation
|
|
39
|
+
|
|
40
|
+
```
|
|
41
|
+
git clone https://github.com/ZPGuiGroupWhu/SUDE-pkg.git
|
|
42
|
+
cd SUDE-pkg
|
|
43
|
+
pip install -e .
|
|
44
|
+
```
|
|
45
|
+
|
|
46
|
+
# How To Run
|
|
47
|
+
The SUDE algorithm package provides the `sude` function for clustering.
|
|
48
|
+
|
|
49
|
+
The description of the hyperparameters for user configuration are presented as follows
|
|
50
|
+
```python
|
|
51
|
+
def sude(
|
|
52
|
+
X,
|
|
53
|
+
no_dims = 2,
|
|
54
|
+
k1 = 20,
|
|
55
|
+
normalize = True,
|
|
56
|
+
large = False,
|
|
57
|
+
initialize = 'le',
|
|
58
|
+
agg_coef = 1.2,
|
|
59
|
+
T_epoch = 50,
|
|
60
|
+
):
|
|
61
|
+
"""
|
|
62
|
+
This function returns representation of the N by D matrix X in the lower-dimensional space. Each row in X
|
|
63
|
+
represents an observation.
|
|
64
|
+
|
|
65
|
+
Parameters are:
|
|
66
|
+
|
|
67
|
+
'no_dims' - A positive integer specifying the number of dimension of the representation Y.
|
|
68
|
+
Default: 2
|
|
69
|
+
'k1' - A non-negative integer specifying the number of nearest neighbors for PPS to
|
|
70
|
+
sample landmarks. It must be smaller than N.
|
|
71
|
+
Default: adaptive
|
|
72
|
+
'normalize' - Logical scalar. If true, normalize X using min-max normalization. If features in
|
|
73
|
+
X are on different scales, 'Normalize' should be set to true because the learning
|
|
74
|
+
process is based on nearest neighbors and features with large scales can override
|
|
75
|
+
the contribution of features with small scales.
|
|
76
|
+
Default: True
|
|
77
|
+
'large' - Logical scalar. If true, the data can be split into multiple blocks to avoid the problem
|
|
78
|
+
of memory overflow, and the gradient can be computed block by block using 'learning_l' function.
|
|
79
|
+
Default: False
|
|
80
|
+
'initialize' - A string specifying the method for initializing Y before manifold learning.
|
|
81
|
+
'le' - Laplacian eigenmaps.
|
|
82
|
+
'pca' - Principal component analysis.
|
|
83
|
+
'mds' - Multidimensional scaling.
|
|
84
|
+
Default: 'le'
|
|
85
|
+
'agg_coef' - A positive scalar specifying the aggregation coefficient.
|
|
86
|
+
Default: 1.2
|
|
87
|
+
'T_epoch' - Maximum number of epochs to take.
|
|
88
|
+
Default: 50
|
|
89
|
+
"""
|
|
90
|
+
```
|
|
91
|
+
|
|
92
|
+
After installing the SUDE library, you can use this function as follows:
|
|
93
|
+
```python
|
|
94
|
+
import pandas as pd
|
|
95
|
+
import numpy as np
|
|
96
|
+
from sude import sude
|
|
97
|
+
import time
|
|
98
|
+
import matplotlib.pyplot as plt
|
|
99
|
+
|
|
100
|
+
# Input data
|
|
101
|
+
data = np.array(pd.read_csv('benchmarks/rice.csv', header=None))
|
|
102
|
+
|
|
103
|
+
# Obtain data size and true annotations
|
|
104
|
+
m = data.shape[1]
|
|
105
|
+
X = data[:, :m - 1]
|
|
106
|
+
ref = data[:, m - 1]
|
|
107
|
+
|
|
108
|
+
# Perform SUDE embedding
|
|
109
|
+
start_time = time.time()
|
|
110
|
+
Y = sude(X, k1=10)
|
|
111
|
+
end_time = time.time()
|
|
112
|
+
print("Elapsed time:", end_time - start_time, 's')
|
|
113
|
+
|
|
114
|
+
plt.scatter(Y[:, 0], Y[:, 1], c=ref, cmap='tab10', s=4)
|
|
115
|
+
plt.show()
|
|
116
|
+
```
|
|
117
|
+
|
|
118
|
+
# Citation Request
|
|
119
|
+
Peng, D., Gui, Z., Wei, W. et al. Sampling-enabled scalable manifold learning unveils the discriminative cluster structure of high-dimensional data. Nat. Mach. Intell. (2025). https://doi.org/10.1038/s42256-025-01112-9
|
|
120
|
+
|
|
121
|
+
|
|
122
|
+
# License
|
|
123
|
+
SUDE is released under the MIT License.
|
|
124
|
+
|
|
125
|
+
|
|
126
|
+
|
|
127
|
+
|
|
128
|
+
|
sude-0.1.0/README.md
ADDED
|
@@ -0,0 +1,105 @@
|
|
|
1
|
+
# Sampling-enabled scalable manifold learning unveils the discriminative cluster structure of high-dimensional data (SUDE)
|
|
2
|
+
We propose a scalable manifold learning (SUDE) method that can cope with large-scale and high-dimensional data in an efficient manner. It starts by seeking a set of landmarks to construct the low-dimensional skeleton of the entire data, and then incorporates the non-landmarks into this skeleton based on the constrained locally linear embedding. This project provides the ***Python version of SUDE***, and the MATLAB version can be found at https://github.com/ZPGuiGroupWhu/sude. This paper has been published in ***Nature Machine Intelligence***, and more details can be seen https://www.nature.com/articles/s42256-025-01112-9.
