priorcons 0.1.0__tar.gz
This diff represents the content of publicly available package versions that have been released to one of the supported registries. The information contained in this diff is provided for informational purposes only and reflects changes between package versions as they appear in their respective public registries.
- priorcons-0.1.0/LICENSE +29 -0
- priorcons-0.1.0/PKG-INFO +276 -0
- priorcons-0.1.0/README.md +229 -0
- priorcons-0.1.0/priorcons/__init__.py +13 -0
- priorcons-0.1.0/priorcons/build_priors.py +39 -0
- priorcons-0.1.0/priorcons/cli.py +49 -0
- priorcons-0.1.0/priorcons/integrate_consensus.py +154 -0
- priorcons-0.1.0/priorcons/utils.py +188 -0
- priorcons-0.1.0/priorcons/utils_integrate_consensus.py +387 -0
- priorcons-0.1.0/priorcons.egg-info/PKG-INFO +276 -0
- priorcons-0.1.0/priorcons.egg-info/SOURCES.txt +16 -0
- priorcons-0.1.0/priorcons.egg-info/dependency_links.txt +1 -0
- priorcons-0.1.0/priorcons.egg-info/entry_points.txt +2 -0
- priorcons-0.1.0/priorcons.egg-info/requires.txt +4 -0
- priorcons-0.1.0/priorcons.egg-info/top_level.txt +1 -0
- priorcons-0.1.0/pyproject.toml +26 -0
- priorcons-0.1.0/setup.cfg +26 -0
priorcons-0.1.0/LICENSE
ADDED
|
@@ -0,0 +1,29 @@
|
|
|
1
|
+
Attribution Permissive License (APL) 1.0
|
|
2
|
+
|
|
3
|
+
Copyright (c) 2025 Germán Vallejo Palma
|
|
4
|
+
|
|
5
|
+
Permission is hereby granted, free of charge, to any person obtaining a copy
|
|
6
|
+
of this software and associated documentation files (the "Software"), to use,
|
|
7
|
+
copy, modify, merge, publish, distribute, sublicense, and/or sell copies of the
|
|
8
|
+
Software, and to permit persons to whom the Software is furnished to do so,
|
|
9
|
+
subject to the following condition:
|
|
10
|
+
|
|
11
|
+
* ATTRIBUTION: Redistributions of source or binary form, modified or
|
|
12
|
+
unmodified, must retain the following attribution notice in a conspicuous
|
|
13
|
+
location (for example, in the repository README, the package metadata,
|
|
14
|
+
or in a NOTICE file shipped with binaries):
|
|
15
|
+
|
|
16
|
+
"This software was developed by Germán Vallejo Palma at the Instituto de
|
|
17
|
+
Salud Carlos III — National Centre of Microbiology (Respiratory Viruses
|
|
18
|
+
and Influenza Unit)."
|
|
19
|
+
|
|
20
|
+
The above copyright notice and this permission notice shall be included in all
|
|
21
|
+
copies or substantial portions of the Software.
|
|
22
|
+
|
|
23
|
+
THE SOFTWARE IS PROVIDED "AS IS", WITHOUT WARRANTY OF ANY KIND, EXPRESS OR
|
|
24
|
+
IMPLIED, INCLUDING BUT NOT LIMITED TO THE WARRANTIES OF MERCHANTABILITY,
|
|
25
|
+
FITNESS FOR A PARTICULAR PURPOSE AND NONINFRINGEMENT. IN NO EVENT SHALL THE
|
|
26
|
+
AUTHORS OR COPYRIGHT HOLDERS BE LIABLE FOR ANY CLAIM, DAMAGES OR OTHER
|
|
27
|
+
LIABILITY, WHETHER IN AN ACTION OF CONTRACT, TORT OR OTHERWISE, ARISING FROM,
|
|
28
|
+
OUT OF OR IN CONNECTION WITH THE SOFTWARE OR THE USE OR OTHER DEALINGS IN THE
|
|
29
|
+
SOFTWARE.
|
priorcons-0.1.0/PKG-INFO
ADDED
|
@@ -0,0 +1,276 @@
|
|
|
1
|
+
Metadata-Version: 2.4
|
|
2
|
+
Name: priorcons
|
|
3
|
+
Version: 0.1.0
|
|
4
|
+
Summary: Tool for the integration of viral consensus sequences obtained by de novo and mapping strategies, supported by prior information.
