scTimeBench 0.1.0__tar.gz
This diff represents the content of publicly available package versions that have been released to one of the supported registries. The information contained in this diff is provided for informational purposes only and reflects changes between package versions as they appear in their respective public registries.
- sctimebench-0.1.0/LICENSE +21 -0
- sctimebench-0.1.0/PKG-INFO +150 -0
- sctimebench-0.1.0/README.md +118 -0
- sctimebench-0.1.0/pyproject.toml +61 -0
- sctimebench-0.1.0/setup.cfg +4 -0
- sctimebench-0.1.0/src/scTimeBench/config.py +318 -0
- sctimebench-0.1.0/src/scTimeBench/database.py +675 -0
- sctimebench-0.1.0/src/scTimeBench/main.py +166 -0
- sctimebench-0.1.0/src/scTimeBench/method_utils/__init__.py +0 -0
- sctimebench-0.1.0/src/scTimeBench/method_utils/method_runner.py +240 -0
- sctimebench-0.1.0/src/scTimeBench/method_utils/ot_method_runner.py +247 -0
- sctimebench-0.1.0/src/scTimeBench/metrics/__init__.py +9 -0
- sctimebench-0.1.0/src/scTimeBench/metrics/base.py +622 -0
- sctimebench-0.1.0/src/scTimeBench/metrics/embeddings/__init__.py +0 -0
- sctimebench-0.1.0/src/scTimeBench/metrics/embeddings/aggregate/__init__.py +0 -0
- sctimebench-0.1.0/src/scTimeBench/metrics/embeddings/aggregate/ari.py +99 -0
- sctimebench-0.1.0/src/scTimeBench/metrics/embeddings/aggregate/base.py +15 -0
- sctimebench-0.1.0/src/scTimeBench/metrics/embeddings/base.py +49 -0
- sctimebench-0.1.0/src/scTimeBench/metrics/embeddings/trajectory/__init__.py +0 -0
- sctimebench-0.1.0/src/scTimeBench/metrics/embeddings/trajectory/base.py +173 -0
- sctimebench-0.1.0/src/scTimeBench/metrics/gex_prediction/__init__.py +13 -0
- sctimebench-0.1.0/src/scTimeBench/metrics/gex_prediction/base.py +96 -0
- sctimebench-0.1.0/src/scTimeBench/metrics/gex_prediction/ot_eval/__init__.py +11 -0
- sctimebench-0.1.0/src/scTimeBench/metrics/gex_prediction/ot_eval/base.py +199 -0
- sctimebench-0.1.0/src/scTimeBench/metrics/gex_prediction/ot_eval/energy.py +39 -0
- sctimebench-0.1.0/src/scTimeBench/metrics/gex_prediction/ot_eval/hausdorff.py +36 -0
- sctimebench-0.1.0/src/scTimeBench/metrics/gex_prediction/ot_eval/mmd.py +39 -0
- sctimebench-0.1.0/src/scTimeBench/metrics/gex_prediction/ot_eval/wass.py +42 -0
- sctimebench-0.1.0/src/scTimeBench/metrics/method_manager.py +80 -0
- sctimebench-0.1.0/src/scTimeBench/metrics/ontology_based/__init__.py +0 -0
- sctimebench-0.1.0/src/scTimeBench/metrics/ontology_based/base.py +45 -0
- sctimebench-0.1.0/src/scTimeBench/metrics/ontology_based/graph_sim/__init__.py +0 -0
- sctimebench-0.1.0/src/scTimeBench/metrics/ontology_based/graph_sim/average_shortest_path_diff.py +80 -0
- sctimebench-0.1.0/src/scTimeBench/metrics/ontology_based/graph_sim/base.py +382 -0
- sctimebench-0.1.0/src/scTimeBench/metrics/ontology_based/graph_sim/confusion_matrix.py +126 -0
- sctimebench-0.1.0/src/scTimeBench/metrics/ontology_based/graph_sim/ged.py +41 -0
- sctimebench-0.1.0/src/scTimeBench/metrics/ontology_based/graph_sim/graph_viz.py +278 -0
- sctimebench-0.1.0/src/scTimeBench/metrics/ontology_based/graph_sim/jaccard_similarity.py +34 -0
- sctimebench-0.1.0/src/scTimeBench/metrics/ontology_based/graph_sim/utils.py +59 -0
- sctimebench-0.1.0/src/scTimeBench/shared/__init__.py +0 -0
- sctimebench-0.1.0/src/scTimeBench/shared/constants.py +26 -0
- sctimebench-0.1.0/src/scTimeBench/shared/dataset/__init__.py +9 -0
- sctimebench-0.1.0/src/scTimeBench/shared/dataset/base.py +262 -0
- sctimebench-0.1.0/src/scTimeBench/shared/dataset/preprocessors/__init__.py +0 -0
- sctimebench-0.1.0/src/scTimeBench/shared/dataset/preprocessors/copy_train_test.py +24 -0
- sctimebench-0.1.0/src/scTimeBench/shared/dataset/preprocessors/lineage.py +70 -0
- sctimebench-0.1.0/src/scTimeBench/shared/dataset/preprocessors/log_norm.py +60 -0
- sctimebench-0.1.0/src/scTimeBench/shared/dataset/preprocessors/percentile_split.py +39 -0
- sctimebench-0.1.0/src/scTimeBench/shared/dataset/preprocessors/pseudotime.py +406 -0
- sctimebench-0.1.0/src/scTimeBench/shared/dataset/preprocessors/round_cells_to_timepoint.py +129 -0
- sctimebench-0.1.0/src/scTimeBench/shared/dataset/preprocessors/test_timepoint_selection.py +52 -0
- sctimebench-0.1.0/src/scTimeBench/shared/dataset/registry/__init__.py +19 -0
- sctimebench-0.1.0/src/scTimeBench/shared/dataset/registry/drosophila.py +31 -0
- sctimebench-0.1.0/src/scTimeBench/shared/dataset/registry/dummy.py +13 -0
- sctimebench-0.1.0/src/scTimeBench/shared/dataset/registry/garcia_alonso.py +32 -0
- sctimebench-0.1.0/src/scTimeBench/shared/dataset/registry/ma.py +29 -0
- sctimebench-0.1.0/src/scTimeBench/shared/dataset/registry/maolaniru.py +31 -0
- sctimebench-0.1.0/src/scTimeBench/shared/dataset/registry/mef.py +31 -0
- sctimebench-0.1.0/src/scTimeBench/shared/dataset/registry/olaniru.py +31 -0
- sctimebench-0.1.0/src/scTimeBench/shared/dataset/registry/suo.py +28 -0
- sctimebench-0.1.0/src/scTimeBench/shared/dataset/registry/zebrafish.py +31 -0
- sctimebench-0.1.0/src/scTimeBench/shared/helpers.py +118 -0
- sctimebench-0.1.0/src/scTimeBench/shared/utils.py +224 -0
- sctimebench-0.1.0/src/scTimeBench/trajectory_infer/__init__.py +9 -0
- sctimebench-0.1.0/src/scTimeBench/trajectory_infer/base.py +546 -0
- sctimebench-0.1.0/src/scTimeBench/trajectory_infer/classifier.py +199 -0
- sctimebench-0.1.0/src/scTimeBench/trajectory_infer/kNN.py +127 -0
- sctimebench-0.1.0/src/scTimeBench/trajectory_infer/ot.py +156 -0
- sctimebench-0.1.0/src/scTimeBench.egg-info/PKG-INFO +150 -0
- sctimebench-0.1.0/src/scTimeBench.egg-info/SOURCES.txt +73 -0
- sctimebench-0.1.0/src/scTimeBench.egg-info/dependency_links.txt +1 -0
- sctimebench-0.1.0/src/scTimeBench.egg-info/entry_points.txt +2 -0
- sctimebench-0.1.0/src/scTimeBench.egg-info/requires.txt +17 -0
- sctimebench-0.1.0/src/scTimeBench.egg-info/top_level.txt +1 -0
- sctimebench-0.1.0/test/test_suite_helpers.py +254 -0
|
@@ -0,0 +1,21 @@
|
|
|
1
|
+
MIT License
|
|
2
|
+
|
|
3
|
+
Copyright (c) 2026 Eric Haoran Huang and Adrien Osakwe
|
|
4
|
+
|
|
5
|
+
Permission is hereby granted, free of charge, to any person obtaining a copy
|
|
6
|
+
of this software and associated documentation files (the "Software"), to deal
|
|
7
|
+
in the Software without restriction, including without limitation the rights
|
|
8
|
+
to use, copy, modify, merge, publish, distribute, sublicense, and/or sell
|
|
9
|
+
copies of the Software, and to permit persons to whom the Software is
|
|
10
|
+
furnished to do so, subject to the following conditions:
|
|
11
|
+
|
|
12
|
+
The above copyright notice and this permission notice shall be included in all
|
|
13
|
+
copies or substantial portions of the Software.