|
|
3
|
+
|
|
4
|
+

|
|
5
|
+
|
|
6
|
+
# Installation
|
|
7
|
+
Supported `python` versions are `3.8` and above.
|
|
8
|
+
|
|
9
|
+
This project has been uploaded to [PyPI](https://pypi.org/project/sude/), supporting direct download and installation from pypi
|
|
10
|
+
|
|
11
|
+
```
|
|
12
|
+
pip install sude
|
|
13
|
+
```
|
|
14
|
+
|
|
15
|
+
## Manual Installation
|
|
16
|
+
|
|
17
|
+
```
|
|
18
|
+
git clone https://github.com/ZPGuiGroupWhu/SUDE-pkg.git
|
|
19
|
+
cd SUDE-pkg
|
|
20
|
+
pip install -e .
|
|
21
|
+
```
|
|
22
|
+
|
|
23
|
+
# How To Run
|
|
24
|
+
The SUDE algorithm package provides the `sude` function for clustering.
|
|
25
|
+
|
|
26
|
+
The description of the hyperparameters for user configuration are presented as follows
|
|
27
|
+
```python
|
|
28
|
+
def sude(
|
|
29
|
+
X,
|
|
30
|
+
no_dims = 2,
|
|
31
|
+
k1 = 20,
|
|
32
|
+
normalize = True,
|
|
33
|
+
large = False,
|
|
34
|
+
initialize = 'le',
|
|
35
|
+
agg_coef = 1.2,
|
|
36
|
+
T_epoch = 50,
|
|
37
|
+
):
|
|
38
|
+
"""
|
|
39
|
+
This function returns representation of the N by D matrix X in the lower-dimensional space. Each row in X
|
|
40
|
+
represents an observation.
|
|
41
|
+
|
|
42
|
+
Parameters are:
|
|
43
|
+
|
|
44
|
+
'no_dims' - A positive integer specifying the number of dimension of the representation Y.
|
|
45
|
+
Default: 2
|
|
46
|
+
'k1' - A non-negative integer specifying the number of nearest neighbors for PPS to
|
|
47
|
+
sample landmarks. It must be smaller than N.
|
|
48
|
+
Default: adaptive
|
|
49
|
+
'normalize' - Logical scalar. If true, normalize X using min-max normalization. If features in
|
|
50
|
+
X are on different scales, 'Normalize' should be set to true because the learning
|
|
51
|
+
process is based on nearest neighbors and features with large scales can override
|
|
52
|
+
the contribution of features with small scales.
|
|
53
|
+
Default: True
|
|
54
|
+
'large' - Logical scalar. If true, the data can be split into multiple blocks to avoid the problem
|
|
55
|
+
of memory overflow, and the gradient can be computed block by block using 'learning_l' function.
|
|
56
|
+
Default: False
|
|
57
|
+
'initialize' - A string specifying the method for initializing Y before manifold learning.
|
|
58
|
+
'le' - Laplacian eigenmaps.
|
|
59
|
+
'pca' - Principal component analysis.
|
|
60
|
+
'mds' - Multidimensional scaling.
|
|
61
|
+
Default: 'le'
|
|
62
|
+
'agg_coef' - A positive scalar specifying the aggregation coefficient.
|
|
63
|
+
Default: 1.2
|
|
64
|
+
'T_epoch' - Maximum number of epochs to take.
|
|
65
|
+
Default: 50
|
|
66
|
+
"""
|
|
67
|
+
```
|
|
68
|
+
|
|
69
|
+
After installing the SUDE library, you can use this function as follows:
|
|
70
|
+
```python
|
|
71
|
+
import pandas as pd
|
|
72
|
+
import numpy as np
|
|
73
|
+
from sude import sude
|
|
74
|
+
import time
|
|
75
|
+
import matplotlib.pyplot as plt
|
|
76
|
+
|
|
77
|
+
# Input data
|
|
78
|
+
data = np.array(pd.read_csv('benchmarks/rice.csv', header=None))
|
|
79
|
+
|
|
80
|
+
# Obtain data size and true annotations
|
|
81
|
+
m = data.shape[1]
|
|
82
|
+
X = data[:, :m - 1]
|
|
83
|
+
ref = data[:, m - 1]
|
|
84
|
+
|
|
85
|
+
# Perform SUDE embedding
|
|
86
|
+
start_time = time.time()
|
|
87
|
+
Y = sude(X, k1=10)
|
|
88
|
+
end_time = time.time()
|
|
89
|
+
print("Elapsed time:", end_time - start_time, 's')
|
|
90
|
+
|
|
91
|
+
plt.scatter(Y[:, 0], Y[:, 1], c=ref, cmap='tab10', s=4)
|
|
92
|
+
plt.show()
|
|
93
|
+
```
|
|
94
|
+
|
|
95
|
+
# Citation Request
|
|
96
|
+
Peng, D., Gui, Z., Wei, W. et al. Sampling-enabled scalable manifold learning unveils the discriminative cluster structure of high-dimensional data. Nat. Mach. Intell. (2025). https://doi.org/10.1038/s42256-025-01112-9
|
|
97
|
+
|
|
98
|
+
|
|
99
|
+
# License
|
|
100
|
+
SUDE is released under the MIT License.
|
|
101
|
+
|
|
102
|
+
|
|
103
|
+
|
|
104
|
+
|
|
105
|
+
|