|
|
5
|
+
Home-page: https://github.com/GERMAN00VP/priorCons
|
|
6
|
+
Author: Germán Vallejo Palma
|
|
7
|
+
Author-email: Germán Vallejo Palma <german.vallejo@isciii.es>
|
|
8
|
+
License: Attribution Permissive License (APL) 1.0
|
|
9
|
+
|
|
10
|
+
Copyright (c) 2025 Germán Vallejo Palma
|
|
11
|
+
|
|
12
|
+
Permission is hereby granted, free of charge, to any person obtaining a copy
|
|
13
|
+
of this software and associated documentation files (the "Software"), to use,
|
|
14
|
+
copy, modify, merge, publish, distribute, sublicense, and/or sell copies of the
|
|
15
|
+
Software, and to permit persons to whom the Software is furnished to do so,
|
|
16
|
+
subject to the following condition:
|
|
17
|
+
|
|
18
|
+
* ATTRIBUTION: Redistributions of source or binary form, modified or
|
|
19
|
+
unmodified, must retain the following attribution notice in a conspicuous
|
|
20
|
+
location (for example, in the repository README, the package metadata,
|
|
21
|
+
or in a NOTICE file shipped with binaries):
|
|
22
|
+
|
|
23
|
+
"This software was developed by Germán Vallejo Palma at the Instituto de
|
|
24
|
+
Salud Carlos III — National Centre of Microbiology (Respiratory Viruses
|
|
25
|
+
and Influenza Unit)."
|
|
26
|
+
|
|
27
|
+
The above copyright notice and this permission notice shall be included in all
|
|
28
|
+
copies or substantial portions of the Software.
|
|
29
|
+
|
|
30
|
+
THE SOFTWARE IS PROVIDED "AS IS", WITHOUT WARRANTY OF ANY KIND, EXPRESS OR
|
|
31
|
+
IMPLIED, INCLUDING BUT NOT LIMITED TO THE WARRANTIES OF MERCHANTABILITY,
|
|
32
|
+
FITNESS FOR A PARTICULAR PURPOSE AND NONINFRINGEMENT. IN NO EVENT SHALL THE
|
|
33
|
+
AUTHORS OR COPYRIGHT HOLDERS BE LIABLE FOR ANY CLAIM, DAMAGES OR OTHER
|
|
34
|
+
LIABILITY, WHETHER IN AN ACTION OF CONTRACT, TORT OR OTHERWISE, ARISING FROM,
|
|
35
|
+
OUT OF OR IN CONNECTION WITH THE SOFTWARE OR THE USE OR OTHER DEALINGS IN THE
|
|
36
|
+
SOFTWARE.
|
|
37
|
+
|
|
38
|
+
Project-URL: Homepage, https://github.com/GERMAN00VP/priorCons/
|
|
39
|
+
Requires-Python: >=3.8
|
|
40
|
+
Description-Content-Type: text/markdown
|
|
41
|
+
License-File: LICENSE
|
|
42
|
+
Requires-Dist: biopython>=1.79
|
|
43
|
+
Requires-Dist: numpy>=1.21
|
|
44
|
+
Requires-Dist: pandas>=1.3
|
|
45
|
+
Requires-Dist: pyarrow>=12.0
|
|
46
|
+
Dynamic: license-file
|
|
47
|
+
|
|
48
|
+
# PriorCons
|
|
49
|
+
|
|
50
|
+
This repository provides tools to:
|
|
51
|
+
|
|
52
|
+
1. **Generate Integrated Consensus (`integrate_consensus.py`)**
|
|
53
|
+
Produces a high-quality viral consensus by strategically using **ABACAS** sequences to fill missing regions in the **mapping** consensus. It employs a sliding-window approach that verifies the evolutionary plausibility of ABACAS content against empirical priors before incorporation.
|
|
54
|
+
|
|
55
|
+
2. **Build Evolutionary Priors (`build_priors.py`)**
|
|
56
|
+
Constructs empirical prior distributions from large multiple-sequence alignments. These priors model expected genetic variation across genomic windows and provide likelihood thresholds for quality control during consensus integration.
|
|
57
|
+
|
|
58
|
+
3. **Access Supporting Utilities (`utils` scripts)**
|
|
59
|
+
Provides modular helper functions for alignment processing, window scoring, and consensus construction used by both main workflows.