|
|
14
|
+
|
|
15
|
+
THE SOFTWARE IS PROVIDED "AS IS", WITHOUT WARRANTY OF ANY KIND, EXPRESS OR
|
|
16
|
+
IMPLIED, INCLUDING BUT NOT LIMITED TO THE WARRANTIES OF MERCHANTABILITY,
|
|
17
|
+
FITNESS FOR A PARTICULAR PURPOSE AND NONINFRINGEMENT. IN NO EVENT SHALL THE
|
|
18
|
+
AUTHORS OR COPYRIGHT HOLDERS BE LIABLE FOR ANY CLAIM, DAMAGES OR OTHER
|
|
19
|
+
LIABILITY, WHETHER IN AN ACTION OF CONTRACT, TORT OR OTHERWISE, ARISING FROM,
|
|
20
|
+
OUT OF OR IN CONNECTION WITH THE SOFTWARE OR THE USE OR OTHER DEALINGS IN THE
|
|
21
|
+
SOFTWARE.
|
|
@@ -0,0 +1,150 @@
|
|
|
1
|
+
Metadata-Version: 2.4
|
|
2
|
+
Name: scTimeBench
|
|
3
|
+
Version: 0.1.0
|
|
4
|
+
Summary: A streamlined benchmarking platform for single-cell time-series analysis
|
|
5
|
+
Author-email: Eric Haoran Huang <eric.h.huang@mail.mcgill.ca>, Adrien Osakwe <adrien.osakwe@mail.mcgill.ca>, Yue Li <yueli@cs.mcgill.ca>
|
|
6
|
+
License-Expression: MIT
|
|
7
|
+
Project-URL: Homepage, https://github.com/li-lab-mcgill/scTimeBench
|
|
8
|
+
Project-URL: Issues, https://github.com/li-lab-mcgill/scTimeBench/issues
|
|
9
|
+
Classifier: Development Status :: 4 - Beta
|
|
10
|
+
Classifier: Intended Audience :: Science/Research
|
|
11
|
+
Classifier: Topic :: Scientific/Engineering :: Bio-Informatics
|
|
12
|
+
Classifier: Programming Language :: Python :: 3.10
|
|
13
|
+
Requires-Python: >=3.10
|
|
14
|
+
Description-Content-Type: text/markdown
|
|
15
|
+
License-File: LICENSE
|
|
16
|
+
Requires-Dist: PyYAML
|
|
17
|
+
Requires-Dist: scanpy
|
|
18
|
+
Requires-Dist: scikit-learn
|
|
19
|
+
Requires-Dist: fastparquet<2026.3.0
|
|
20
|
+
Provides-Extra: dev
|
|
21
|
+
Requires-Dist: pre-commit; extra == "dev"
|
|
22
|
+
Requires-Dist: pytest; extra == "dev"
|
|
23
|
+
Provides-Extra: benchmark
|
|
24
|
+
Requires-Dist: networkx; extra == "benchmark"
|
|
25
|
+
Requires-Dist: numpy; extra == "benchmark"
|
|
26
|
+
Requires-Dist: geomloss; extra == "benchmark"
|
|
27
|
+
Requires-Dist: pykeops; extra == "benchmark"
|
|
28
|
+
Requires-Dist: scanpy[leiden]; extra == "benchmark"
|
|
29
|
+
Requires-Dist: graphviz; extra == "benchmark"
|
|
30
|
+
Requires-Dist: celltypist; extra == "benchmark"
|
|
31
|
+
Dynamic: license-file
|
|
32
|
+
|
|
33
|
+
<!-- for this to work on Pypi, we need to point to the absolute path -->
|
|
34
|
+
<h1>
|
|
35
|
+
<img src="https://raw.githubusercontent.com/li-lab-mcgill/scTimeBench/refs/heads/main/assets/logo.png" alt="scTimeBench-Logo" height="150" align="absmiddle" /> scTimeBench
|
|
36
|
+
</h1>
|
|
37
|
+
|
|
38
|
+
[](https://www.python.org/downloads/release/python-31012/)
|
|
39
|
+
[](https://opensource.org/license/mit)
|
|
40
|
+
[](https://www.biorxiv.org/content/10.64898/2026.03.16.712069v1)
|
|
41
|
+
[](https://colab.research.google.com/drive/1J-yNXu_FcSnhrCwTDQKjWCBSHsmdbohJ?usp=sharing)
|
|
42
|
+
<!-- TODO: --> <!-- []() -->
|
|
43
|
+
|
|
44
|
+
|
|
45
|
+
|
|
46
|
+
|
|
47
|
+
## Table of Contents
|
|
48
|
+
- [Environment Setup](#environment-setup)
|
|
49
|
+
- [Suggested UV Installation](#suggested-uv-installation)
|
|
50
|
+
- [Standard Pip](#standard-pip)
|
|
51
|
+
- [Benchmark Architecture](#benchmark-architecture)
|
|
52
|
+
- [Detailed Layout of File Structure](#detailed-layout-of-file-structure)
|
|
53
|
+
- [Command-Line Interface Details](#command-line-interface-details)
|
|
54
|
+
- [Example Run](#example-run)
|
|
55
|
+
- [Contributing to scTimeBench](#contributing-to-sctimebench)
|
|
56
|
+
- [Citation](#citation)
|
|
57
|
+
|
|
58
|
+
## Environment Setup
|
|
59
|
+
scTimeBench was tested and supported using Python 3.10. If any other version that is 3.10+ does not work when using the benchmark, please submit an issue to this GitHub.
