seraplot 2.3.89__tar.gz → 2.4.1__tar.gz
This diff represents the content of publicly available package versions that have been released to one of the supported registries. The information contained in this diff is provided for informational purposes only and reflects changes between package versions as they appear in their respective public registries.
- {seraplot-2.3.89 → seraplot-2.4.1}/Cargo.lock +1 -1
- {seraplot-2.3.89 → seraplot-2.4.1}/Cargo.toml +9 -1
- {seraplot-2.3.89 → seraplot-2.4.1}/PKG-INFO +24 -35
- {seraplot-2.3.89 → seraplot-2.4.1}/README.md +23 -34
- seraplot-2.4.1/maturin_output.log +0 -0
- seraplot-2.4.1/publish_v241.log +0 -0
- {seraplot-2.3.89 → seraplot-2.4.1}/pyproject.toml +1 -1
- seraplot-2.4.1/src/.gitignore +4 -0
- {seraplot-2.3.89 → seraplot-2.4.1}/src/bindings/commands/charts.rs +374 -4
- {seraplot-2.3.89 → seraplot-2.4.1}/src/bindings/commands/ml.rs +477 -0
- {seraplot-2.3.89 → seraplot-2.4.1}/src/bindings/registry_macro.rs +72 -0
- {seraplot-2.3.89 → seraplot-2.4.1}/src/book.toml +1 -1
- seraplot-2.4.1/src/docs/SUMMARY.md +151 -0
- seraplot-2.4.1/src/docs/about/index.md +13 -0
- seraplot-2.4.1/src/docs/about/support.md +96 -0
- {seraplot-2.3.89 → seraplot-2.4.1}/src/docs/charts/2d/area.md +5 -1
- {seraplot-2.3.89 → seraplot-2.4.1}/src/docs/charts/2d/bar.md +5 -1
- {seraplot-2.3.89 → seraplot-2.4.1}/src/docs/charts/2d/boxplot.md +5 -1
- {seraplot-2.3.89 → seraplot-2.4.1}/src/docs/charts/2d/bubble.md +5 -1
- {seraplot-2.3.89 → seraplot-2.4.1}/src/docs/charts/2d/bullet.md +5 -1
- {seraplot-2.3.89 → seraplot-2.4.1}/src/docs/charts/2d/candlestick.md +5 -1
- {seraplot-2.3.89 → seraplot-2.4.1}/src/docs/charts/2d/donut.md +5 -1
- {seraplot-2.3.89 → seraplot-2.4.1}/src/docs/charts/2d/dumbbell.md +5 -1
- {seraplot-2.3.89 → seraplot-2.4.1}/src/docs/charts/2d/funnel.md +5 -1
- {seraplot-2.3.89 → seraplot-2.4.1}/src/docs/charts/2d/gauge.md +5 -1
- {seraplot-2.3.89 → seraplot-2.4.1}/src/docs/charts/2d/grid.md +5 -1
- {seraplot-2.3.89 → seraplot-2.4.1}/src/docs/charts/2d/grouped-bar.md +5 -1
- {seraplot-2.3.89 → seraplot-2.4.1}/src/docs/charts/2d/hbar.md +5 -1
- {seraplot-2.3.89 → seraplot-2.4.1}/src/docs/charts/2d/heatmap.md +5 -1
- {seraplot-2.3.89 → seraplot-2.4.1}/src/docs/charts/2d/histogram-overlay.md +5 -1
- {seraplot-2.3.89 → seraplot-2.4.1}/src/docs/charts/2d/histogram.md +5 -1
- {seraplot-2.3.89 → seraplot-2.4.1}/src/docs/charts/2d/kde.md +5 -1
- {seraplot-2.3.89 → seraplot-2.4.1}/src/docs/charts/2d/line.md +5 -1
- {seraplot-2.3.89 → seraplot-2.4.1}/src/docs/charts/2d/lollipop.md +5 -1
- {seraplot-2.3.89 → seraplot-2.4.1}/src/docs/charts/2d/multiline.md +5 -1
- {seraplot-2.3.89 → seraplot-2.4.1}/src/docs/charts/2d/parallel.md +5 -1
- {seraplot-2.3.89 → seraplot-2.4.1}/src/docs/charts/2d/pie.md +5 -1
- {seraplot-2.3.89 → seraplot-2.4.1}/src/docs/charts/2d/radar.md +5 -1
- {seraplot-2.3.89 → seraplot-2.4.1}/src/docs/charts/2d/ridgeline.md +5 -1
- {seraplot-2.3.89 → seraplot-2.4.1}/src/docs/charts/2d/scatter.md +5 -1
- {seraplot-2.3.89 → seraplot-2.4.1}/src/docs/charts/2d/slideshow.md +5 -1
- {seraplot-2.3.89 → seraplot-2.4.1}/src/docs/charts/2d/slope.md +5 -1
- {seraplot-2.3.89 → seraplot-2.4.1}/src/docs/charts/2d/stacked-bar.md +5 -1
- {seraplot-2.3.89 → seraplot-2.4.1}/src/docs/charts/2d/sunburst.md +5 -1
- {seraplot-2.3.89 → seraplot-2.4.1}/src/docs/charts/2d/treemap.md +5 -1
- {seraplot-2.3.89 → seraplot-2.4.1}/src/docs/charts/2d/violin.md +5 -1
- {seraplot-2.3.89 → seraplot-2.4.1}/src/docs/charts/2d/waterfall.md +5 -1
- {seraplot-2.3.89 → seraplot-2.4.1}/src/docs/charts/2d/wordcloud.md +5 -1
- {seraplot-2.3.89 → seraplot-2.4.1}/src/docs/charts/3d/bar3d.md +5 -1
- {seraplot-2.3.89 → seraplot-2.4.1}/src/docs/charts/3d/bubble3d.md +5 -1
- {seraplot-2.3.89 → seraplot-2.4.1}/src/docs/charts/3d/candlestick3d.md +5 -1
- {seraplot-2.3.89 → seraplot-2.4.1}/src/docs/charts/3d/dumbbell3d.md +5 -1
- {seraplot-2.3.89 → seraplot-2.4.1}/src/docs/charts/3d/funnel3d.md +5 -1
- {seraplot-2.3.89 → seraplot-2.4.1}/src/docs/charts/3d/globe3d.md +5 -1
- {seraplot-2.3.89 → seraplot-2.4.1}/src/docs/charts/3d/heatmap3d.md +5 -1
- {seraplot-2.3.89 → seraplot-2.4.1}/src/docs/charts/3d/kde3d.md +5 -1
- {seraplot-2.3.89 → seraplot-2.4.1}/src/docs/charts/3d/line3d.md +5 -1
- {seraplot-2.3.89 → seraplot-2.4.1}/src/docs/charts/3d/lollipop3d.md +5 -1
- {seraplot-2.3.89 → seraplot-2.4.1}/src/docs/charts/3d/pie3d.md +5 -1
- {seraplot-2.3.89 → seraplot-2.4.1}/src/docs/charts/3d/radar3d.md +5 -1
- {seraplot-2.3.89 → seraplot-2.4.1}/src/docs/charts/3d/ridgeline3d.md +5 -1
- {seraplot-2.3.89 → seraplot-2.4.1}/src/docs/charts/3d/scatter3d.md +49 -8
- {seraplot-2.3.89 → seraplot-2.4.1}/src/docs/charts/3d/stacked-bar3d.md +5 -1
- {seraplot-2.3.89 → seraplot-2.4.1}/src/docs/charts/3d/sunburst3d.md +5 -1
- {seraplot-2.3.89 → seraplot-2.4.1}/src/docs/charts/3d/violin3d.md +5 -1
- seraplot-2.4.1/src/docs/charts/index.md +13 -0
- {seraplot-2.3.89 → seraplot-2.4.1}/src/docs/charts/map/bubble-map.md +5 -1
- {seraplot-2.3.89 → seraplot-2.4.1}/src/docs/charts/map/choropleth.md +5 -1
- {seraplot-2.3.89 → seraplot-2.4.1}/src/docs/config/automl.md +6 -2
- seraplot-2.4.1/src/docs/config/index.md +13 -0
- {seraplot-2.3.89 → seraplot-2.4.1}/src/docs/config/pickle.md +26 -0
- seraplot-2.4.1/src/docs/config/themes.md +273 -0
- {seraplot-2.3.89 → seraplot-2.4.1}/src/docs/getting-started/chart-methods.md +784 -503
- {seraplot-2.3.89 → seraplot-2.4.1}/src/docs/getting-started/chart-object.md +2 -10
- seraplot-2.4.1/src/docs/getting-started/index.md +109 -0
- {seraplot-2.3.89 → seraplot-2.4.1}/src/docs/getting-started/installation.md +0 -56
- seraplot-2.4.1/src/docs/getting-started/quickstart.md +105 -0
- seraplot-2.4.1/src/docs/introduction.md +682 -0
- seraplot-2.4.1/src/docs/ml/adaboost.md +203 -0
- seraplot-2.4.1/src/docs/ml/cv-splitters.md +121 -0
- seraplot-2.4.1/src/docs/ml/dbscan-class.md +203 -0
- seraplot-2.4.1/src/docs/ml/dbscan.md +427 -0
- {seraplot-2.3.89 → seraplot-2.4.1}/src/docs/ml/dbscan3d.md +212 -167
- {seraplot-2.3.89 → seraplot-2.4.1}/src/docs/ml/decision-tree.md +10 -10
- {seraplot-2.3.89 → seraplot-2.4.1}/src/docs/ml/decomposition.md +16 -16
- {seraplot-2.3.89 → seraplot-2.4.1}/src/docs/ml/elastic-net.md +174 -174
- {seraplot-2.3.89 → seraplot-2.4.1}/src/docs/ml/gradient-boosting.md +10 -10
- {seraplot-2.3.89 → seraplot-2.4.1}/src/docs/ml/grid-search.md +6 -6
- seraplot-2.4.1/src/docs/ml/isolation-forest.md +129 -0
- {seraplot-2.3.89 → seraplot-2.4.1}/src/docs/ml/kmeans-class.md +55 -1
- seraplot-2.4.1/src/docs/ml/kmeans.md +336 -0
- {seraplot-2.3.89 → seraplot-2.4.1}/src/docs/ml/knn.md +10 -10
- {seraplot-2.3.89 → seraplot-2.4.1}/src/docs/ml/lasso.md +158 -158
- {seraplot-2.3.89 → seraplot-2.4.1}/src/docs/ml/linear-regression.md +150 -150
- {seraplot-2.3.89 → seraplot-2.4.1}/src/docs/ml/logistic-regression.md +170 -170
- seraplot-2.4.1/src/docs/ml/metrics-classification.md +311 -0
- seraplot-2.4.1/src/docs/ml/metrics-clustering.md +203 -0
- seraplot-2.4.1/src/docs/ml/metrics-regression.md +197 -0
- {seraplot-2.3.89 → seraplot-2.4.1}/src/docs/ml/metrics.md +14 -14
- {seraplot-2.3.89 → seraplot-2.4.1}/src/docs/ml/naive-bayes.md +14 -14
- seraplot-2.4.1/src/docs/ml/permutation-importance.md +137 -0
- seraplot-2.4.1/src/docs/ml/preprocessing-advanced.md +359 -0