|
|
60
|
+
|
|
61
|
+
---
|
|
62
|
+
### Installation
|
|
63
|
+
|
|
64
|
+
```bash
|
|
65
|
+
pip install priorcons
|
|
66
|
+
```
|
|
67
|
+
---
|
|
68
|
+
|
|
69
|
+
### CLI usage
|
|
70
|
+
```bash
|
|
71
|
+
# Create priors
|
|
72
|
+
priorcons build-priors --input sequences.fasta --ref REF_ID --output priors.parquet
|
|
73
|
+
|
|
74
|
+
# Run consensus integration
|
|
75
|
+
priorcons integrate-consensus --input alignment.aln --ref REF_ID --prior priors.parquet --output_dir results
|
|
76
|
+
|
|
77
|
+
```
|
|
78
|
+
|
|
79
|
+
---
|
|
80
|
+
|
|
81
|
+
## 🚀 Main Script: integrate_consensus.py
|
|
82
|
+
|
|
83
|
+
This is the entrypoint of the tool. It creates a *integrated consensus sequence* by combining mapping consensus and ABACAS output, both aligned to a reference sequence, but only after performing quality control (QC) at the window level.
|
|
84
|
+
|
|
85
|
+
### 🔑 Inputs
|
|
86
|
+
|
|
87
|
+
- `--input` → path to an alignment file (`.aln`) containing at least:
|
|
88
|
+
- **1º Reference sequence**
|
|
89
|
+
- **2º Mapping consensus sequence**
|
|
90
|
+
- **3º ABACAS consensus sequence**
|
|
91
|
+
|
|
92
|
+
**The sequences in the alignment file must be provided in the specified order, as they will be identified by their position.**
|
|
93
|
+
|
|
94
|
+
|
|
95
|
+
- `--ref` → ID of the reference sequence in the alignment.
|
|
96
|
+
|
|
97
|
+
- `--prior` → path to a priors table (`.parquet`) generated with `build_priors.py`.
|
|
98
|
+
|
|
99
|
+
- `--output_dir` → directory to save the results.
|
|
100
|
+
|
|
101
|
+
|
|
102
|
+
---
|
|
103
|
+
|
|
104
|
+
### 🧪 Workflow
|
|
105
|
+
|
|
106
|
+
1. **Start with mapping consensus** as the baseline
|
|
107
|
+
2. **Identify missing/unreliable regions** in mapping consensus
|
|
108
|
+
3. **For each window**:
|
|
109
|
+
- If mapping has coverage → keep mapping sequence
|
|
110
|
+
- If mapping has missing data → evaluate ABACAS for that window:
|
|
111
|
+
* Check fragmentation and quality
|
|
112
|
+
* Verify evolutionary plausibility using priors [(nLL score)](README.md#-methodology-build_priorspy)
|
|
113
|
+
* If ABACAS passes QC → use ABACAS to fill missing regions
|
|
114
|
+
4. **Construct final consensus** combining mapping baseline with validated ABACAS fills
|
|
115
|
+
5. **Restore mapping-specific insertions**
|
|
116
|
+
6. **QC reporting**: compute coverage, substitutions, and insertion metrics comparing the final integrated consensus to MAPPING.
|
|
117
|
+
|
|
118
|
+
---
|
|
119
|
+
|
|
120
|
+
### 📦 Outputs
|
|
121
|
+
|
|
122
|
+
The script produces **three files** inside `--output_dir`:
|
|
123
|
+
|
|
124
|
+
1. **Integrated consensus FASTA**
|
|
125
|
+
- File: `<basename>-INTEGRATED.fasta`
|
|
126
|
+
- Contains the final consensus sequence after merging and reinserting insertions.
|
|
127
|
+
|
|
128
|
+
2. **Window QC trace (CSV)**
|
|
129
|
+
- File: `windows_trace.csv`
|
|
130
|
+
- One row per window, recording:
|
|
131
|
+
- `start`, `end` → genomic coordinates.
|
|
132
|
+
- `MISSING_MAPPING`, `MISSING_ABACAS` → counts of missing bases.
|
|
133
|
+
- `ABACAS_MORE_INFO` → whether ABACAS has fewer missing bases than MAPPING.
|
|
134
|
+
- `ABACAS_FRAGMENTS` → fragmentation level of ABACAS in this window (keep: 0 < n fragments < 3 ).
|
|
135
|
+
- `WINDOW_PRIOR_nLL_p95` → threshold from priors.
|
|
136
|
+
- `WINDOW_SCORE_nLL` → score of ABACAS in this window.
|
|
137
|
+
- `WINDOW_QC_PASSED` → True/False decision.