|
|
60
|
+
|
|
61
|
+
Important Note: this setup is needed twice. Once for the user to run the benchmark metrics, and the second time for the method itself needing to read from `scTimeBench.method_utils.method_runner` and other important shared constants. In short, this means we need to install scTimeBench into two separate virtual environments:
|
|
62
|
+
1. Your normal pip installation where you'll be running the benchmark from. This will require the extra "\[benchmark\]" group installation (pip) or the extra group installation `--extra benchmark` (uv).
|
|
63
|
+
2. For each method's virtual environment, you need to install the scTimeBench.
|
|
64
|
+
|
|
65
|
+
### Suggested UV Installation
|
|
66
|
+
Due to external dependencies and a more complex setup, we have decided to package everything under `uv` (see: https://github.com/astral-sh/uv). To start with, you need to get all the necessary extern dependencies, which can be done either by running:
|
|
67
|
+
```
|
|
68
|
+
git submodule update --init extern/
|
|
69
|
+
```
|
|
70
|
+
If you wish to benchmark across all methods, feel free to clone the submodules for all the methods as well with:
|
|
71
|
+
```
|
|
72
|
+
git submodule update --init
|
|
73
|
+
```
|
|
74
|
+
Then, install `uv` and run the following:
|
|
75
|
+
```
|
|
76
|
+
uv sync --extra benchmark
|
|
77
|
+
```
|
|
78
|
+
If you're using uv under a method's virtual environment, either the pip installation or the following will suffice:
|
|
79
|
+
```
|
|
80
|
+
uv sync
|
|
81
|
+
```
|
|
82
|
+
|
|
83
|
+
### Standard Pip
|
|
84
|
+
If the external dependencies such as pypsupertime or sceptic are not used (which they are not used by default), you can install using pip as follows:
|
|
85
|
+
```
|
|
86
|
+
pip install -e ".[benchmark]"
|
|
87
|
+
```
|
|
88
|
+
to run the benchmark. For your own method, simply install without the extra benchmarking requirements with
|
|
89
|
+
```
|
|
90
|
+
pip install -e .
|
|
91
|
+
```
|
|
92
|
+
There are extra dependencies that can be found under `pyproject.toml`.
|
|
93
|
+
|
|
94
|
+
## Benchmark Architecture
|
|
95
|
+
|
|
96
|
+
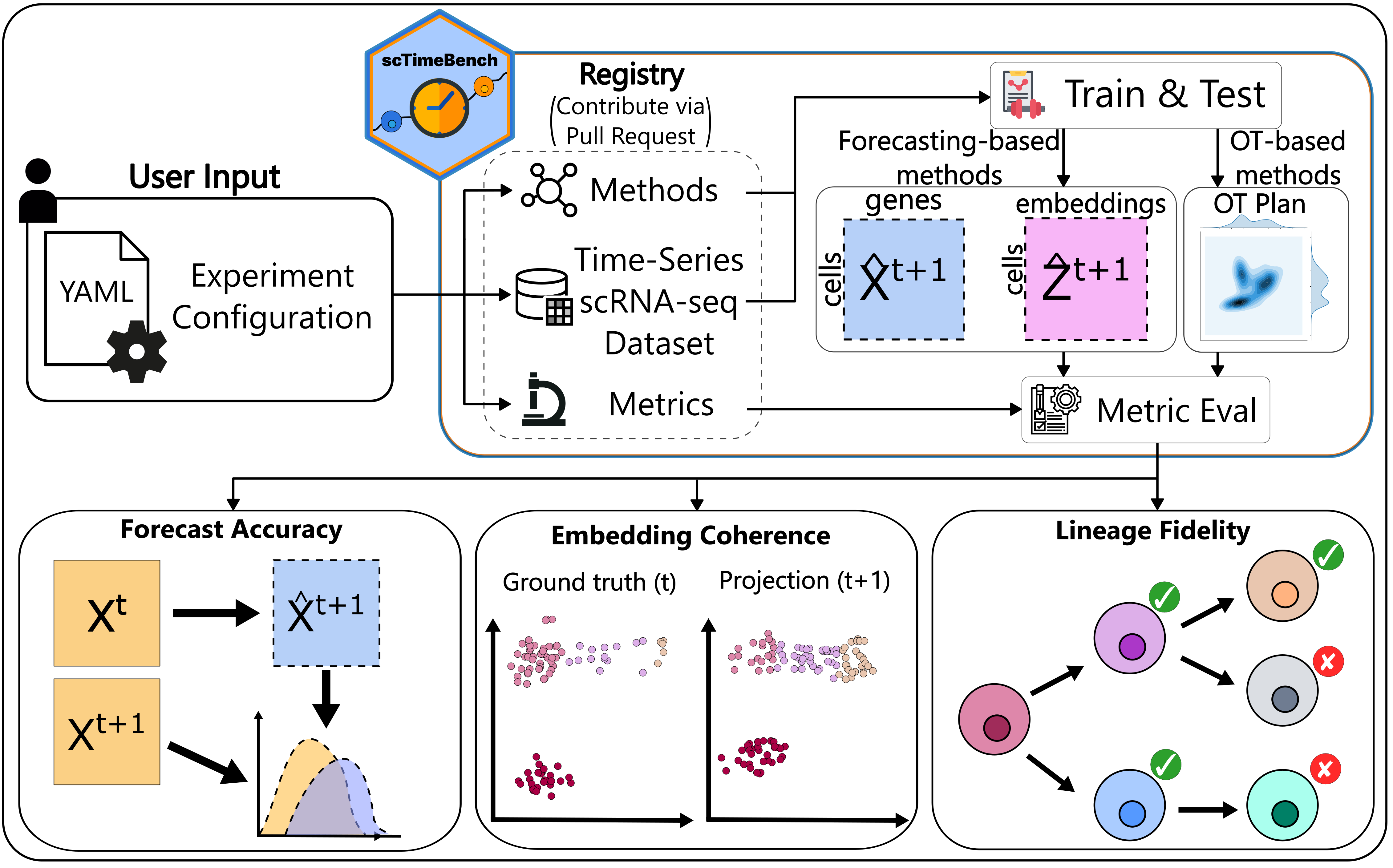
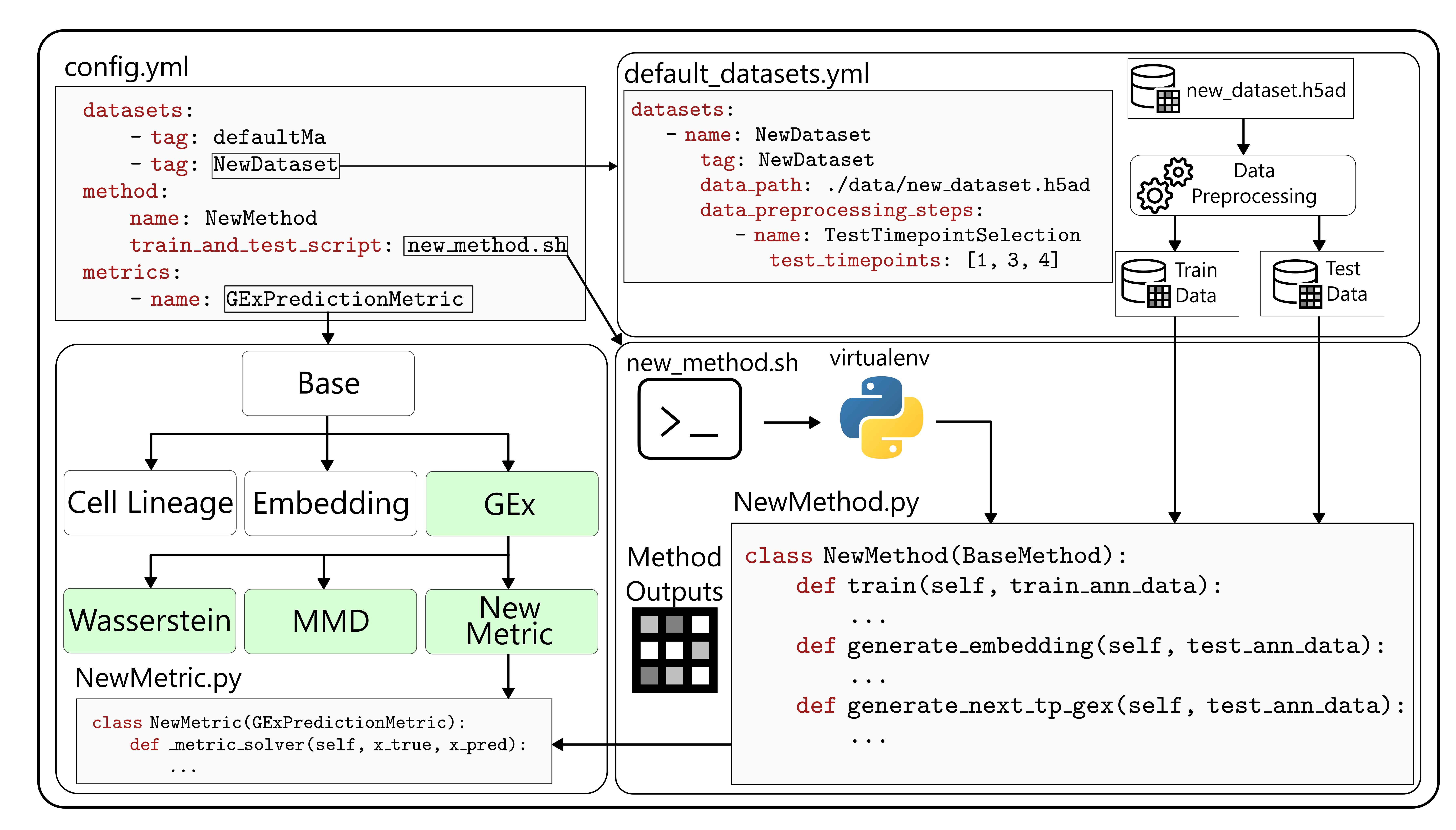
scTimeBench is controlled by a central configuration file which determines which datasets, methods, and metrics to run. An example of this can be found under `configs/scNODE/gex.yaml`.