- {seraplot-2.3.89 → seraplot-2.4.1}/src/docs/ml/preprocessing.md +12 -12
- {seraplot-2.3.89 → seraplot-2.4.1}/src/docs/ml/random-forest.md +10 -10
- {seraplot-2.3.89 → seraplot-2.4.1}/src/docs/ml/ridge.md +176 -176
- {seraplot-2.3.89 → seraplot-2.4.1}/src/docs/ml/sgd.md +6 -6
- {seraplot-2.3.89 → seraplot-2.4.1}/src/docs/ml/svm.md +8 -8
- {seraplot-2.3.89 → seraplot-2.4.1}/src/docs/ml/train-test-split.md +6 -6
- {seraplot-2.3.89 → seraplot-2.4.1}/src/docs/theme/custom.css +28 -0
- seraplot-2.4.1/src/docs/theme/donate-popup.js +93 -0
- seraplot-2.4.1/src/docs/theme/lang-switcher.js +173 -0
- seraplot-2.4.1/src/docs/tooling/index.md +13 -0
- seraplot-2.4.1/src/docs/tooling/vscode.md +165 -0
- seraplot-2.4.1/src/ml/anomaly/isolation_forest.rs +153 -0
- seraplot-2.4.1/src/ml/anomaly/mod.rs +3 -0
- seraplot-2.4.1/src/ml/metrics/classification.rs +347 -0
- seraplot-2.4.1/src/ml/metrics/clustering.rs +273 -0
- {seraplot-2.3.89 → seraplot-2.4.1}/src/ml/metrics/mod.rs +2 -0
- {seraplot-2.3.89 → seraplot-2.4.1}/src/ml/metrics/regression.rs +46 -0
- {seraplot-2.3.89 → seraplot-2.4.1}/src/ml/mod.rs +2 -0
- {seraplot-2.3.89 → seraplot-2.4.1}/src/ml/model_selection/grid_search.rs +1 -32
- {seraplot-2.3.89 → seraplot-2.4.1}/src/ml/model_selection/mod.rs +2 -0
- seraplot-2.4.1/src/ml/model_selection/permutation.rs +68 -0
- {seraplot-2.3.89 → seraplot-2.4.1}/src/ml/model_selection/split.rs +60 -0
- {seraplot-2.3.89 → seraplot-2.4.1}/src/ml/preprocessing/encoders.rs +50 -0
- {seraplot-2.3.89 → seraplot-2.4.1}/src/ml/preprocessing/mod.rs +2 -0
- {seraplot-2.3.89 → seraplot-2.4.1}/src/ml/preprocessing/scalers.rs +87 -3
- seraplot-2.4.1/src/ml/preprocessing/transformers.rs +312 -0
- seraplot-2.3.89/src/docs/SUMMARY.md +0 -149
- seraplot-2.3.89/src/docs/getting-started/quickstart.md +0 -288
- seraplot-2.3.89/src/docs/introduction.md +0 -790
- seraplot-2.3.89/src/docs/ml/adaboost.md +0 -185
- seraplot-2.3.89/src/docs/ml/dbscan-class.md +0 -191
- seraplot-2.3.89/src/docs/ml/dbscan.md +0 -214
- seraplot-2.3.89/src/docs/ml/kmeans.md +0 -189
- seraplot-2.3.89/src/docs/theme/lang-switcher.js +0 -141
- seraplot-2.3.89/src/ml/metrics/classification.rs +0 -125
- {seraplot-2.3.89 → seraplot-2.4.1}/.gitignore +0 -0
- {seraplot-2.3.89 → seraplot-2.4.1}/05--Diabetes_SeraPlot.ipynb +0 -0
- {seraplot-2.3.89 → seraplot-2.4.1}/07--OpenFoodFacts_SeraPlot_Benchmark.ipynb +0 -0
- {seraplot-2.3.89 → seraplot-2.4.1}/TEST_1M_Battle.ipynb +0 -0
- {seraplot-2.3.89 → seraplot-2.4.1}/bench_all.py +0 -0
- {seraplot-2.3.89 → seraplot-2.4.1}/bench_bigdata.py +0 -0
- {seraplot-2.3.89 → seraplot-2.4.1}/bench_full.py +0 -0
- {seraplot-2.3.89 → seraplot-2.4.1}/bench_issues.json +0 -0
- {seraplot-2.3.89 → seraplot-2.4.1}/bench_knn_sgd.py +0 -0
- {seraplot-2.3.89 → seraplot-2.4.1}/bench_perf.py +0 -0
- {seraplot-2.3.89 → seraplot-2.4.1}/bench_quick.py +0 -0
- {seraplot-2.3.89 → seraplot-2.4.1}/bench_quick2.py +0 -0
- {seraplot-2.3.89 → seraplot-2.4.1}/bench_scale.py +0 -0
- {seraplot-2.3.89 → seraplot-2.4.1}/bench_seraplot.py +0 -0
- {seraplot-2.3.89 → seraplot-2.4.1}/bench_sklearn.py +0 -0
- {seraplot-2.3.89 → seraplot-2.4.1}/bench_svc.py +0 -0
- {seraplot-2.3.89 → seraplot-2.4.1}/bench_svc2.py +0 -0
- {seraplot-2.3.89 → seraplot-2.4.1}/bench_test.py +0 -0
- {seraplot-2.3.89 → seraplot-2.4.1}/benchmark_gridsearch.py +0 -0
- {seraplot-2.3.89 → seraplot-2.4.1}/benchmark_gridsearch2.py +0 -0
- {seraplot-2.3.89 → seraplot-2.4.1}/benchmark_kmeans_heavy.ipynb +0 -0
- {seraplot-2.3.89 → seraplot-2.4.1}/benchmark_kmeans_openfoodfacts.ipynb +0 -0
- {seraplot-2.3.89 → seraplot-2.4.1}/benchmark_ml.ipynb +0 -0
- {seraplot-2.3.89 → seraplot-2.4.1}/benchmark_ml_openfoodfacts.ipynb +0 -0
- {seraplot-2.3.89 → seraplot-2.4.1}/benchmark_ml_out.ipynb +0 -0
- {seraplot-2.3.89 → seraplot-2.4.1}/build.log +0 -0
- {seraplot-2.3.89 → seraplot-2.4.1}/gen_bench_slides.py +0 -0
- {seraplot-2.3.89 → seraplot-2.4.1}/images/2d/1.png +0 -0
- {seraplot-2.3.89 → seraplot-2.4.1}/images/2d/2.png +0 -0
- {seraplot-2.3.89 → seraplot-2.4.1}/images/2d/3.png +0 -0
- {seraplot-2.3.89 → seraplot-2.4.1}/images/2d/4.png +0 -0
- {seraplot-2.3.89 → seraplot-2.4.1}/images/2d/5.png +0 -0
- {seraplot-2.3.89 → seraplot-2.4.1}/images/2d/6.png +0 -0
- {seraplot-2.3.89 → seraplot-2.4.1}/images/2d/7.png +0 -0
- {seraplot-2.3.89 → seraplot-2.4.1}/images/2d/8.png +0 -0
- {seraplot-2.3.89 → seraplot-2.4.1}/images/3d/1.png +0 -0
- {seraplot-2.3.89 → seraplot-2.4.1}/images/3d/2.png +0 -0
- {seraplot-2.3.89 → seraplot-2.4.1}/images/3d/3.png +0 -0
- {seraplot-2.3.89 → seraplot-2.4.1}/images/3d/4.png +0 -0
- {seraplot-2.3.89 → seraplot-2.4.1}/images/3d/5.png +0 -0
- {seraplot-2.3.89 → seraplot-2.4.1}/images/3d/6.png +0 -0
- {seraplot-2.3.89 → seraplot-2.4.1}/images/3d/7.png +0 -0
- {seraplot-2.3.89 → seraplot-2.4.1}/images/3d/8.png +0 -0
- {seraplot-2.3.89 → seraplot-2.4.1}/images/README.md +0 -0
- {seraplot-2.3.89 → seraplot-2.4.1}/images/benchmark/1.png +0 -0
- {seraplot-2.3.89 → seraplot-2.4.1}/images/benchmark/2.png +0 -0
- {seraplot-2.3.89 → seraplot-2.4.1}/images/benchmark/3.png +0 -0
- {seraplot-2.3.89 → seraplot-2.4.1}/images/logo.png +0 -0
- {seraplot-2.3.89 → seraplot-2.4.1}/kmeans_test.html +0 -0
- {seraplot-2.3.89 → seraplot-2.4.1}/publish_2378.log +0 -0
- {seraplot-2.3.89 → seraplot-2.4.1}/publish_log.txt +0 -0
- {seraplot-2.3.89 → seraplot-2.4.1}/python/seraplot/__init__.py +0 -0
- {seraplot-2.3.89 → seraplot-2.4.1}/python/seraplot/matplotlib.py +0 -0
- {seraplot-2.3.89 → seraplot-2.4.1}/python/seraplot/seraplot.cp311-win_amd64.pyd.old +0 -0
- {seraplot-2.3.89 → seraplot-2.4.1}/results_seraplot.json +0 -0
- {seraplot-2.3.89 → seraplot-2.4.1}/results_sklearn.json +0 -0
- {seraplot-2.3.89 → seraplot-2.4.1}/src/.github/workflows/mdbook.yml +0 -0
- {seraplot-2.3.89 → seraplot-2.4.1}/src/README.md +0 -0
- {seraplot-2.3.89 → seraplot-2.4.1}/src/asset/SVG World Map with labels.svg +0 -0
- {seraplot-2.3.89 → seraplot-2.4.1}/src/asset/logo.png +0 -0
- {seraplot-2.3.89 → seraplot-2.4.1}/src/asset/world.svg +0 -0
- {seraplot-2.3.89 → seraplot-2.4.1}/src/bindings/builder_template.rs +0 -0
- {seraplot-2.3.89 → seraplot-2.4.1}/src/bindings/chart_types.rs +0 -0
- {seraplot-2.3.89 → seraplot-2.4.1}/src/bindings/commands/docs.rs +0 -0
- {seraplot-2.3.89 → seraplot-2.4.1}/src/bindings/commands/mod.rs +0 -0
- {seraplot-2.3.89 → seraplot-2.4.1}/src/bindings/commands/native.rs +0 -0
- {seraplot-2.3.89 → seraplot-2.4.1}/src/bindings/commands/registry.rs +0 -0
- {seraplot-2.3.89 → seraplot-2.4.1}/src/bindings/export_builder.rs +0 -0
- {seraplot-2.3.89 → seraplot-2.4.1}/src/bindings/fast_export_c.rs +0 -0
- {seraplot-2.3.89 → seraplot-2.4.1}/src/bindings/fast_render.rs +0 -0
- {seraplot-2.3.89 → seraplot-2.4.1}/src/bindings/memory_pool.rs +0 -0
- {seraplot-2.3.89 → seraplot-2.4.1}/src/bindings/mod.rs +0 -0
- {seraplot-2.3.89 → seraplot-2.4.1}/src/bindings/unified_builder.rs +0 -0
- {seraplot-2.3.89 → seraplot-2.4.1}/src/bindings/unified_config.rs +0 -0
- {seraplot-2.3.89 → seraplot-2.4.1}/src/bindings/utils/arena_alloc.rs +0 -0
- {seraplot-2.3.89 → seraplot-2.4.1}/src/bindings/utils/bitset.rs +0 -0
- {seraplot-2.3.89 → seraplot-2.4.1}/src/bindings/utils/compact_state.rs +0 -0
- {seraplot-2.3.89 → seraplot-2.4.1}/src/bindings/utils/data_processor.rs +0 -0