|
|
138
|
+
|
|
139
|
+
3. **Consensus QC summary (JSON)**
|
|
140
|
+
- File: `qc.json`
|
|
141
|
+
- Provides overall metrics comparing the MAPPING consensus and the integrated consensus:
|
|
142
|
+
- `MAPPING_COVERAGE` → % of genome covered in MAPPING.
|
|
143
|
+
- `FINAL_COVERAGE` → % of genome covered in integrated consensus.
|
|
144
|
+
- `MAPPING_SUBSTITUTIONS` → substitutions vs. reference in MAPPING.
|
|
145
|
+
- `FINAL_SUBSTITUTIONS` → substitutions vs. reference in integrated consensus.
|
|
146
|
+
- `EXPECTED_SUBSTITUTIONS` → expected number of substitutions, extrapolated from mapping.
|
|
147
|
+
- `OBS-EXP_SUBSTITUTIONS` → difference between observed and expected substitutions.
|
|
148
|
+
- `N_INSERTIONS` → number of insertions added back.
|
|
149
|
+
- `TOTAL_INSERTIONS_LENGTH` → total inserted length.
|
|
150
|
+
- `INSERTIONS` → list of insertions with their coordinates.
|
|
151
|
+
|
|
152
|
+
---
|
|
153
|
+
|
|
154
|
+
### ▶️ Example run
|
|
155
|
+
|
|
156
|
+
```bash
|
|
157
|
+
python integrate_consensus.py \
|
|
158
|
+
--input /path/to/<sample_name>.aln \
|
|
159
|
+
--ref RSV_BD \
|
|
160
|
+
--prior /path/to/RSVBD_win100_ovlp50_priors.parquet \
|
|
161
|
+
--output_dir results
|
|
162
|
+
```
|
|
163
|
+
----
|
|
164
|
+
|
|
165
|
+
This will generate:
|
|
166
|
+
|
|
167
|
+
- `results/<sample_name>-INTEGRATED.fasta`
|
|
168
|
+
- `results/windows_trace.csv`
|
|
169
|
+
- `results/qc.json`
|
|
170
|
+
|
|
171
|
+
---
|
|
172
|
+
|
|
173
|
+
## 🛠 Script: build_priors.py
|
|
174
|
+
|
|
175
|
+
This script creates **empirical priors** (overlapped windows) from a large multiple sequence alignment.
|
|
176
|
+
These priors are later used by `integrate_consensus.py` to evaluate windows.
|
|
177
|
+
|
|
178
|
+
### 🔑 Inputs
|
|
179
|
+
|
|
180
|
+
- `-i / --input` → aligned FASTA file with multiple sequences.
|
|
181
|
+
- `-r / --ref` → ID of the reference sequence.
|
|
182
|
+
- `-o / --output` → output file (`.parquet`).
|
|
183
|
+
- `--win` → window size (default: 100).
|
|
184
|
+
- `--overlap` → overlap size (default: 10).
|
|
185
|
+
|
|
186
|
+
### ▶️ Example run
|
|
187
|
+
|
|
188
|
+
```bash
|
|
189
|
+
python build_priors.py \
|
|
190
|
+
-i alignment.fasta \
|
|
191
|
+
-r ReferenceID \
|
|
192
|
+
-o priors.parquet \
|
|
193
|
+
--win 100 \
|
|
194
|
+
--overlap 10
|
|
195
|
+
```
|
|
196
|
+
----
|
|
197
|
+
|
|
198
|
+
### 📦 Output
|
|
199
|
+
|
|
200
|
+
A `.parquet` file with one row per window, containing:
|
|
201
|
+
|
|
202
|
+
- `start`, `end` → window coordinates.
|
|
203
|
+
- `nLL_p95`, `nLL_p99` → empirical thresholds.
|
|
204
|
+
- `profile` → base probability distributions for each position in the window.
|
|
205
|
+
|
|
206
|
+
## 🧮 Methodology (build_priors.py)
|
|
207
|
+
|
|
208
|
+
### 1. Probability distributions per position
|
|
209
|
+
|
|
210
|
+
For each window of size `W` bases (e.g., `W = 100`), and for each position `j` within that window, we compute the probability of observing each nucleotide:
|
|
211
|
+
|
|
212
|
+
=\frac{c_j(b)+\alpha}{\sum_{x\in\{A,C,G,T\}}(c_j(x)+\alpha)})
|
|
213
|
+
|
|
214
|
+
Where:
|
|
215
|
+
- ) = number of sequences with base 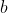 at position 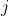.
|
|
216
|
+
- 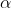 = pseudocount (Laplace smoothing, default ) to avoid zero probabilities.