|
|
97
|
+
|
|
98
|
+
### Detailed Layout of File Structure
|
|
99
|
+
* `configs/`: possible yaml config files to use as a starting point
|
|
100
|
+
* `extern/`: external packages that required edits for compatability such as pyPsupertime and Sceptic.
|
|
101
|
+
* `methods/`: the different methods that are possible to use, including defined submodules. Add your own methodology here.
|
|
102
|
+
* `src/`: where the scTimeBench package lies. See `src/ReadMe.md` for more documentation on the modules that exist there.
|
|
103
|
+
* `test/`: unit tests for each method, dataset, metric, and other important modules.
|
|
104
|
+
|
|
105
|
+
### Command-Line Interface Details
|
|
106
|
+
The entrypoint for the benchmark is defined as `scTimeBench`. Run `scTimeBench --help` for more details, or refer to `src/scTimeBench/config.py` and the documentation.
|
|
107
|
+
|
|
108
|
+
### Example Run
|
|
109
|
+
Run the package with:
|
|
110
|
+
```
|
|
111
|
+
scTimeBench --config configs/scNODE/gex.yaml
|
|
112
|
+
```
|
|
113
|
+
|
|
114
|
+
For a full running example using scNODE, refer to our example [Jupyter Notebook](https://colab.research.google.com/drive/1J-yNXu_FcSnhrCwTDQKjWCBSHsmdbohJ?usp=sharing).
|
|
115
|
+
|
|
116
|
+
## Contributing to scTimeBench
|
|
117
|
+
If you want to contribute, please install the dev environments with:
|
|
118
|
+
```
|
|
119
|
+
uv sync --extra dev --extra benchmark
|
|
120
|
+
```
|
|
121
|
+
or
|
|
122
|
+
```
|
|
123
|
+
pip install -e ".[dev, benchmark]"
|
|
124
|
+
```
|
|
125
|
+
|
|
126
|
+
To enable the autoformatting, please run:
|
|
127
|
+
```
|
|
128
|
+
pre-commit install
|
|
129
|
+
```
|
|
130
|
+
before committing.
|
|
131
|
+
|
|
132
|
+
Follow our example tutorials on adding new methods, datasets, and metrics in our documentation here: TODO-ADD-THIS.
|
|
133
|
+
|
|
134
|
+
### Testing
|
|
135
|
+
If your change heavily modifies the architecture, please run the necessary tests under the `test/` environment using pytest. Read more on the different available tests under `test/ReadMe.md`. See more information on the pytest documentation: https://docs.pytest.org/en/stable/. A useful flag is `-s` to view the entire output of the test.