- {seraplot-2.3.89 → seraplot-2.4.1}/src/bindings/utils/image_processor.rs +0 -0
- {seraplot-2.3.89 → seraplot-2.4.1}/src/bindings/utils/lazy_builders.rs +0 -0
- {seraplot-2.3.89 → seraplot-2.4.1}/src/bindings/utils/memory_pool.rs +0 -0
- {seraplot-2.3.89 → seraplot-2.4.1}/src/bindings/utils/mod.rs +0 -0
- {seraplot-2.3.89 → seraplot-2.4.1}/src/bindings/utils/simd_ops.rs +0 -0
- {seraplot-2.3.89 → seraplot-2.4.1}/src/bindings/utils/state_export.rs +0 -0
- {seraplot-2.3.89 → seraplot-2.4.1}/src/core/builders.rs +0 -0
- {seraplot-2.3.89 → seraplot-2.4.1}/src/core/math.rs +0 -0
- {seraplot-2.3.89 → seraplot-2.4.1}/src/core/mod.rs +0 -0
- {seraplot-2.3.89 → seraplot-2.4.1}/src/data/conversion.rs +0 -0
- {seraplot-2.3.89 → seraplot-2.4.1}/src/data/index.rs +0 -0
- {seraplot-2.3.89 → seraplot-2.4.1}/src/data/loader.rs +0 -0
- {seraplot-2.3.89 → seraplot-2.4.1}/src/data/mod.rs +0 -0
- {seraplot-2.3.89 → seraplot-2.4.1}/src/data/processor.rs +0 -0
- {seraplot-2.3.89 → seraplot-2.4.1}/src/docs/api/index.md +0 -0
- {seraplot-2.3.89 → seraplot-2.4.1}/src/docs/charts/2d/index.md +0 -0
- {seraplot-2.3.89 → seraplot-2.4.1}/src/docs/charts/3d/index.md +0 -0
- {seraplot-2.3.89 → seraplot-2.4.1}/src/docs/charts/map/index.md +0 -0
- {seraplot-2.3.89 → seraplot-2.4.1}/src/docs/config/a11y.md +0 -0
- {seraplot-2.3.89 → seraplot-2.4.1}/src/docs/config/auto-display.md +0 -0
- {seraplot-2.3.89 → seraplot-2.4.1}/src/docs/config/background.md +0 -0
- {seraplot-2.3.89 → seraplot-2.4.1}/src/docs/config/csp.md +0 -0
- {seraplot-2.3.89 → seraplot-2.4.1}/src/docs/config/diff.md +0 -0
- {seraplot-2.3.89 → seraplot-2.4.1}/src/docs/config/downsample.md +0 -0
- {seraplot-2.3.89 → seraplot-2.4.1}/src/docs/config/drift.md +0 -0
- {seraplot-2.3.89 → seraplot-2.4.1}/src/docs/config/export.md +0 -0
- {seraplot-2.3.89 → seraplot-2.4.1}/src/docs/config/facet.md +0 -0
- {seraplot-2.3.89 → seraplot-2.4.1}/src/docs/config/hover.md +0 -0
- {seraplot-2.3.89 → seraplot-2.4.1}/src/docs/config/ml-persistence.md +0 -0
- {seraplot-2.3.89 → seraplot-2.4.1}/src/docs/config/palette.md +0 -0
- {seraplot-2.3.89 → seraplot-2.4.1}/src/docs/config/streaming.md +0 -0
- {seraplot-2.3.89 → seraplot-2.4.1}/src/docs/ml/bayes-index.md +0 -0
- {seraplot-2.3.89 → seraplot-2.4.1}/src/docs/ml/clustering-index.md +0 -0
- {seraplot-2.3.89 → seraplot-2.4.1}/src/docs/ml/decomposition-index.md +0 -0
- {seraplot-2.3.89 → seraplot-2.4.1}/src/docs/ml/evaluation-index.md +0 -0
- {seraplot-2.3.89 → seraplot-2.4.1}/src/docs/ml/index.md +0 -0
- {seraplot-2.3.89 → seraplot-2.4.1}/src/docs/ml/linear-index.md +0 -0
- {seraplot-2.3.89 → seraplot-2.4.1}/src/docs/ml/neighbors-index.md +0 -0
- {seraplot-2.3.89 → seraplot-2.4.1}/src/docs/ml/preprocessing-index.md +0 -0
- {seraplot-2.3.89 → seraplot-2.4.1}/src/docs/ml/selection-index.md +0 -0
- {seraplot-2.3.89 → seraplot-2.4.1}/src/docs/ml/svm-index.md +0 -0
- {seraplot-2.3.89 → seraplot-2.4.1}/src/docs/ml/tree-index.md +0 -0
- {seraplot-2.3.89 → seraplot-2.4.1}/src/docs/previews/area.html +0 -0
- {seraplot-2.3.89 → seraplot-2.4.1}/src/docs/previews/bar.html +0 -0
- {seraplot-2.3.89 → seraplot-2.4.1}/src/docs/previews/bar3d.html +0 -0
- {seraplot-2.3.89 → seraplot-2.4.1}/src/docs/previews/bench-matplotlib.html +0 -0
- {seraplot-2.3.89 → seraplot-2.4.1}/src/docs/previews/bench-plotly.html +0 -0
- {seraplot-2.3.89 → seraplot-2.4.1}/src/docs/previews/bench-seraplot.html +0 -0
- {seraplot-2.3.89 → seraplot-2.4.1}/src/docs/previews/boxplot.html +0 -0
- {seraplot-2.3.89 → seraplot-2.4.1}/src/docs/previews/bubble-map.html +0 -0
- {seraplot-2.3.89 → seraplot-2.4.1}/src/docs/previews/bubble.html +0 -0
- {seraplot-2.3.89 → seraplot-2.4.1}/src/docs/previews/bubble3d.html +0 -0
- {seraplot-2.3.89 → seraplot-2.4.1}/src/docs/previews/bullet.html +0 -0
- {seraplot-2.3.89 → seraplot-2.4.1}/src/docs/previews/candlestick.html +0 -0
- {seraplot-2.3.89 → seraplot-2.4.1}/src/docs/previews/candlestick3d.html +0 -0
- {seraplot-2.3.89 → seraplot-2.4.1}/src/docs/previews/choropleth.html +0 -0
- {seraplot-2.3.89 → seraplot-2.4.1}/src/docs/previews/dbscan-class.html +0 -0
- {seraplot-2.3.89 → seraplot-2.4.1}/src/docs/previews/dbscan.html +0 -0
- {seraplot-2.3.89 → seraplot-2.4.1}/src/docs/previews/dbscan3d.html +0 -0
- {seraplot-2.3.89 → seraplot-2.4.1}/src/docs/previews/donut.html +0 -0
- {seraplot-2.3.89 → seraplot-2.4.1}/src/docs/previews/dumbbell.html +0 -0
- {seraplot-2.3.89 → seraplot-2.4.1}/src/docs/previews/dumbbell3d.html +0 -0
- {seraplot-2.3.89 → seraplot-2.4.1}/src/docs/previews/funnel.html +0 -0
- {seraplot-2.3.89 → seraplot-2.4.1}/src/docs/previews/funnel3d.html +0 -0
- {seraplot-2.3.89 → seraplot-2.4.1}/src/docs/previews/gauge.html +0 -0
- {seraplot-2.3.89 → seraplot-2.4.1}/src/docs/previews/globe3d.html +0 -0
- {seraplot-2.3.89 → seraplot-2.4.1}/src/docs/previews/grid.html +0 -0
- {seraplot-2.3.89 → seraplot-2.4.1}/src/docs/previews/grouped-bar.html +0 -0
- {seraplot-2.3.89 → seraplot-2.4.1}/src/docs/previews/hbar.html +0 -0
- {seraplot-2.3.89 → seraplot-2.4.1}/src/docs/previews/heatmap.html +0 -0
- {seraplot-2.3.89 → seraplot-2.4.1}/src/docs/previews/heatmap3d.html +0 -0
- {seraplot-2.3.89 → seraplot-2.4.1}/src/docs/previews/histogram-overlay.html +0 -0
- {seraplot-2.3.89 → seraplot-2.4.1}/src/docs/previews/histogram.html +0 -0
- {seraplot-2.3.89 → seraplot-2.4.1}/src/docs/previews/kde.html +0 -0
- {seraplot-2.3.89 → seraplot-2.4.1}/src/docs/previews/kde3d.html +0 -0
- {seraplot-2.3.89 → seraplot-2.4.1}/src/docs/previews/line.html +0 -0
- {seraplot-2.3.89 → seraplot-2.4.1}/src/docs/previews/line3d.html +0 -0
- {seraplot-2.3.89 → seraplot-2.4.1}/src/docs/previews/lollipop.html +0 -0
- {seraplot-2.3.89 → seraplot-2.4.1}/src/docs/previews/lollipop3d.html +0 -0
- {seraplot-2.3.89 → seraplot-2.4.1}/src/docs/previews/multiline.html +0 -0
- {seraplot-2.3.89 → seraplot-2.4.1}/src/docs/previews/parallel.html +0 -0
- {seraplot-2.3.89 → seraplot-2.4.1}/src/docs/previews/pie.html +0 -0
- {seraplot-2.3.89 → seraplot-2.4.1}/src/docs/previews/pie3d.html +0 -0
- {seraplot-2.3.89 → seraplot-2.4.1}/src/docs/previews/radar.html +0 -0
- {seraplot-2.3.89 → seraplot-2.4.1}/src/docs/previews/radar3d.html +0 -0
- {seraplot-2.3.89 → seraplot-2.4.1}/src/docs/previews/ridgeline.html +0 -0
- {seraplot-2.3.89 → seraplot-2.4.1}/src/docs/previews/ridgeline3d.html +0 -0
- {seraplot-2.3.89 → seraplot-2.4.1}/src/docs/previews/scatter.html +0 -0
- {seraplot-2.3.89 → seraplot-2.4.1}/src/docs/previews/scatter3d.html +0 -0
- {seraplot-2.3.89 → seraplot-2.4.1}/src/docs/previews/slideshow.html +0 -0
- {seraplot-2.3.89 → seraplot-2.4.1}/src/docs/previews/slope.html +0 -0
- {seraplot-2.3.89 → seraplot-2.4.1}/src/docs/previews/stacked-bar.html +0 -0
- {seraplot-2.3.89 → seraplot-2.4.1}/src/docs/previews/stacked-bar3d.html +0 -0
- {seraplot-2.3.89 → seraplot-2.4.1}/src/docs/previews/sunburst.html +0 -0
- {seraplot-2.3.89 → seraplot-2.4.1}/src/docs/previews/sunburst3d.html +0 -0
- {seraplot-2.3.89 → seraplot-2.4.1}/src/docs/previews/treemap.html +0 -0
- {seraplot-2.3.89 → seraplot-2.4.1}/src/docs/previews/violin.html +0 -0
- {seraplot-2.3.89 → seraplot-2.4.1}/src/docs/previews/violin3d.html +0 -0
- {seraplot-2.3.89 → seraplot-2.4.1}/src/docs/previews/waterfall.html +0 -0