|
|
217
|
+
- Bases `N` are ignored in the counts.
|
|
218
|
+
|
|
219
|
+
This gives a **per-position categorical distribution**.
|
|
220
|
+
|
|
221
|
+
---
|
|
222
|
+
|
|
223
|
+
### 2. Log-likelihood of a sequence in a window
|
|
224
|
+
|
|
225
|
+
Given a query sequence 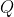, we compute its probability under the window profile.
|
|
226
|
+
For each valid (non-`N`) position 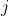 with observed base :
|
|
227
|
+
|
|
228
|
+
=\sum_{j=1}^{W}\log%20P_j(q_j))
|
|
229
|
+
|
|
230
|
+
The **normalized negative log-likelihood (nLL)** is:
|
|
231
|
+
|
|
232
|
+
=-\frac{1}{N_{\text{valid}}}\sum_{j=1}^{W}\log%20P_j(q_j))
|
|
233
|
+
|
|
234
|
+
Where:
|
|
235
|
+
-  = number of positions in the window where 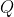 has a non-`N` base.
|
|
236
|
+
|
|
237
|
+
Smaller nLL values indicate sequences more likely under the empirical profile.
|
|
238
|
+
|
|
239
|
+
|
|
240
|
+
### 3. Empirical priors
|
|
241
|
+
|
|
242
|
+
To characterize "normal variation" for each window:
|
|
243
|
+
|
|
244
|
+
1. Score **all sequences** from the alignment against the window profile.
|
|
245
|
+
2. Collect the distribution of nLL values.
|
|
246
|
+
3. Extract percentiles (e.g., 95th and 99th) to serve as thresholds.
|
|
247
|
+
|
|
248
|
+
Thus, for each window we store:
|
|
249
|
+
- The **distribution (profile)**.
|
|
250
|
+
- Empirical thresholds: `nLL_p95` and `nLL_p99`.
|
|
251
|
+
|
|
252
|
+
A new sequence can later be compared:
|
|
253
|
+
- If `nLL < nLL_p95` → typical.
|
|
254
|
+
- If `nLL > nLL_p99` → unusually variable, possibly unreliable region.
|
|
255
|
+
|
|
256
|
+
---
|
|
257
|
+
|
|
258
|
+
---
|
|
259
|
+
|
|
260
|
+
## � Supporting utils
|
|
261
|
+
|
|
262
|
+
Several utility scripts provide reusable functions for both processes:
|
|
263
|
+
|
|
264
|
+
- **utils.py** → basic alignment and scoring functions:
|
|
265
|
+
- `load_alignment`, `extract_ref_positions`, `sliding_windows`, `score_window`.
|
|
266
|
+
|
|
267
|
+
- **utils_integrate_consensus.py** → additional helpers for consensus integration:
|
|
268
|
+
- missingness and fragmentation counts,
|
|
269
|
+
- insertion handling,
|
|
270
|
+
- QC calculations,
|
|
271
|
+
- consensus merging,
|
|
272
|
+
- window evaluation wrapper.
|
|
273
|
+
|
|
274
|
+
These modular functions keep the pipeline clean and reusable.
|
|
275
|
+
|
|
276
|
+
|
|
@@ -0,0 +1,229 @@
|
|
|
1
|
+
# PriorCons
|
|
2
|
+
|
|
3
|
+
This repository provides tools to:
|
|
4
|
+
|
|
5
|
+
1. **Generate Integrated Consensus (`integrate_consensus.py`)**
|
|
6
|
+
Produces a high-quality viral consensus by strategically using **ABACAS** sequences to fill missing regions in the **mapping** consensus. It employs a sliding-window approach that verifies the evolutionary plausibility of ABACAS content against empirical priors before incorporation.
|
|
7
|
+
|
|
8
|
+
2. **Build Evolutionary Priors (`build_priors.py`)**
|
|
9
|
+
Constructs empirical prior distributions from large multiple-sequence alignments. These priors model expected genetic variation across genomic windows and provide likelihood thresholds for quality control during consensus integration.
|
|
10
|
+
|
|
11
|
+
3. **Access Supporting Utilities (`utils` scripts)**
|
|
12
|
+
Provides modular helper functions for alignment processing, window scoring, and consensus construction used by both main workflows.