|
|
136
|
+
|
|
137
|
+
## Citation
|
|
138
|
+
```bibtex
|
|
139
|
+
@article {scTimeBench,
|
|
140
|
+
author = {Osakwe, Adrien and Huang, Eric Haoran and Li, Yue},
|
|
141
|
+
title = {scTimeBench: A streamlined benchmarking platform for single-cell time-series analysis},
|
|
142
|
+
elocation-id = {2026.03.16.712069},
|
|
143
|
+
year = {2026},
|
|
144
|
+
doi = {10.64898/2026.03.16.712069},
|
|
145
|
+
publisher = {Cold Spring Harbor Laboratory},
|
|
146
|
+
URL = {https://www.biorxiv.org/content/early/2026/03/18/2026.03.16.712069},
|
|
147
|
+
eprint = {https://www.biorxiv.org/content/early/2026/03/18/2026.03.16.712069.full.pdf},
|
|
148
|
+
journal = {bioRxiv}
|
|
149
|
+
}
|
|
150
|
+
```
|
|
@@ -0,0 +1,118 @@
|
|
|
1
|
+
<!-- for this to work on Pypi, we need to point to the absolute path -->
|
|
2
|
+
<h1>
|
|
3
|
+
<img src="https://raw.githubusercontent.com/li-lab-mcgill/scTimeBench/refs/heads/main/assets/logo.png" alt="scTimeBench-Logo" height="150" align="absmiddle" /> scTimeBench
|
|
4
|
+
</h1>
|
|
5
|
+
|
|
6
|
+
[](https://www.python.org/downloads/release/python-31012/)
|
|
7
|
+
[](https://opensource.org/license/mit)
|
|
8
|
+
[](https://www.biorxiv.org/content/10.64898/2026.03.16.712069v1)
|
|
9
|
+
[](https://colab.research.google.com/drive/1J-yNXu_FcSnhrCwTDQKjWCBSHsmdbohJ?usp=sharing)
|
|
10
|
+
<!-- TODO: --> <!-- []() -->
|
|
11
|
+
|
|
12
|
+
|
|
13
|
+

|
|
14
|
+
|
|
15
|
+
## Table of Contents
|
|
16
|
+
- [Environment Setup](#environment-setup)
|
|
17
|
+
- [Suggested UV Installation](#suggested-uv-installation)
|
|
18
|
+
- [Standard Pip](#standard-pip)
|
|
19
|
+
- [Benchmark Architecture](#benchmark-architecture)
|
|
20
|
+
- [Detailed Layout of File Structure](#detailed-layout-of-file-structure)
|
|
21
|
+
- [Command-Line Interface Details](#command-line-interface-details)
|
|
22
|
+
- [Example Run](#example-run)
|
|
23
|
+
- [Contributing to scTimeBench](#contributing-to-sctimebench)
|
|
24
|
+
- [Citation](#citation)
|
|
25
|
+
|
|
26
|
+
## Environment Setup
|
|
27
|
+
scTimeBench was tested and supported using Python 3.10. If any other version that is 3.10+ does not work when using the benchmark, please submit an issue to this GitHub.
|
|
28
|
+
|
|
29
|
+
Important Note: this setup is needed twice. Once for the user to run the benchmark metrics, and the second time for the method itself needing to read from `scTimeBench.method_utils.method_runner` and other important shared constants. In short, this means we need to install scTimeBench into two separate virtual environments:
|
|
30
|
+
1. Your normal pip installation where you'll be running the benchmark from. This will require the extra "\[benchmark\]" group installation (pip) or the extra group installation `--extra benchmark` (uv).
|
|
31
|
+
2. For each method's virtual environment, you need to install the scTimeBench.
|
|
32
|
+
|
|
33
|
+
### Suggested UV Installation
|
|
34
|
+
Due to external dependencies and a more complex setup, we have decided to package everything under `uv` (see: https://github.com/astral-sh/uv). To start with, you need to get all the necessary extern dependencies, which can be done either by running:
|
|
35
|
+
```
|
|
36
|
+
git submodule update --init extern/
|
|
37
|
+
```
|
|
38
|
+
If you wish to benchmark across all methods, feel free to clone the submodules for all the methods as well with:
|
|
39
|
+
```
|
|
40
|
+
git submodule update --init
|
|
41
|
+
```
|
|
42
|
+
Then, install `uv` and run the following:
|
|
43
|
+
```
|
|
44
|
+
uv sync --extra benchmark
|
|
45
|
+
```
|
|
46
|
+
If you're using uv under a method's virtual environment, either the pip installation or the following will suffice:
|
|
47
|
+
```
|
|
48
|
+
uv sync
|
|
49
|
+
```
|
|
50
|
+
|
|
51
|
+
### Standard Pip
|
|
52
|
+
If the external dependencies such as pypsupertime or sceptic are not used (which they are not used by default), you can install using pip as follows:
|
|
53
|
+
```
|
|
54
|
+
pip install -e ".[benchmark]"
|
|
55
|
+
```
|
|
56
|
+
to run the benchmark. For your own method, simply install without the extra benchmarking requirements with
|
|
57
|
+
```
|
|
58
|
+
pip install -e .
|
|
59
|
+
```
|
|
60
|
+
There are extra dependencies that can be found under `pyproject.toml`.
|
|
61
|
+
|
|
62
|
+
## Benchmark Architecture
|
|
63
|
+

|
|
64
|
+
scTimeBench is controlled by a central configuration file which determines which datasets, methods, and metrics to run. An example of this can be found under `configs/scNODE/gex.yaml`.
|
|
65
|
+
|
|
66
|
+
### Detailed Layout of File Structure
|
|
67
|
+
* `configs/`: possible yaml config files to use as a starting point
|
|
68
|
+
* `extern/`: external packages that required edits for compatability such as pyPsupertime and Sceptic.
|
|
69
|
+
* `methods/`: the different methods that are possible to use, including defined submodules. Add your own methodology here.
|
|
70
|
+
* `src/`: where the scTimeBench package lies. See `src/ReadMe.md` for more documentation on the modules that exist there.
|
|
71
|
+
* `test/`: unit tests for each method, dataset, metric, and other important modules.
|
|
72
|
+
|
|
73
|
+
### Command-Line Interface Details
|
|
74
|
+
The entrypoint for the benchmark is defined as `scTimeBench`. Run `scTimeBench --help` for more details, or refer to `src/scTimeBench/config.py` and the documentation.
|
|
75
|
+
|
|
76
|
+
### Example Run
|
|
77
|
+
Run the package with:
|
|
78
|
+
```
|
|
79
|
+
scTimeBench --config configs/scNODE/gex.yaml
|
|
80
|
+
```
|
|
81
|
+
|
|
82
|
+
For a full running example using scNODE, refer to our example [Jupyter Notebook](https://colab.research.google.com/drive/1J-yNXu_FcSnhrCwTDQKjWCBSHsmdbohJ?usp=sharing).
|
|
83
|
+
|
|
84
|
+
## Contributing to scTimeBench
|
|
85
|
+
If you want to contribute, please install the dev environments with:
|
|
86
|
+
```
|
|
87
|
+
uv sync --extra dev --extra benchmark
|
|
88
|
+
```
|
|
89
|
+
or
|
|
90
|
+
```
|
|
91
|
+
pip install -e ".[dev, benchmark]"
|
|
92
|
+
```
|
|
93
|
+
|
|
94
|
+
To enable the autoformatting, please run:
|
|
95
|
+
```
|
|
96
|
+
pre-commit install
|
|
97
|
+
```
|
|
98
|
+
before committing.
|
|
99
|
+
|
|
100
|
+
Follow our example tutorials on adding new methods, datasets, and metrics in our documentation here: TODO-ADD-THIS.