- {seraplot-2.3.89 → seraplot-2.4.1}/src/docs/previews/wordcloud.html +0 -0
- {seraplot-2.3.89 → seraplot-2.4.1}/src/html/assets.rs +0 -0
- {seraplot-2.3.89 → seraplot-2.4.1}/src/html/fast_builders.rs +0 -0
- {seraplot-2.3.89 → seraplot-2.4.1}/src/html/fast_exporter.rs +0 -0
- {seraplot-2.3.89 → seraplot-2.4.1}/src/html/hover.rs +0 -0
- {seraplot-2.3.89 → seraplot-2.4.1}/src/html/html_export.rs +0 -0
- {seraplot-2.3.89 → seraplot-2.4.1}/src/html/html_template.rs +0 -0
- {seraplot-2.3.89 → seraplot-2.4.1}/src/html/js_3d.rs +0 -0
- {seraplot-2.3.89 → seraplot-2.4.1}/src/html/mod.rs +0 -0
- {seraplot-2.3.89 → seraplot-2.4.1}/src/lib.rs +0 -0
- {seraplot-2.3.89 → seraplot-2.4.1}/src/ml/cache.rs +0 -0
- {seraplot-2.3.89 → seraplot-2.4.1}/src/ml/decomposition/mod.rs +0 -0
- {seraplot-2.3.89 → seraplot-2.4.1}/src/ml/decomposition/pca.rs +0 -0
- {seraplot-2.3.89 → seraplot-2.4.1}/src/ml/linalg.rs +0 -0
- {seraplot-2.3.89 → seraplot-2.4.1}/src/ml/linear/elastic_net.rs +0 -0
- {seraplot-2.3.89 → seraplot-2.4.1}/src/ml/linear/lasso.rs +0 -0
- {seraplot-2.3.89 → seraplot-2.4.1}/src/ml/linear/logistic.rs +0 -0
- {seraplot-2.3.89 → seraplot-2.4.1}/src/ml/linear/mod.rs +0 -0
- {seraplot-2.3.89 → seraplot-2.4.1}/src/ml/linear/ols.rs +0 -0
- {seraplot-2.3.89 → seraplot-2.4.1}/src/ml/linear/ridge.rs +0 -0
- {seraplot-2.3.89 → seraplot-2.4.1}/src/ml/linear/sgd.rs +0 -0
- {seraplot-2.3.89 → seraplot-2.4.1}/src/ml/model_selection/cross_val.rs +0 -0
- {seraplot-2.3.89 → seraplot-2.4.1}/src/ml/naive_bayes/bernoulli.rs +0 -0
- {seraplot-2.3.89 → seraplot-2.4.1}/src/ml/naive_bayes/gaussian.rs +0 -0
- {seraplot-2.3.89 → seraplot-2.4.1}/src/ml/naive_bayes/mod.rs +0 -0
- {seraplot-2.3.89 → seraplot-2.4.1}/src/ml/naive_bayes/multinomial.rs +0 -0
- {seraplot-2.3.89 → seraplot-2.4.1}/src/ml/neighbors/knn.rs +0 -0
- {seraplot-2.3.89 → seraplot-2.4.1}/src/ml/neighbors/mod.rs +0 -0
- {seraplot-2.3.89 → seraplot-2.4.1}/src/ml/svm/mod.rs +0 -0
- {seraplot-2.3.89 → seraplot-2.4.1}/src/ml/svm/svm.rs +0 -0
- {seraplot-2.3.89 → seraplot-2.4.1}/src/ml/tree/adaboost.rs +0 -0
- {seraplot-2.3.89 → seraplot-2.4.1}/src/ml/tree/decision_tree.rs +0 -0
- {seraplot-2.3.89 → seraplot-2.4.1}/src/ml/tree/decision_tree_backup.rs +0 -0
- {seraplot-2.3.89 → seraplot-2.4.1}/src/ml/tree/gradient_boosting.rs +0 -0
- {seraplot-2.3.89 → seraplot-2.4.1}/src/ml/tree/mod.rs +0 -0
- {seraplot-2.3.89 → seraplot-2.4.1}/src/ml/tree/random_forest.rs +0 -0
- {seraplot-2.3.89 → seraplot-2.4.1}/src/plot/camera.rs +0 -0
- {seraplot-2.3.89 → seraplot-2.4.1}/src/plot/canvas.rs +0 -0
- {seraplot-2.3.89 → seraplot-2.4.1}/src/plot/containers_3d.rs +0 -0
- {seraplot-2.3.89 → seraplot-2.4.1}/src/plot/controller/chart_controller.rs +0 -0
- {seraplot-2.3.89 → seraplot-2.4.1}/src/plot/controller/mod.rs +0 -0
- {seraplot-2.3.89 → seraplot-2.4.1}/src/plot/controller/plot_3d_controller.rs +0 -0
- {seraplot-2.3.89 → seraplot-2.4.1}/src/plot/default/_3d/bar_3d.rs +0 -0
- {seraplot-2.3.89 → seraplot-2.4.1}/src/plot/default/_3d/line_3d.rs +0 -0
- {seraplot-2.3.89 → seraplot-2.4.1}/src/plot/default/_3d/mod.rs +0 -0
- {seraplot-2.3.89 → seraplot-2.4.1}/src/plot/default/_3d/plot_3d_types.rs +0 -0
- {seraplot-2.3.89 → seraplot-2.4.1}/src/plot/default/_3d/scatter_3d.rs +0 -0
- {seraplot-2.3.89 → seraplot-2.4.1}/src/plot/default/bar.rs +0 -0
- {seraplot-2.3.89 → seraplot-2.4.1}/src/plot/default/chart.rs +0 -0
- {seraplot-2.3.89 → seraplot-2.4.1}/src/plot/default/kmeans.rs +0 -0
- {seraplot-2.3.89 → seraplot-2.4.1}/src/plot/default/line.rs +0 -0
- {seraplot-2.3.89 → seraplot-2.4.1}/src/plot/default/mod.rs +0 -0
- {seraplot-2.3.89 → seraplot-2.4.1}/src/plot/default/scatter.rs +0 -0
- {seraplot-2.3.89 → seraplot-2.4.1}/src/plot/default/svg.rs +0 -0
- {seraplot-2.3.89 → seraplot-2.4.1}/src/plot/generic.rs +0 -0
- {seraplot-2.3.89 → seraplot-2.4.1}/src/plot/map/_3d/globe.rs +0 -0
- {seraplot-2.3.89 → seraplot-2.4.1}/src/plot/map/_3d/globe_html.rs +0 -0
- {seraplot-2.3.89 → seraplot-2.4.1}/src/plot/map/_3d/globe_types.rs +0 -0
- {seraplot-2.3.89 → seraplot-2.4.1}/src/plot/map/_3d/mod.rs +0 -0
- {seraplot-2.3.89 → seraplot-2.4.1}/src/plot/map/bubble_map.rs +0 -0
- {seraplot-2.3.89 → seraplot-2.4.1}/src/plot/map/chart.rs +0 -0
- {seraplot-2.3.89 → seraplot-2.4.1}/src/plot/map/choropleth.rs +0 -0
- {seraplot-2.3.89 → seraplot-2.4.1}/src/plot/map/mod.rs +0 -0
- {seraplot-2.3.89 → seraplot-2.4.1}/src/plot/map/svg_parser.rs +0 -0
- {seraplot-2.3.89 → seraplot-2.4.1}/src/plot/map/world_data.rs +0 -0
- {seraplot-2.3.89 → seraplot-2.4.1}/src/plot/mod.rs +0 -0
- {seraplot-2.3.89 → seraplot-2.4.1}/src/plot/projection.rs +0 -0
- {seraplot-2.3.89 → seraplot-2.4.1}/src/plot/renderers.rs +0 -0
- {seraplot-2.3.89 → seraplot-2.4.1}/src/plot/scale_renderer.rs +0 -0
- {seraplot-2.3.89 → seraplot-2.4.1}/src/plot/seaborn/_3d/mod.rs +0 -0
- {seraplot-2.3.89 → seraplot-2.4.1}/src/plot/seaborn/_3d/plot_3d_types.rs +0 -0
- {seraplot-2.3.89 → seraplot-2.4.1}/src/plot/seaborn/chart.rs +0 -0
- {seraplot-2.3.89 → seraplot-2.4.1}/src/plot/seaborn/mod.rs +0 -0
- {seraplot-2.3.89 → seraplot-2.4.1}/src/plot/statistical/_3d/candlestick3d.rs +0 -0
- {seraplot-2.3.89 → seraplot-2.4.1}/src/plot/statistical/_3d/dumbbell3d.rs +0 -0
- {seraplot-2.3.89 → seraplot-2.4.1}/src/plot/statistical/_3d/funnel3d.rs +0 -0
- {seraplot-2.3.89 → seraplot-2.4.1}/src/plot/statistical/_3d/heatmap3d.rs +0 -0
- {seraplot-2.3.89 → seraplot-2.4.1}/src/plot/statistical/_3d/kde3d.rs +0 -0
- {seraplot-2.3.89 → seraplot-2.4.1}/src/plot/statistical/_3d/lollipop3d.rs +0 -0
- {seraplot-2.3.89 → seraplot-2.4.1}/src/plot/statistical/_3d/mod.rs +0 -0
- {seraplot-2.3.89 → seraplot-2.4.1}/src/plot/statistical/_3d/pie3d.rs +0 -0
- {seraplot-2.3.89 → seraplot-2.4.1}/src/plot/statistical/_3d/plot_3d_types.rs +0 -0
- {seraplot-2.3.89 → seraplot-2.4.1}/src/plot/statistical/_3d/radar3d.rs +0 -0
- {seraplot-2.3.89 → seraplot-2.4.1}/src/plot/statistical/_3d/ridgeline3d.rs +0 -0
- {seraplot-2.3.89 → seraplot-2.4.1}/src/plot/statistical/_3d/stacked_bar3d.rs +0 -0
- {seraplot-2.3.89 → seraplot-2.4.1}/src/plot/statistical/_3d/sunburst3d.rs +0 -0
- {seraplot-2.3.89 → seraplot-2.4.1}/src/plot/statistical/_3d/violin3d.rs +0 -0
- {seraplot-2.3.89 → seraplot-2.4.1}/src/plot/statistical/area.rs +0 -0
- {seraplot-2.3.89 → seraplot-2.4.1}/src/plot/statistical/boxplot.rs +0 -0
- {seraplot-2.3.89 → seraplot-2.4.1}/src/plot/statistical/bubble.rs +0 -0
- {seraplot-2.3.89 → seraplot-2.4.1}/src/plot/statistical/bullet.rs +0 -0
- {seraplot-2.3.89 → seraplot-2.4.1}/src/plot/statistical/candlestick.rs +0 -0
- {seraplot-2.3.89 → seraplot-2.4.1}/src/plot/statistical/common.rs +0 -0
- {seraplot-2.3.89 → seraplot-2.4.1}/src/plot/statistical/dumbbell.rs +0 -0
- {seraplot-2.3.89 → seraplot-2.4.1}/src/plot/statistical/funnel.rs +0 -0
- {seraplot-2.3.89 → seraplot-2.4.1}/src/plot/statistical/gauge.rs +0 -0
- {seraplot-2.3.89 → seraplot-2.4.1}/src/plot/statistical/grouped_bar.rs +0 -0
- {seraplot-2.3.89 → seraplot-2.4.1}/src/plot/statistical/heatmap.rs +0 -0
- {seraplot-2.3.89 → seraplot-2.4.1}/src/plot/statistical/histogram.rs +0 -0
- {seraplot-2.3.89 → seraplot-2.4.1}/src/plot/statistical/kde.rs +0 -0
- {seraplot-2.3.89 → seraplot-2.4.1}/src/plot/statistical/lollipop.rs +0 -0
- {seraplot-2.3.89 → seraplot-2.4.1}/src/plot/statistical/mod.rs +0 -0