|
|
101
|
+
|
|
102
|
+
### Testing
|
|
103
|
+
If your change heavily modifies the architecture, please run the necessary tests under the `test/` environment using pytest. Read more on the different available tests under `test/ReadMe.md`. See more information on the pytest documentation: https://docs.pytest.org/en/stable/. A useful flag is `-s` to view the entire output of the test.
|
|
104
|
+
|
|
105
|
+
## Citation
|
|
106
|
+
```bibtex
|
|
107
|
+
@article {scTimeBench,
|
|
108
|
+
author = {Osakwe, Adrien and Huang, Eric Haoran and Li, Yue},
|
|
109
|
+
title = {scTimeBench: A streamlined benchmarking platform for single-cell time-series analysis},
|
|
110
|
+
elocation-id = {2026.03.16.712069},
|
|
111
|
+
year = {2026},
|
|
112
|
+
doi = {10.64898/2026.03.16.712069},
|
|
113
|
+
publisher = {Cold Spring Harbor Laboratory},
|
|
114
|
+
URL = {https://www.biorxiv.org/content/early/2026/03/18/2026.03.16.712069},
|
|
115
|
+
eprint = {https://www.biorxiv.org/content/early/2026/03/18/2026.03.16.712069.full.pdf},
|
|
116
|
+
journal = {bioRxiv}
|
|
117
|
+
}
|
|
118
|
+
```
|
|
@@ -0,0 +1,61 @@
|
|
|
1
|
+
[build-system]
|
|
2
|
+
requires = ["setuptools", "wheel"]
|
|
3
|
+
build-backend = "setuptools.build_meta"
|
|
4
|
+
|
|
5
|
+
[project]
|
|
6
|
+
name = "scTimeBench" # This is the name you use for 'pip install'
|
|
7
|
+
version = "0.1.0"
|
|
8
|
+
dependencies = [
|
|
9
|
+
"PyYAML",
|
|
10
|
+
"scanpy",
|
|
11
|
+
"scikit-learn",
|
|
12
|
+
"fastparquet<2026.3.0",
|
|
13
|
+
]
|
|
14
|
+
requires-python = ">=3.10"
|
|
15
|
+
authors = [
|
|
16
|
+
{ name="Eric Haoran Huang", email="eric.h.huang@mail.mcgill.ca" },
|
|
17
|
+
{ name="Adrien Osakwe", email="adrien.osakwe@mail.mcgill.ca" },
|
|
18
|
+
{ name="Yue Li", email="yueli@cs.mcgill.ca" },
|
|
19
|
+
]
|
|
20
|
+
description = "A streamlined benchmarking platform for single-cell time-series analysis"
|
|
21
|
+
classifiers = [
|
|
22
|
+
"Development Status :: 4 - Beta",
|
|
23
|
+
"Intended Audience :: Science/Research",
|
|
24
|
+
"Topic :: Scientific/Engineering :: Bio-Informatics",
|
|
25
|
+
"Programming Language :: Python :: 3.10",
|
|
26
|
+
]
|
|
27
|
+
license = "MIT"
|
|
28
|
+
license-files = ["LICENSE"]
|
|
29
|
+
readme = "README.md"
|
|
30
|
+
|
|
31
|
+
[project.urls]
|
|
32
|
+
Homepage = "https://github.com/li-lab-mcgill/scTimeBench"
|
|
33
|
+
Issues = "https://github.com/li-lab-mcgill/scTimeBench/issues"
|
|
34
|
+
# TODO: have documentation here!
|
|
35
|
+
|
|
36
|
+
[tool.setuptools.packages.find]
|
|
37
|
+
where = ["src"] # Tells Python: "The actual code lives inside the src folder"
|
|
38
|
+
|
|
39
|
+
[tool.setuptools.package-data]
|
|
40
|
+
scTimeBench = [
|
|
41
|
+
"metrics/shared/*.yaml",
|
|
42
|
+
"metrics/shared/cell_lineages/**/*.txt",
|
|
43
|
+
]
|
|
44
|
+
|
|
45
|
+
[project.optional-dependencies]
|
|
46
|
+
dev = [
|
|
47
|
+
"pre-commit",
|
|
48
|
+
"pytest",
|
|
49
|
+
]
|
|
50
|
+
benchmark = [
|
|
51
|
+
"networkx",
|
|
52
|
+
"numpy",
|
|
53
|
+
"geomloss",
|
|
54
|
+
"pykeops",
|
|
55
|
+
"scanpy[leiden]",
|
|
56
|
+
"graphviz",
|
|
57
|
+
"celltypist"
|
|
58
|
+
]
|
|
59
|
+
|
|
60
|
+
[project.scripts]
|
|
61
|
+
scTimeBench = "scTimeBench.main:main"
|
|
@@ -0,0 +1,318 @@
|
|
|
1
|
+
"""
|
|
2
|
+
config.py
|
|
3
|
+
|
|
4
|
+
Configuration management for YAML-based configs, similar to the tf-binding project.
|
|
5
|
+
Handles both YAML file loading and command-line argument parsing.
|
|
6
|
+
"""
|
|
7
|
+
|
|
8
|
+
import argparse
|
|
9
|
+
import logging
|
|
10
|
+
import os
|
|
11
|
+
import yaml
|
|
12
|
+
|
|
13
|
+
from enum import Enum
|
|
14
|
+
|
|
15
|
+
|
|
16
|
+
# enum for the different run types, primarily:
|
|
17
|
+
# 1) auto_train_test: automatically run training and testing for methods that support it,
|
|
18
|
+
# by running the training and testing script specified.
|
|
19
|
+
# 2) preprocess: we preprocess the data and then save out a yaml file specifying requirements.
|
|
20
|
+
# The user handles training and testing outside of this framework.
|
|
21
|
+
# 3) eval_only: we only evaluate the metric on already generated data based on step 2).
|
|
22
|
+
class RunType(Enum):
|
|
23
|
+
AUTO_TRAIN_TEST = "auto_train_test"
|
|
24
|
+
PREPROCESS = "preprocess"
|
|
25
|
+
EVAL_ONLY = "eval_only"
|
|
26
|
+
TRAIN_ONLY = "train_only"
|
|
27
|
+
|
|
28
|
+
|
|
29
|
+
class Config:
|
|
30
|
+
"""Config class for both yaml and cli arguments."""
|
|
31
|
+
|
|
32
|
+
def __init__(self):
|
|
33
|
+
"""
|
|
34
|
+
Initialize config by parsing YAML file and command-line arguments.
|
|
35
|
+
CLI arguments override YAML settings.