- {seraplot-2.3.89 → seraplot-2.4.1}/src/plot/statistical/multiline.rs +0 -0
- {seraplot-2.3.89 → seraplot-2.4.1}/src/plot/statistical/parallel.rs +0 -0
- {seraplot-2.3.89 → seraplot-2.4.1}/src/plot/statistical/pie.rs +0 -0
- {seraplot-2.3.89 → seraplot-2.4.1}/src/plot/statistical/radar.rs +0 -0
- {seraplot-2.3.89 → seraplot-2.4.1}/src/plot/statistical/ridgeline.rs +0 -0
- {seraplot-2.3.89 → seraplot-2.4.1}/src/plot/statistical/slope.rs +0 -0
- {seraplot-2.3.89 → seraplot-2.4.1}/src/plot/statistical/sunburst.rs +0 -0
- {seraplot-2.3.89 → seraplot-2.4.1}/src/plot/statistical/treemap.rs +0 -0
- {seraplot-2.3.89 → seraplot-2.4.1}/src/plot/statistical/violin.rs +0 -0
- {seraplot-2.3.89 → seraplot-2.4.1}/src/plot/statistical/waterfall.rs +0 -0
- {seraplot-2.3.89 → seraplot-2.4.1}/src/plot/statistical/wordcloud.rs +0 -0
- {seraplot-2.3.89 → seraplot-2.4.1}/src/viewer/cache.rs +0 -0
- {seraplot-2.3.89 → seraplot-2.4.1}/src/viewer/chart.rs +0 -0
- {seraplot-2.3.89 → seraplot-2.4.1}/src/viewer/gui.rs +0 -0
- {seraplot-2.3.89 → seraplot-2.4.1}/src/viewer/hybrid.rs +0 -0
- {seraplot-2.3.89 → seraplot-2.4.1}/src/viewer/manager/button_manager.rs +0 -0
- {seraplot-2.3.89 → seraplot-2.4.1}/src/viewer/manager/mod.rs +0 -0
- {seraplot-2.3.89 → seraplot-2.4.1}/src/viewer/mod.rs +0 -0
- {seraplot-2.3.89 → seraplot-2.4.1}/src/viewer/render/advanced_render.rs +0 -0
- {seraplot-2.3.89 → seraplot-2.4.1}/src/viewer/render/fast_render_gui.rs +0 -0
- {seraplot-2.3.89 → seraplot-2.4.1}/src/viewer/render/mod.rs +0 -0
- {seraplot-2.3.89 → seraplot-2.4.1}/src/viewer/render/pipeline.rs +0 -0
- {seraplot-2.3.89 → seraplot-2.4.1}/src/viewer/render/viewer_3d.rs +0 -0
- {seraplot-2.3.89 → seraplot-2.4.1}/src/viewer/render/wiki_viewer.rs +0 -0
- {seraplot-2.3.89 → seraplot-2.4.1}/src/viewer/utils/image_loader.rs +0 -0
- {seraplot-2.3.89 → seraplot-2.4.1}/src/viewer/utils/mod.rs +0 -0
- {seraplot-2.3.89 → seraplot-2.4.1}/src/wiki/api.rs +0 -0
- {seraplot-2.3.89 → seraplot-2.4.1}/src/wiki/extractor.rs +0 -0
- {seraplot-2.3.89 → seraplot-2.4.1}/src/wiki/language.rs +0 -0
- {seraplot-2.3.89 → seraplot-2.4.1}/src/wiki/macros.rs +0 -0
- {seraplot-2.3.89 → seraplot-2.4.1}/src/wiki/metadata.rs +0 -0
- {seraplot-2.3.89 → seraplot-2.4.1}/src/wiki/mod.rs +0 -0
- {seraplot-2.3.89 → seraplot-2.4.1}/test_accuracy.py +0 -0
- {seraplot-2.3.89 → seraplot-2.4.1}/test_universal_ml.js +0 -0
|
@@ -1,6 +1,6 @@
|
|
|
1
1
|
[package]
|
|
2
2
|
name = "seraplot"
|
|
3
|
-
version = "2.
|
|
3
|
+
version = "2.4.1"
|
|
4
4
|
edition = "2021"
|
|
5
5
|
authors = ["feur25"]
|
|
6
6
|
description = "Rust data visualization framework - The modern Plotly alternative"
|
|
@@ -44,6 +44,14 @@ ffi = ["dep:egui"]
|
|
|
44
44
|
all = ["python"]
|
|
45
45
|
pyo3 = ["dep:pyo3"]
|
|
46
46
|
|
|
47
|
+
[[example]]
|
|
48
|
+
name = "csv_test"
|
|
49
|
+
path = "examples/csv_test.rs"
|
|
50
|
+
|
|
51
|
+
[[example]]
|
|
52
|
+
name = "csv_viewer"
|
|
53
|
+
path = "examples/csv_viewer.rs"
|
|
54
|
+
|
|
47
55
|
[profile.release]
|
|
48
56
|
lto = "thin"
|
|
49
57
|
codegen-units = 1
|
|
@@ -1,6 +1,6 @@
|
|
|
1
1
|
Metadata-Version: 2.4
|
|
2
2
|
Name: seraplot
|
|
3
|
-
Version: 2.
|
|
3
|
+
Version: 2.4.1
|
|
4
4
|
Classifier: Development Status :: 5 - Production/Stable
|
|
5
5
|
Classifier: Environment :: Win32 (MS Windows)
|
|
6
6
|
Classifier: Intended Audience :: Developers
|
|
@@ -30,42 +30,47 @@ License: MIT
|
|
|
30
30
|
Requires-Python: >=3.8
|
|
31
31
|
Description-Content-Type: text/markdown; charset=UTF-8; variant=GFM
|
|
32
32
|
|
|
33
|
-
SeraPlot
|
|
33
|
+
SeraPlot — High-Performance Data Visualization & ML Framework
|
|
34
34
|
|
|
35
|
-
SeraPlot is a framework
|
|
36
|
-
alternative to Plotly, designed specifically for data visualization. This library is distributed
|
|
37
|
-
across multiple programming languages (Python, C#, C++, JavaScript), regularly maintained and
|
|
38
|
-
updated, offering superior speed and significantly lower memory consumption compared to competitors.
|
|
35
|
+
**SeraPlot v2.4.0+** is a production-grade framework written in **Rust**, delivering blazing-fast interactive charts and built-in machine learning pipelines. Designed as a modern alternative to Plotly + scikit-learn, it combines visualization and ML preprocessing in a single Rust binary — no dependencies, no bloat.
|
|
39
36
|
|
|
40
|
-
|
|
37
|
+
📖 **Documentation:** https://feur25.github.io/seraplot/introduction.html
|
|
41
38
|
|
|
42
39
|
---
|
|
43
40
|
|
|
44
|
-
|
|
41
|
+
## Why Choose SeraPlot?
|
|
45
42
|
|
|
46
|
-
|
|
47
|
-
|
|
48
|
-
|
|
49
|
-
|
|
50
|
-
|
|
51
|
-
|
|
43
|
+
**100–8000× faster** than Plotly & Matplotlib on chart generation
|
|
44
|
+
**Minimal memory footprint** — runs on edge devices, embedded systems, low-power servers
|
|
45
|
+
**Production-ready** — enterprise-grade stability, zero fluff, maximum efficiency
|
|
46
|
+
**Multi-language** — Python, JavaScript/WebAssembly, C/C++, C#
|
|
47
|
+
**60+ chart types** — 2D, 3D, maps, statistical plots, all GPU-accelerated
|
|
48
|
+
**ML preprocessing & metrics** — StandardScaler with Welford `partial_fit` (online learning), Pipeline with `score`/`predict_proba`/`decision_function`, OneHotEncoder/OrdinalEncoder with incremental category union
|
|
49
|
+
**WebAssembly** — npm package `@seraplot/wasm` for browser visualization & ML inference
|
|
50
|
+
**Streaming data** — online scalers and encoders for incremental model training
|
|
52
51
|
|
|
53
52
|
---
|
|
54
|
-
|
|
55
|
-
|
|
53
|
+
|
|
54
|
+
## Installation
|
|
55
|
+
|
|
56
|
+
**Python** (PyPI — wheel for CPython 3.11+)
|
|
56
57
|
|
|
57
58
|
```bash
|
|
58
59
|
pip install seraplot
|
|
59
60
|
```
|
|
60
61
|
|
|
61
|
-
|
|
62
|
-
|
|
62
|
+
Alternative package managers:
|
|
63
63
|
```bash
|
|
64
64
|
conda install -c conda-forge seraplot
|
|
65
|
+
uv pip install seraplot
|
|
65
66
|
```
|
|
67
|
+
|
|
68
|
+
**JavaScript/WebAssembly** (npm)
|
|
69
|
+
|
|
66
70
|
```bash
|
|
67
|
-
|
|
71
|
+
npm install seraplot
|
|
68
72
|
```
|
|
73
|
+
|
|
69
74
|
---
|
|
70
75
|
|
|
71
76
|
### Gallery — Chart Types
|
|
@@ -91,20 +96,4 @@ uv pip install seraplot
|
|
|
91
96
|
| 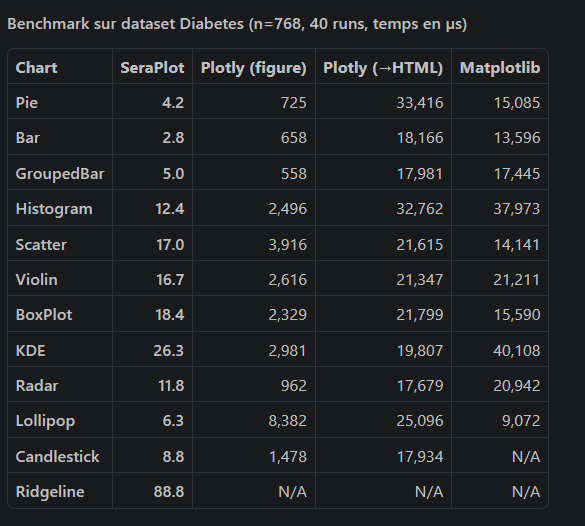 | 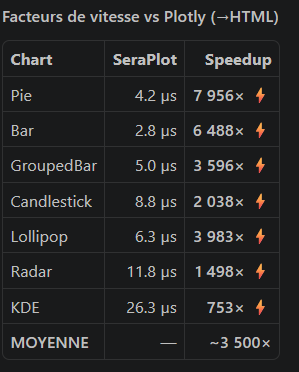 | 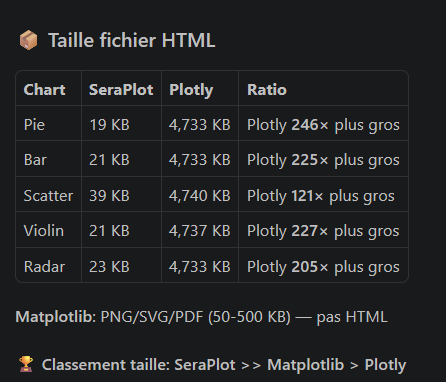 |
|
|
92
97
|
|
|
93
98
|
**SeraPlot outperforms Plotly and Matplotlib by 100–8000× on chart generation speed.**
|
|
94
|
-
|
|
95
|
-
---
|
|
96
|
-
|
|
97
|
-
### Quick Start
|
|
98
|
-
|
|
99
|
-
```python
|
|
100
|
-
from seraplot import build_bar_chart
|
|
101
|
-
|
|
102
|
-
# Create a simple bar chart
|
|
103
|
-
build_bar_chart(
|
|
104
|
-
'Sales by Region',
|
|
105
|
-
['North', 'South', 'East', 'West'],
|
|
106
|
-
[120, 95, 150, 110],
|
|
107
|
-
width=900, height=500
|
|
108
|
-
)
|
|
109
|
-
```
|
|
110
99
|
|
|
@@ -1,39 +1,44 @@
|
|
|
1
|
-
SeraPlot
|
|
1
|
+
SeraPlot — High-Performance Data Visualization & ML Framework
|
|
2
2
|
|
|
3
|
-
SeraPlot is a framework
|
|
4
|
-
alternative to Plotly, designed specifically for data visualization. This library is distributed
|
|
5
|
-
across multiple programming languages (Python, C#, C++, JavaScript), regularly maintained and
|
|
6
|
-
updated, offering superior speed and significantly lower memory consumption compared to competitors.
|
|
3
|
+
**SeraPlot v2.4.0+** is a production-grade framework written in **Rust**, delivering blazing-fast interactive charts and built-in machine learning pipelines. Designed as a modern alternative to Plotly + scikit-learn, it combines visualization and ML preprocessing in a single Rust binary — no dependencies, no bloat.
|
|
7
4
|
|
|
8
|
-
|
|
5
|
+
📖 **Documentation:** https://feur25.github.io/seraplot/introduction.html
|
|
9
6
|
|
|
10
7
|
---
|
|
11
8
|
|
|
12
|
-
|
|
9
|
+
## Why Choose SeraPlot?
|
|
13
10
|
|
|
14
|
-
|
|
15
|
-
|
|
16
|
-
|
|
17
|
-
|
|
18
|
-
|
|
19
|
-
|
|
11
|
+
**100–8000× faster** than Plotly & Matplotlib on chart generation
|
|
12
|
+
**Minimal memory footprint** — runs on edge devices, embedded systems, low-power servers
|
|
13
|
+
**Production-ready** — enterprise-grade stability, zero fluff, maximum efficiency
|
|
14
|
+
**Multi-language** — Python, JavaScript/WebAssembly, C/C++, C#
|
|
15
|
+
**60+ chart types** — 2D, 3D, maps, statistical plots, all GPU-accelerated
|
|
16
|
+
**ML preprocessing & metrics** — StandardScaler with Welford `partial_fit` (online learning), Pipeline with `score`/`predict_proba`/`decision_function`, OneHotEncoder/OrdinalEncoder with incremental category union
|
|
17
|
+
**WebAssembly** — npm package `@seraplot/wasm` for browser visualization & ML inference
|
|
18
|
+
**Streaming data** — online scalers and encoders for incremental model training
|
|
20
19
|
|
|
21
20
|
---
|
|
22
|
-
|
|
23
|
-
|
|
21
|
+
|
|
22
|
+
## Installation
|
|
23
|
+
|
|
24
|
+
**Python** (PyPI — wheel for CPython 3.11+)
|
|
24
25
|
|
|
25
26
|
```bash
|
|
26
27
|
pip install seraplot
|
|
27
28
|
```
|
|
28
29
|
|
|
29
|
-
|
|
30
|
-
|
|
30
|
+
Alternative package managers:
|
|
31
31
|
```bash
|
|
32
32
|
conda install -c conda-forge seraplot
|
|
33
|
+
uv pip install seraplot
|
|
33
34
|
```
|
|
35
|
+
|
|
36
|
+
**JavaScript/WebAssembly** (npm)
|
|
37
|
+
|
|
34
38
|
```bash
|
|
35
|
-
|
|
39
|
+
npm install seraplot
|
|
36
40
|
```
|
|
41
|
+
|
|
37
42
|
---
|
|
38
43
|
|
|
39
44
|
### Gallery — Chart Types
|
|
@@ -59,19 +64,3 @@ uv pip install seraplot
|
|
|
59
64
|
|  |  |  |
|
|
60
65
|
|
|
61
66
|
**SeraPlot outperforms Plotly and Matplotlib by 100–8000× on chart generation speed.**
|
|
62
|
-
|
|
63
|
-
---
|
|
64
|
-
|
|
65
|
-
### Quick Start
|
|
66
|
-
|
|
67
|
-
```python
|
|
68
|
-
from seraplot import build_bar_chart
|
|
69
|
-
|
|
70
|
-
# Create a simple bar chart
|
|
71
|
-
build_bar_chart(
|
|
72
|
-
'Sales by Region',
|
|
73
|
-
['North', 'South', 'East', 'West'],
|
|
74
|
-
[120, 95, 150, 110],
|
|
75
|
-
width=900, height=500
|
|
76
|
-
)
|
|
77
|
-
```
|
|
Binary file
|
|
Binary file
|
|
@@ -319,9 +319,12 @@ pub fn build_kmeans_chart(input: &str) -> String {
|
|
|
319
319
|
pub fn build_scatter_chart(input: &str) -> String {
|
|
320
320
|
let (title_s, a, o) = parse_all(input);
|
|
321
321
|
let title = title_s.as_str();
|
|
322
|
-
let
|
|
323
|
-
let
|
|
324
|
-
|
|
322
|
+
let raw_labels = a.labels.unwrap_or_default();
|
|
323
|
+
let x: Vec<f64> = a.x.unwrap_or_else(|| {
|
|
324
|
+
raw_labels.iter().enumerate().map(|(i, s)| s.parse::<f64>().unwrap_or(i as f64)).collect()
|
|
325
|
+
});
|
|
326
|
+
let y: Vec<f64> = a.y.unwrap_or_else(|| a.values.unwrap_or_default());
|
|
327
|
+
let lbls = raw_labels;
|
|
325
328
|
let sz = a.sizes.unwrap_or_default();
|
|
326
329
|
let cgs = o.color_groups.clone().unwrap_or_default();
|
|
327
330
|
let hover = o.hj();
|
|
@@ -591,7 +594,8 @@ pub fn build_area_chart(input: &str) -> String {
|
|
|
591
594
|
let (title_s, a, o) = parse_all(input);
|
|
592
595
|
let title = title_s.as_str();
|
|
593
596
|
let x_labels = a.x_labels.or(a.labels).unwrap_or_default();
|
|
594
|
-
let
|
|
597
|
+
let values_fallback = a.values;
|
|
598
|
+
let series_flat = a.series.unwrap_or_else(|| values_fallback.map(|v| vec![v]).unwrap_or_default());
|
|
595
599
|
use crate::plot::statistical::{AreaConfig, render_area_html};
|
|
596
600
|
let hover = o.hj();
|
|
597
601
|
let sn = o.series_names.clone().unwrap_or_default();
|
|
@@ -1450,6 +1454,372 @@ pub fn ml_kmeans_fit_predict(input: &str) -> String {
|
|
|
1450
1454
|
serde_json::to_string(&labels).unwrap_or_default(), inertia)
|
|
1451
1455
|
}
|
|
1452
1456
|
|
|
1457
|
+
pub fn ml_metric_score(input: &str) -> String {
|
|
1458
|
+
use crate::ml::metrics::*;
|
|
1459
|
+
#[derive(Deserialize, Default)]
|
|
1460
|
+
struct I {
|
|
1461
|
+
name: Option<String>,
|
|
1462
|
+
y_true: Option<Vec<f64>>,
|
|
1463
|
+
y_pred: Option<Vec<f64>>,
|
|
1464
|
+
y_score: Option<Vec<f64>>,
|
|
1465
|
+
labels: Option<Vec<i32>>,
|
|
1466
|
+
labels_true: Option<Vec<i32>>,
|
|
1467
|
+
labels_pred: Option<Vec<i32>>,
|
|
1468
|
+
x: Option<Vec<Vec<f64>>>,
|
|
1469
|
+
average: Option<String>,
|
|
1470
|
+
pos_label: Option<i32>,
|
|
1471
|
+
beta: Option<f64>,
|
|
1472
|
+
alpha: Option<f64>,
|
|
1473
|
+
eps: Option<f64>,
|
|
1474
|
+
}
|
|
1475
|
+
let i: I = serde_json::from_str(input).unwrap_or_default();
|
|
1476
|
+
let name = i.name.unwrap_or_default();
|
|
1477
|
+
let to_i32 = |v: &[f64]| v.iter().map(|x| *x as i32).collect::<Vec<i32>>();
|
|
1478
|
+
let yt_f = i.y_true.clone().unwrap_or_default();
|
|
1479
|
+
let yp_f = i.y_pred.clone().unwrap_or_default();
|
|
1480
|
+
let ys_f = i.y_score.clone().unwrap_or_default();
|
|
1481
|
+
let yt_i = to_i32(&yt_f);
|
|
1482
|
+
let yp_i = to_i32(&yp_f);
|
|
1483
|
+
let pos = i.pos_label.unwrap_or(1);
|
|
1484
|
+
let avg = match i.average.as_deref().unwrap_or("binary") {
|
|
1485
|
+
"macro" => Average::Macro,
|
|
1486
|
+
"weighted" => Average::Weighted,
|
|
1487
|
+
_ => Average::Binary(pos),
|
|
1488
|
+
};
|
|
1489
|
+
let value: f64 = match name.as_str() {
|
|
1490
|
+
"accuracy_score" => accuracy_score(&yt_i, &yp_i),
|
|
1491
|
+
"balanced_accuracy_score" => balanced_accuracy_score(&yt_i, &yp_i),
|
|
1492
|
+
"precision_score" => precision_score(&yt_i, &yp_i, avg),
|
|
1493
|
+
"recall_score" => recall_score(&yt_i, &yp_i, avg),
|
|
1494
|
+
"f1_score" => f1_score(&yt_i, &yp_i, avg),
|
|
1495
|
+
"fbeta_score" => fbeta_score(&yt_i, &yp_i, i.beta.unwrap_or(1.0), avg),
|
|
1496
|
+
"jaccard_score" => jaccard_score(&yt_i, &yp_i, pos),
|
|
1497
|
+
"matthews_corrcoef" => matthews_corrcoef(&yt_i, &yp_i),
|
|
1498
|
+
"cohen_kappa_score" => cohen_kappa_score(&yt_i, &yp_i),
|
|
1499
|
+
"hamming_loss" => hamming_loss(&yt_i, &yp_i),
|
|
1500
|
+
"zero_one_loss" => zero_one_loss(&yt_i, &yp_i),
|
|
1501
|
+
"binary_log_loss" => binary_log_loss(&yt_i, &yp_f, i.eps.unwrap_or(1e-15)),
|
|
1502
|
+
"brier_score_loss" => brier_score_loss(&yt_i, &yp_f),
|
|
1503
|
+
"hinge_loss" => hinge_loss(&yt_i, &yp_f),
|
|
1504
|
+
"roc_auc_score" => roc_auc_score(&yt_i, &ys_f, pos),
|
|
1505
|
+
"average_precision_score" => average_precision_score(&yt_i, &ys_f, pos),
|
|
1506
|
+
"mean_squared_error" => mean_squared_error(&yt_f, &yp_f),
|
|
1507
|
+
"root_mean_squared_error" => root_mean_squared_error(&yt_f, &yp_f),
|
|
1508
|
+
"mean_absolute_error" => mean_absolute_error(&yt_f, &yp_f),
|
|
1509
|
+
"median_absolute_error" => median_absolute_error(&yt_f, &yp_f),
|
|
1510
|
+
"r2_score" => r2_score(&yt_f, &yp_f),
|
|
1511
|
+
"explained_variance_score" => explained_variance_score(&yt_f, &yp_f),
|
|
1512
|
+
"max_error" => max_error(&yt_f, &yp_f),
|
|
1513
|
+
"mean_absolute_percentage_error" => mean_absolute_percentage_error(&yt_f, &yp_f),
|
|
1514
|
+
"mean_squared_log_error" => mean_squared_log_error(&yt_f, &yp_f),
|
|
1515
|
+
"root_mean_squared_log_error" => root_mean_squared_log_error(&yt_f, &yp_f),
|
|
1516
|
+
"mean_pinball_loss" => mean_pinball_loss(&yt_f, &yp_f, i.alpha.unwrap_or(0.5)),