|
|
36
|
+
"""
|
|
37
|
+
# Initiate parser and parse arguments
|
|
38
|
+
parser = argparse.ArgumentParser(
|
|
39
|
+
description="Single-cell trajectory analysis configuration"
|
|
40
|
+
)
|
|
41
|
+
|
|
42
|
+
# Config file argument
|
|
43
|
+
parser.add_argument(
|
|
44
|
+
"-c", "--config", type=str, help="Path to YAML configuration file"
|
|
45
|
+
)
|
|
46
|
+
|
|
47
|
+
# add metrics argument
|
|
48
|
+
parser.add_argument(
|
|
49
|
+
"--metrics",
|
|
50
|
+
type=str,
|
|
51
|
+
nargs="+",
|
|
52
|
+
help="List of metrics to compute",
|
|
53
|
+
)
|
|
54
|
+
|
|
55
|
+
parser.add_argument(
|
|
56
|
+
"--available",
|
|
57
|
+
action="store_true",
|
|
58
|
+
help="Show available methods, datasets, and metrics",
|
|
59
|
+
)
|
|
60
|
+
|
|
61
|
+
parser.add_argument(
|
|
62
|
+
"--print_all",
|
|
63
|
+
action="store_true",
|
|
64
|
+
help="Print all entries in the database tables",
|
|
65
|
+
)
|
|
66
|
+
|
|
67
|
+
parser.add_argument(
|
|
68
|
+
"--graph_sim_to_csv",
|
|
69
|
+
action="store_true",
|
|
70
|
+
help="Print graph similarity evaluations as CSV to stdout",
|
|
71
|
+
)
|
|
72
|
+
|
|
73
|
+
parser.add_argument(
|
|
74
|
+
"--output_csv_path",
|
|
75
|
+
type=str,
|
|
76
|
+
default="graph_sim.csv",
|
|
77
|
+
help="Optional path to save CSV output of graph similarity evaluations; if omitted, outputs to graph_sim.csv",
|
|
78
|
+
)
|
|
79
|
+
|
|
80
|
+
parser.add_argument(
|
|
81
|
+
"--clear_tables",
|
|
82
|
+
action="store_true",
|
|
83
|
+
help="Clear all entries in the database tables",
|
|
84
|
+
)
|
|
85
|
+
|
|
86
|
+
parser.add_argument(
|
|
87
|
+
"--view_evals_by_method",
|
|
88
|
+
action="store_true",
|
|
89
|
+
help="View existing evaluations of all metrics in the database per method set in the configuration",
|
|
90
|
+
)
|
|
91
|
+
|
|
92
|
+
parser.add_argument(
|
|
93
|
+
"--view_evals_by_metric",
|
|
94
|
+
action="store_true",
|
|
95
|
+
help="View existing evaluations of all methods in the database per metric set in the configuration",
|
|
96
|
+
)
|
|
97
|
+
|
|
98
|
+
parser.add_argument(
|
|
99
|
+
"--database_path",
|
|
100
|
+
type=str,
|
|
101
|
+
help="Path to the SQLite database file for storing results",
|
|
102
|
+
)
|
|
103
|
+
|
|
104
|
+
parser.add_argument(
|
|
105
|
+
"--run_type",
|
|
106
|
+
type=str,
|
|
107
|
+
choices=[rt.value for rt in RunType],
|
|
108
|
+
help="Type of run to perform: (default) auto_train_test, preprocess, eval_only, train_only. Defaults to auto_train_test.",
|
|
109
|
+
)
|
|
110
|
+
|
|
111
|
+
parser.add_argument(
|
|
112
|
+
"--output_dir",
|
|
113
|
+
type=str,
|
|
114
|
+
help="Directory to store outputs",
|
|
115
|
+
)
|
|
116
|
+
|
|
117
|
+
parser.add_argument(
|
|
118
|
+
"--log_level",
|
|
119
|
+
type=str,
|
|
120
|
+
choices=["DEBUG", "INFO", "WARNING", "ERROR", "CRITICAL"],
|
|
121
|
+
help="Logging level for the run (default: INFO)",
|
|
122
|
+
)
|
|
123
|
+
|
|
124
|
+
parser.add_argument(
|
|
125
|
+
"--log_file",
|
|
126
|
+
type=str,
|
|
127
|
+
help="Optional path to a log file; if omitted logs only go to stdout",
|
|
128
|
+
)
|
|
129
|
+
|
|
130
|
+
parser.add_argument(
|
|
131
|
+
"--data_dir",
|
|
132
|
+
type=str,
|
|
133
|
+
help="Optional base directory for dataset files, otherwise uses root paths specified in the config. If used, treats paths in config as either absolute or relative to this directory.",
|
|
134
|
+
)
|
|
135
|
+
|
|
136
|
+
parser.add_argument(
|
|
137
|
+
"--force_rerun",
|
|
138
|
+
action="store_true",
|
|
139
|
+
help="Usually duplicate method evaluations are skipped. This flag forces re-running even if evaluations already exist.",
|
|
140
|
+
)
|
|
141
|
+
|
|
142
|
+
parser.add_argument(
|
|
143
|
+
"-cf",
|
|
144
|
+
"--crispy-fishstick",
|
|
145
|
+
action="store_true",
|
|
146
|
+
help=argparse.SUPPRESS, # hide this from help since it's an Easter egg
|
|
147
|
+
)
|
|
148
|
+
|
|
149
|
+
# Parse known arguments
|
|
150
|
+
args = parser.parse_args()
|
|
151
|
+
|
|
152
|
+
# first handle the Easter egg
|
|
153
|
+
if args.crispy_fishstick:
|
|
154
|
+
from scTimeBench.shared.utils import animate, restore_interrupts
|
|
155
|
+
import sys
|
|
156
|
+
|
|
157
|
+
try:
|
|
158
|
+
animate()
|
|
159
|
+
finally:
|
|
160
|
+
restore_interrupts()
|
|
161
|
+
sys.stdout.write("\033[?25h") # Re-show the cursor if you hid it
|
|
162
|
+
exit()
|
|
163
|
+
|
|
164
|
+
# Get all config keys
|
|
165
|
+
config_keys = list(args.__dict__.keys())
|
|
166
|
+
|
|
167
|
+
# other keys to add from the yaml file
|
|
168
|
+
config_keys.extend(["method", "datasets"])
|
|
169
|
+
|
|
170
|
+
# First read the config file if provided
|
|
171
|
+
assert (
|
|
172
|
+
args.config is not None
|
|
173
|
+
), "Config file path must be provided with --config"
|
|
174
|
+
if not os.path.exists(args.config):
|
|
175
|
+
raise FileNotFoundError(f"Config file not found: {args.config}")
|
|
176
|
+
|
|
177
|
+
with open(args.config, "r") as file:
|
|
178
|
+
data = yaml.safe_load(file)
|
|
179
|
+
|
|
180
|
+
# Set attributes from YAML file
|
|
181
|
+
for key in config_keys:
|
|
182
|
+
if key in data.keys():
|
|
183
|
+
setattr(self, key, data[key])
|
|
184
|
+
|
|
185
|
+
# Override with command-line arguments
|
|
186
|
+
for key, value in args._get_kwargs():
|
|
187
|
+
if value is not None:
|
|
188
|
+
setattr(self, key, value)
|
|
189
|
+
|
|
190
|
+
# Set defaults for optional parameters
|
|
191
|
+
defaults = {
|
|
192
|
+
"database_path": "scTimeBench.db",
|
|
193
|
+
"run_type": RunType.AUTO_TRAIN_TEST.value,
|
|
194
|
+
"output_dir": "outputs/",
|
|
195
|
+
"datasets": [],
|
|
196
|
+
"log_level": "INFO",
|
|
197
|
+
"log_file": None,
|
|
198
|
+
}
|
|
199
|
+
|
|
200
|
+
for key, value in defaults.items():
|
|
201
|
+
if not hasattr(self, key) or getattr(self, key) is None:
|
|
202
|
+
setattr(self, key, value)