|
|
1517
|
+
"d2_absolute_error_score" => d2_absolute_error_score(&yt_f, &yp_f),
|
|
1518
|
+
"silhouette_score" | "davies_bouldin_score" | "calinski_harabasz_score" => {
|
|
1519
|
+
let rows = i.x.clone().unwrap_or_default();
|
|
1520
|
+
let n = rows.len();
|
|
1521
|
+
let p = if n > 0 { rows[0].len() } else { 0 };
|
|
1522
|
+
let mut flat = Vec::with_capacity(n * p);
|
|
1523
|
+
for r in &rows { flat.extend_from_slice(&r[..p.min(r.len())]); if r.len() < p { flat.extend(std::iter::repeat(0.0).take(p - r.len())); } }
|
|
1524
|
+
let labs = i.labels.clone().unwrap_or_default();
|
|
1525
|
+
match name.as_str() {
|
|
1526
|
+
"silhouette_score" => silhouette_score(&flat, &labs, n, p),
|
|
1527
|
+
"davies_bouldin_score" => davies_bouldin_score(&flat, &labs, n, p),
|
|
1528
|
+
_ => calinski_harabasz_score(&flat, &labs, n, p),
|
|
1529
|
+
}
|
|
1530
|
+
}
|
|
1531
|
+
"adjusted_rand_score" | "normalized_mutual_info_score" | "fowlkes_mallows_score"
|
|
1532
|
+
| "homogeneity_score" | "completeness_score" | "v_measure_score" => {
|
|
1533
|
+
let lt = i.labels_true.clone().unwrap_or_default();
|
|
1534
|
+
let lp = i.labels_pred.clone().unwrap_or_default();
|
|
1535
|
+
match name.as_str() {
|
|
1536
|
+
"adjusted_rand_score" => adjusted_rand_score(<, &lp),
|
|
1537
|
+
"normalized_mutual_info_score" => normalized_mutual_info_score(<, &lp),
|
|
1538
|
+
"fowlkes_mallows_score" => fowlkes_mallows_score(<, &lp),
|
|
1539
|
+
"homogeneity_score" => homogeneity_score(<, &lp),
|
|
1540
|
+
"completeness_score" => completeness_score(<, &lp),
|
|
1541
|
+
_ => v_measure_score(<, &lp),
|
|
1542
|
+
}
|
|
1543
|
+
}
|
|
1544
|
+
_ => f64::NAN,
|
|
1545
|
+
};
|
|
1546
|
+
if value.is_nan() { format!("{{\"error\":\"unknown metric: {}\"}}", name) }
|
|
1547
|
+
else { format!("{{\"value\":{}}}", value) }
|
|
1548
|
+
}
|
|
1549
|
+
|
|
1550
|
+
pub fn ml_metric_curve(input: &str) -> String {
|
|
1551
|
+
use crate::ml::metrics::*;
|
|
1552
|
+
#[derive(Deserialize, Default)]
|
|
1553
|
+
struct I {
|
|
1554
|
+
name: Option<String>,
|
|
1555
|
+
y_true: Option<Vec<f64>>,
|
|
1556
|
+
y_score: Option<Vec<f64>>,
|
|
1557
|
+
pos_label: Option<i32>,
|
|
1558
|
+
}
|
|
1559
|
+
let i: I = serde_json::from_str(input).unwrap_or_default();
|
|
1560
|
+
let yt: Vec<i32> = i.y_true.unwrap_or_default().iter().map(|v| *v as i32).collect();
|
|
1561
|
+
let ys = i.y_score.unwrap_or_default();
|
|
1562
|
+
let pos = i.pos_label.unwrap_or(1);
|
|
1563
|
+
match i.name.as_deref().unwrap_or("") {
|
|
1564
|
+
"roc_curve" => {
|
|
1565
|
+
let (a, b, c) = roc_curve(&yt, &ys, pos);
|
|
1566
|
+
format!("{{\"fpr\":{},\"tpr\":{},\"thresholds\":{}}}",
|
|
1567
|
+
serde_json::to_string(&a).unwrap_or_default(),
|
|
1568
|
+
serde_json::to_string(&b).unwrap_or_default(),
|
|
1569
|
+
serde_json::to_string(&c).unwrap_or_default())
|
|
1570
|
+
}
|
|
1571
|
+
"precision_recall_curve" => {
|
|
1572
|
+
let (a, b, c) = precision_recall_curve(&yt, &ys, pos);
|
|
1573
|
+
format!("{{\"precision\":{},\"recall\":{},\"thresholds\":{}}}",
|
|
1574
|
+
serde_json::to_string(&a).unwrap_or_default(),
|
|
1575
|
+
serde_json::to_string(&b).unwrap_or_default(),
|
|
1576
|
+
serde_json::to_string(&c).unwrap_or_default())
|
|
1577
|
+
}
|
|
1578
|
+
n => format!("{{\"error\":\"unknown curve: {}\"}}", n),
|
|
1579
|
+
}
|
|
1580
|
+
}
|
|
1581
|
+
|
|
1582
|
+
pub fn ml_fit_transform(input: &str) -> String {
|
|
1583
|
+
use crate::ml::preprocessing::transformers::*;
|
|
1584
|
+
use crate::ml::preprocessing::scalers::*;
|
|
1585
|
+
#[derive(Deserialize, Default)]
|
|
1586
|
+
struct I {
|
|
1587
|
+
name: Option<String>,
|
|
1588
|
+
data: Option<Vec<Vec<f64>>>,
|
|
1589
|
+
strategy: Option<String>,
|
|
1590
|
+
fill_value: Option<f64>,
|
|
1591
|
+
degree: Option<usize>,
|
|
1592
|
+
interaction_only: Option<bool>,
|
|
1593
|
+
include_bias: Option<bool>,
|
|
1594
|
+
n_bins: Option<usize>,
|
|
1595
|
+
method: Option<String>,
|
|
1596
|
+
n_quantiles: Option<usize>,
|
|
1597
|
+
output_distribution: Option<String>,
|
|
1598
|
+
}
|
|
1599
|
+
let i: I = serde_json::from_str(input).unwrap_or_default();
|
|
1600
|
+
let rows = i.data.unwrap_or_default();
|
|
1601
|
+
let n = rows.len();
|
|
1602
|
+
let p = if n > 0 { rows[0].len() } else { 0 };
|
|
1603
|
+
let mut flat = Vec::with_capacity(n * p);
|
|
1604
|
+
for r in &rows {
|
|
1605
|
+
flat.extend_from_slice(&r[..p.min(r.len())]);
|
|
1606
|
+
if r.len() < p { flat.extend(std::iter::repeat(0.0).take(p - r.len())); }
|
|
1607
|
+
}
|
|
1608
|
+
let (out, cols, extra): (Vec<f64>, usize, String) = match i.name.as_deref().unwrap_or("") {
|
|
1609
|
+
"SimpleImputer" => {
|
|
1610
|
+
let mut t = SimpleImputer::new(i.strategy.as_deref().unwrap_or("mean"), i.fill_value.unwrap_or(0.0));
|
|
1611
|
+
t.fit(&flat, n, p);
|
|
1612
|
+
let o = t.transform(&flat, n, p);
|
|
1613
|
+
(o, p, format!(",\"statistics\":{}", serde_json::to_string(&t.statistics).unwrap_or_default()))
|
|
1614
|
+
}
|
|
1615
|
+
"PolynomialFeatures" => {
|
|
1616
|
+
let mut t = PolynomialFeatures::new(i.degree.unwrap_or(2), i.interaction_only.unwrap_or(false), i.include_bias.unwrap_or(true));
|
|
1617
|
+
t.fit(&flat, n, p);
|
|
1618
|
+
let o = t.transform(&flat, n, p);
|
|
1619
|
+
let nf = t.n_output_features();
|
|
1620
|
+
(o, nf, format!(",\"n_features_out\":{}", nf))
|
|
1621
|
+
}
|
|
1622
|
+
"KBinsDiscretizer" => {
|
|
1623
|
+
let mut t = KBinsDiscretizer::new(i.n_bins.unwrap_or(5), i.strategy.as_deref().unwrap_or("quantile"));
|
|
1624
|
+
t.fit(&flat, n, p);
|
|
1625
|
+
let o = t.transform(&flat, n, p);
|
|
1626
|
+
(o, p, String::new())
|
|
1627
|
+
}
|
|
1628
|
+
"PowerTransformer" => {
|
|
1629
|
+
let mut t = PowerTransformer::new(i.method.as_deref().unwrap_or("yeo-johnson"));
|
|
1630
|
+
t.fit(&flat, n, p);
|
|
1631
|
+
let o = t.transform(&flat, n, p);
|
|
1632
|
+
(o, p, format!(",\"lambdas\":{}", serde_json::to_string(&t.lambdas).unwrap_or_default()))
|
|
1633
|
+
}
|
|
1634
|
+
"QuantileTransformer" => {
|
|
1635
|
+
let mut t = QuantileTransformer::new(i.n_quantiles.unwrap_or(1000), i.output_distribution.as_deref().unwrap_or("uniform"));
|
|
1636
|
+
t.fit(&flat, n, p);
|
|
1637
|
+
let o = t.transform(&flat, n, p);
|
|
1638
|
+
(o, p, String::new())
|
|
1639
|
+
}
|
|
1640
|
+
"StandardScaler" => {
|
|
1641
|
+
let mut t = StandardScaler::new(true, true);
|
|
1642
|
+
t.fit(&flat, n, p);
|
|
1643
|
+
let o = t.transform(&flat, n, p);
|
|
1644
|
+
(o, p, String::new())
|
|
1645
|
+
}
|
|
1646
|
+
"MinMaxScaler" => {
|
|
1647
|
+
let mut t = MinMaxScaler::new((0.0, 1.0));
|
|
1648
|
+
t.fit(&flat, n, p);
|
|
1649
|
+
let o = t.transform(&flat, n, p);
|
|
1650
|
+
(o, p, String::new())
|
|
1651
|
+
}
|
|
1652
|
+
"RobustScaler" => {
|
|
1653
|
+
let mut t = RobustScaler::new(true, true);
|
|
1654
|
+
t.fit(&flat, n, p);
|
|
1655
|
+
let o = t.transform(&flat, n, p);
|
|
1656
|
+
(o, p, String::new())
|
|
1657
|
+
}
|
|
1658
|
+
"MaxAbsScaler" => {
|
|
1659
|
+
let mut t = MaxAbsScaler::new();
|
|
1660
|
+
t.fit(&flat, n, p);
|
|
1661
|
+
let o = t.transform(&flat, n, p);
|
|
1662
|
+
(o, p, String::new())
|
|
1663
|
+
}
|
|
1664
|
+
n_ => return format!("{{\"error\":\"unknown transformer: {}\"}}", n_),
|
|
1665
|
+
};
|
|
1666
|
+
let mut data: Vec<Vec<f64>> = Vec::with_capacity(n);
|
|
1667
|
+
for r in 0..n {
|
|
1668
|
+
data.push(out[r * cols..(r + 1) * cols].to_vec());
|
|
1669
|
+
}
|
|
1670
|
+
format!("{{\"data\":{},\"n\":{},\"cols\":{}{}}}",
|
|
1671
|
+
serde_json::to_string(&data).unwrap_or_default(), n, cols, extra)
|
|
1672
|
+
}
|
|
1673
|
+
|
|
1674
|
+
pub fn ml_kfold_split(input: &str) -> String {
|
|
1675
|
+
use crate::ml::model_selection::split::*;
|
|
1676
|
+
#[derive(Deserialize, Default)]
|
|
1677
|
+
struct I {
|
|
1678
|
+
kind: Option<String>,
|
|
1679
|
+
n: Option<usize>,
|
|
1680
|
+
k: Option<usize>,
|
|
1681
|
+
seed: Option<u64>,
|
|
1682
|
+
y: Option<Vec<i32>>,
|
|
1683
|
+
groups: Option<Vec<i32>>,
|
|
1684
|
+
}
|
|
1685
|
+
let i: I = serde_json::from_str(input).unwrap_or_default();
|
|
1686
|
+
let kind = i.kind.unwrap_or_else(|| "kfold".to_string());
|
|
1687
|
+
let k = i.k.unwrap_or(5);
|
|
1688
|
+
let seed = i.seed.unwrap_or(0);
|
|
1689
|
+
let folds = match kind.as_str() {
|
|
1690
|
+
"stratified" => stratified_kfold_indices(&i.y.unwrap_or_default(), k, seed),
|
|
1691
|
+
"group" => group_kfold_indices(&i.groups.unwrap_or_default(), k),
|
|
1692
|
+
_ => kfold_indices(i.n.unwrap_or(0), k, seed),
|
|
1693
|
+
};
|
|
1694
|
+
let payload: Vec<serde_json::Value> = folds.into_iter().map(|(tr, te)| {
|
|
1695
|
+
serde_json::json!({"train": tr, "test": te})
|
|
1696
|
+
}).collect();
|
|
1697
|
+
serde_json::to_string(&payload).unwrap_or_else(|_| "[]".to_string())
|
|
1698
|
+
}
|
|
1699
|
+
|
|
1700
|
+
pub fn ml_isolation_forest(input: &str) -> String {
|
|
1701
|
+
use crate::ml::anomaly::isolation_forest::IsolationForest;
|
|
1702
|
+
#[derive(Deserialize, Default)]
|
|
1703
|
+
struct I {
|
|
1704
|
+
data: Option<Vec<Vec<f64>>>,
|
|
1705
|
+
x_test: Option<Vec<Vec<f64>>>,
|
|
1706
|
+
n_estimators: Option<usize>,
|
|
1707
|
+
max_samples: Option<usize>,
|
|
1708
|
+
contamination: Option<f64>,
|
|
1709
|
+
seed: Option<u64>,
|
|
1710
|
+
}
|
|
1711
|
+
let i: I = serde_json::from_str(input).unwrap_or_default();
|
|
1712
|
+
let rows = i.data.unwrap_or_default();
|
|
1713
|
+
let n = rows.len();
|
|
1714
|
+
if n == 0 { return "{\"labels\":[],\"scores\":[],\"threshold\":0.0}".to_string(); }
|
|
1715
|
+
let p = rows[0].len();
|
|
1716
|
+
let mut flat = Vec::with_capacity(n * p);
|
|
1717
|
+
for r in &rows { flat.extend_from_slice(&r[..p.min(r.len())]); if r.len() < p { flat.extend(std::iter::repeat(0.0).take(p - r.len())); } }
|
|
1718
|
+
let mut model = IsolationForest::new(
|
|
1719
|
+
i.n_estimators.unwrap_or(100),
|
|
1720
|
+
i.max_samples.unwrap_or(256),
|
|
1721
|
+
i.contamination.unwrap_or(0.1),
|
|
1722
|
+
i.seed.unwrap_or(42),
|
|
1723
|
+
);
|
|
1724
|
+
model.fit(&flat, n, p);
|
|
1725
|
+
let labels = model.predict(&flat, n, p);
|
|
1726
|
+
let scores = model.score_samples(&flat, n, p);
|
|
1727
|
+
let test_payload = if let Some(rows_t) = i.x_test {
|
|
1728
|
+
let nt = rows_t.len();
|
|
1729
|
+
if nt == 0 {
|
|
1730
|
+
String::new()
|
|
1731
|
+
} else {
|
|
1732
|
+
let mut flat_t = Vec::with_capacity(nt * p);
|
|
1733
|
+
for r in &rows_t { flat_t.extend_from_slice(&r[..p.min(r.len())]); if r.len() < p { flat_t.extend(std::iter::repeat(0.0).take(p - r.len())); } }
|
|
1734
|
+
let lt = model.predict(&flat_t, nt, p);
|
|
1735
|
+
let st = model.score_samples(&flat_t, nt, p);
|
|
1736
|
+
format!(",\"test_labels\":{},\"test_scores\":{}",
|
|
1737
|
+
serde_json::to_string(<).unwrap_or_default(),
|
|
1738
|
+
serde_json::to_string(&st).unwrap_or_default())
|
|
1739
|
+
}
|
|
1740
|
+
} else { String::new() };
|
|
1741
|
+
format!("{{\"labels\":{},\"scores\":{},\"threshold\":{}{}}}",
|
|
1742
|
+
serde_json::to_string(&labels).unwrap_or_default(),
|
|
1743
|
+
serde_json::to_string(&scores).unwrap_or_default(),
|
|
1744
|
+
model.threshold_,
|
|
1745
|
+
test_payload)
|
|
1746
|
+
}
|
|
1747
|
+
|
|
1748
|
+
pub fn ml_permutation_importance(input: &str) -> String {
|
|
1749
|
+
use crate::ml::model_selection::permutation::*;
|
|
1750
|
+
#[derive(Deserialize, Default)]
|
|
1751
|
+
struct I {
|
|
1752
|
+
data: Option<Vec<Vec<f64>>>,
|
|
1753
|
+
y: Option<Vec<f64>>,
|
|
1754
|
+
baseline_pred: Option<Vec<f64>>,
|
|
1755
|
+
perm_preds: Option<Vec<Vec<Vec<f64>>>>,
|
|
1756
|
+
task: Option<String>,
|
|
1757
|
+
n_repeats: Option<usize>,
|
|
1758
|
+
seed: Option<u64>,
|
|
1759
|
+
}
|
|
1760
|
+
let i: I = serde_json::from_str(input).unwrap_or_default();
|
|
1761
|
+
let rows = i.data.unwrap_or_default();
|
|
1762
|
+
let n = rows.len();
|
|
1763
|
+
if n == 0 { return "{\"importances_mean\":[],\"importances_std\":[]}".to_string(); }
|
|
1764
|
+
let p = rows[0].len();
|
|
1765
|
+
let mut flat = Vec::with_capacity(n * p);
|
|
1766
|
+
for r in &rows { flat.extend_from_slice(&r[..p.min(r.len())]); if r.len() < p { flat.extend(std::iter::repeat(0.0).take(p - r.len())); } }
|
|
1767
|
+
let yf = i.y.clone().unwrap_or_default();
|
|
1768
|
+
let task = i.task.unwrap_or_else(|| "regression".to_string());
|
|
1769
|
+
let n_repeats = i.n_repeats.unwrap_or(5);
|
|
1770
|
+
let seed = i.seed.unwrap_or(0);
|
|
1771
|
+
let perm_preds = i.perm_preds.unwrap_or_default();
|
|
1772
|
+
let baseline_pred = i.baseline_pred.unwrap_or_default();
|
|
1773
|
+
let _ = (n_repeats, seed, flat, n);
|
|
1774
|
+
if perm_preds.is_empty() || baseline_pred.is_empty() {
|
|
1775
|
+
return "{\"importances_mean\":[],\"importances_std\":[],\"error\":\"baseline_pred and perm_preds[p][n_repeats][n] are required\"}".to_string();
|
|
1776
|
+
}
|
|
1777
|
+
if task == "classification" {
|
|
1778
|
+
let yi: Vec<i32> = yf.iter().map(|v| *v as i32).collect();
|
|
1779
|
+
let baseline_i: Vec<i32> = baseline_pred.iter().map(|v| *v as i32).collect();
|
|
1780
|
+
let baseline = crate::ml::metrics::classification::accuracy_score(&yi, &baseline_i);
|
|
1781
|
+
let mut means = vec![0.0; p];
|
|
1782
|
+
let mut stds = vec![0.0; p];
|
|
1783
|
+
for j in 0..p.min(perm_preds.len()) {
|
|
1784
|
+
let reps = &perm_preds[j];
|
|
1785
|
+
let mut diffs = Vec::with_capacity(reps.len());
|
|
1786
|
+
for rep in reps {
|
|
1787
|
+
let pi: Vec<i32> = rep.iter().map(|v| *v as i32).collect();
|
|
1788
|
+
let s = crate::ml::metrics::classification::accuracy_score(&yi, &pi);
|
|
1789
|
+
diffs.push(baseline - s);
|
|
1790
|
+
}
|
|
1791
|
+
let m = diffs.iter().sum::<f64>() / diffs.len().max(1) as f64;
|
|
1792
|
+
let v = diffs.iter().map(|x| (x - m).powi(2)).sum::<f64>() / diffs.len().max(1) as f64;
|
|
1793
|
+
means[j] = m;
|
|
1794
|
+
stds[j] = v.sqrt();
|
|
1795
|
+
}
|
|
1796
|
+
format!("{{\"importances_mean\":{},\"importances_std\":{},\"baseline\":{}}}",
|
|
1797
|
+
serde_json::to_string(&means).unwrap_or_default(),
|
|
1798
|
+
serde_json::to_string(&stds).unwrap_or_default(),
|
|
1799
|
+
baseline)
|
|
1800
|
+
} else {
|
|
1801
|
+
let baseline = crate::ml::metrics::regression::r2_score(&yf, &baseline_pred);
|
|
1802
|
+
let mut means = vec![0.0; p];
|
|
1803
|
+
let mut stds = vec![0.0; p];
|
|
1804
|
+
for j in 0..p.min(perm_preds.len()) {
|
|
1805
|
+
let reps = &perm_preds[j];
|
|
1806
|
+
let mut diffs = Vec::with_capacity(reps.len());
|
|
1807
|
+
for rep in reps {
|
|
1808
|
+
let s = crate::ml::metrics::regression::r2_score(&yf, rep);
|
|
1809
|
+
diffs.push(baseline - s);
|
|
1810
|
+
}
|
|
1811
|
+
let m = diffs.iter().sum::<f64>() / diffs.len().max(1) as f64;
|
|
1812
|
+
let v = diffs.iter().map(|x| (x - m).powi(2)).sum::<f64>() / diffs.len().max(1) as f64;
|
|
1813
|
+
means[j] = m;
|
|
1814
|
+
stds[j] = v.sqrt();
|
|
1815
|
+
}
|
|
1816
|
+
format!("{{\"importances_mean\":{},\"importances_std\":{},\"baseline\":{}}}",
|
|
1817
|
+
serde_json::to_string(&means).unwrap_or_default(),
|
|
1818
|
+
serde_json::to_string(&stds).unwrap_or_default(),
|
|
1819
|
+
baseline)
|
|
1820
|
+
}
|
|
1821
|
+
}
|
|
1822
|
+
|
|
1453
1823
|
pub fn set_global_background(input: &str) -> String {
|
|
1454
1824
|
let color = input.trim().trim_matches('"');
|
|
1455
1825
|
set_global_bg(if color.is_empty() { None } else { Some(color.to_string()) });
|