|
|
203
|
+
|
|
204
|
+
# Configure logging with stdout always enabled and optional file output.
|
|
205
|
+
resolved_log_level = getattr(logging, str(self.log_level).upper(), logging.INFO)
|
|
206
|
+
handlers = [logging.StreamHandler()]
|
|
207
|
+
if self.log_file:
|
|
208
|
+
log_dir = os.path.dirname(self.log_file)
|
|
209
|
+
if log_dir:
|
|
210
|
+
os.makedirs(log_dir, exist_ok=True)
|
|
211
|
+
handlers.append(logging.FileHandler(self.log_file))
|
|
212
|
+
|
|
213
|
+
logging.basicConfig(
|
|
214
|
+
level=resolved_log_level,
|
|
215
|
+
format="%(asctime)s | %(levelname)s | %(name)s | %(message)s",
|
|
216
|
+
handlers=handlers,
|
|
217
|
+
)
|
|
218
|
+
|
|
219
|
+
# Validate required fields
|
|
220
|
+
required_fields = ["method", "metrics"]
|
|
221
|
+
for field in required_fields:
|
|
222
|
+
assert (
|
|
223
|
+
hasattr(self, field) and getattr(self, field) is not None
|
|
224
|
+
), f"Required field '{field}' must be specified in config file or as --{field}"
|
|
225
|
+
|
|
226
|
+
# make sure no other fields exist besides this + datasets in the yaml
|
|
227
|
+
allowed_fields = set(required_fields + ["datasets", "metrics_skiplist"])
|
|
228
|
+
for field in data.keys():
|
|
229
|
+
if field not in allowed_fields:
|
|
230
|
+
raise ValueError(
|
|
231
|
+
f"Unknown field '{field}' found in config. Allowed fields are {allowed_fields}."
|
|
232
|
+
)
|
|
233
|
+
|
|
234
|
+
# validate the fields within each larger section
|
|
235
|
+
# N.B.: we don't need them to specify preprocessors because it might already be preprocessed
|
|
236
|
+
# ** DATASETS **
|
|
237
|
+
dataset_required_fields = ["data_path", "name"]
|
|
238
|
+
dataset_optional_fields = ["data_preprocessing_steps"]
|
|
239
|
+
dataset_alternate_field = "tag"
|
|
240
|
+
|
|
241
|
+
for dataset in self.datasets:
|
|
242
|
+
# we want to make sure either all of the required fields are specified,
|
|
243
|
+
# or the alternate field is specified, but not a mix of both
|
|
244
|
+
if dataset_alternate_field in dataset:
|
|
245
|
+
if any([field in dataset for field in dataset_required_fields]):
|
|
246
|
+
raise ValueError(
|
|
247
|
+
f"Dataset config cannot have both '{dataset_alternate_field}' and any other fields {dataset_required_fields}."
|
|
248
|
+
)
|
|
249
|
+
continue
|
|
250
|
+
|
|
251
|
+
for field in dataset_required_fields:
|
|
252
|
+
assert (
|
|
253
|
+
field in dataset
|
|
254
|
+
), f"Required dataset field '{field}' must be specified in config file, or use alternate flag \"tag\" instead."
|
|
255
|
+
|
|
256
|
+
# also ensure that all the fields found in the dataset are only of the required fields
|
|
257
|
+
# this is to ensure caching also works properly
|
|
258
|
+
for field in dataset.keys():
|
|
259
|
+
if (
|
|
260
|
+
field not in dataset_required_fields
|
|
261
|
+
and field not in dataset_optional_fields
|
|
262
|
+
):
|
|
263
|
+
raise ValueError(
|
|
264
|
+
f"Unknown field '{field}' found in dataset config. Allowed fields are {dataset_required_fields} or '{dataset_alternate_field}'."
|
|
265
|
+
)
|
|
266
|
+
|
|
267
|
+
# ** METHOD **
|
|
268
|
+
method_required_fields = ["name"]
|
|
269
|
+
|
|
270
|
+
for field in method_required_fields:
|
|
271
|
+
assert (
|
|
272
|
+
field in self.method
|
|
273
|
+
), f"Required method field '{field}' must be specified in config file"
|
|
274
|
+
|
|
275
|
+
# Validate paths exist
|
|
276
|
+
dataset_path_keys = [
|
|
277
|
+
"data_path",
|
|
278
|
+
"cell_lineage_file",
|
|
279
|
+
"cell_equivalence_file",
|
|
280
|
+
]
|
|
281
|
+
method_path_keys = [
|
|
282
|
+
"train_and_test_script",
|
|
283
|
+
]
|
|
284
|
+
paths = {
|
|
285
|
+
*{
|
|
286
|
+
value
|
|
287
|
+
for dataset in self.datasets
|
|
288
|
+
for key, value in dataset.items()
|
|
289
|
+
if key in dataset_path_keys
|
|
290
|
+
},
|
|
291
|
+
*{value for key, value in self.method.items() if key in method_path_keys},
|
|
292
|
+
}
|
|
293
|
+
|
|
294
|
+
for path in paths:
|
|
295
|
+
assert os.path.exists(path), f"Path for '{path}' does not exist: {path}"
|
|
296
|
+
|
|
297
|
+
# set the run type
|
|
298
|
+
self.run_type = RunType(self.run_type)
|
|
299
|
+
|
|
300
|
+
# verify that the train and test script is specified if auto_train_test is set
|
|
301
|
+
if self.run_type == RunType.AUTO_TRAIN_TEST:
|
|
302
|
+
assert (
|
|
303
|
+
"train_and_test_script" in self.method
|
|
304
|
+
), "Method must specify 'train_and_test_script' to use --auto_train_test"
|
|
305
|
+
|
|
306
|
+
# finally, to be used later for saving the original config
|
|
307
|
+
# into the method output yaml config:
|
|
308
|
+
self.method_yaml_data = data["method"]
|
|
309
|
+
|
|
310
|
+
# then let's also add in the metric skip list
|
|
311
|
+
self.metrics_skiplist = data.get("metrics_skiplist", [])
|
|
312
|
+
self.metrics_skiplist = [
|
|
313
|
+
metric if isinstance(metric, str) else metric.get("name", "")
|
|
314
|
+
for metric in self.metrics_skiplist
|
|
315
|
+
]
|
|
316
|
+
|
|
317
|
+
logging.info("Configuration successfully loaded")
|
|
318
|
+
logging.debug("Configuration details: %s", self.__dict__)
|