|
|
13
|
+
|
|
14
|
+
---
|
|
15
|
+
### Installation
|
|
16
|
+
|
|
17
|
+
```bash
|
|
18
|
+
pip install priorcons
|
|
19
|
+
```
|
|
20
|
+
---
|
|
21
|
+
|
|
22
|
+
### CLI usage
|
|
23
|
+
```bash
|
|
24
|
+
# Create priors
|
|
25
|
+
priorcons build-priors --input sequences.fasta --ref REF_ID --output priors.parquet
|
|
26
|
+
|
|
27
|
+
# Run consensus integration
|
|
28
|
+
priorcons integrate-consensus --input alignment.aln --ref REF_ID --prior priors.parquet --output_dir results
|
|
29
|
+
|
|
30
|
+
```
|
|
31
|
+
|
|
32
|
+
---
|
|
33
|
+
|
|
34
|
+
## 🚀 Main Script: integrate_consensus.py
|
|
35
|
+
|
|
36
|
+
This is the entrypoint of the tool. It creates a *integrated consensus sequence* by combining mapping consensus and ABACAS output, both aligned to a reference sequence, but only after performing quality control (QC) at the window level.
|
|
37
|
+
|
|
38
|
+
### 🔑 Inputs
|
|
39
|
+
|
|
40
|
+
- `--input` → path to an alignment file (`.aln`) containing at least:
|
|
41
|
+
- **1º Reference sequence**
|
|
42
|
+
- **2º Mapping consensus sequence**
|
|
43
|
+
- **3º ABACAS consensus sequence**
|
|
44
|
+
|
|
45
|
+
**The sequences in the alignment file must be provided in the specified order, as they will be identified by their position.**
|
|
46
|
+
|
|
47
|
+
|
|
48
|
+
- `--ref` → ID of the reference sequence in the alignment.
|
|
49
|
+
|
|
50
|
+
- `--prior` → path to a priors table (`.parquet`) generated with `build_priors.py`.
|
|
51
|
+
|
|
52
|
+
- `--output_dir` → directory to save the results.
|
|
53
|
+
|
|
54
|
+
|
|
55
|
+
---
|
|
56
|
+
|
|
57
|
+
### 🧪 Workflow
|
|
58
|
+
|
|
59
|
+
1. **Start with mapping consensus** as the baseline
|
|
60
|
+
2. **Identify missing/unreliable regions** in mapping consensus
|
|
61
|
+
3. **For each window**:
|
|
62
|
+
- If mapping has coverage → keep mapping sequence
|
|
63
|
+
- If mapping has missing data → evaluate ABACAS for that window:
|
|
64
|
+
* Check fragmentation and quality
|
|
65
|
+
* Verify evolutionary plausibility using priors [(nLL score)](README.md#-methodology-build_priorspy)
|
|
66
|
+
* If ABACAS passes QC → use ABACAS to fill missing regions
|
|
67
|
+
4. **Construct final consensus** combining mapping baseline with validated ABACAS fills
|
|
68
|
+
5. **Restore mapping-specific insertions**
|
|
69
|
+
6. **QC reporting**: compute coverage, substitutions, and insertion metrics comparing the final integrated consensus to MAPPING.
|
|
70
|
+
|
|
71
|
+
---
|
|
72
|
+
|
|
73
|
+
### 📦 Outputs
|
|
74
|
+
|
|
75
|
+
The script produces **three files** inside `--output_dir`:
|
|
76
|
+
|
|
77
|
+
1. **Integrated consensus FASTA**
|
|
78
|
+
- File: `<basename>-INTEGRATED.fasta`
|
|
79
|
+
- Contains the final consensus sequence after merging and reinserting insertions.
|
|
80
|
+
|
|
81
|
+
2. **Window QC trace (CSV)**
|
|
82
|
+
- File: `windows_trace.csv`
|
|
83
|
+
- One row per window, recording:
|
|
84
|
+
- `start`, `end` → genomic coordinates.
|
|
85
|
+
- `MISSING_MAPPING`, `MISSING_ABACAS` → counts of missing bases.
|
|
86
|
+
- `ABACAS_MORE_INFO` → whether ABACAS has fewer missing bases than MAPPING.
|
|
87
|
+
- `ABACAS_FRAGMENTS` → fragmentation level of ABACAS in this window (keep: 0 < n fragments < 3 ).
|
|
88
|
+
- `WINDOW_PRIOR_nLL_p95` → threshold from priors.
|
|
89
|
+
- `WINDOW_SCORE_nLL` → score of ABACAS in this window.
|
|
90
|
+
- `WINDOW_QC_PASSED` → True/False decision.
|
|
91
|
+
|
|
92
|
+
3. **Consensus QC summary (JSON)**
|
|
93
|
+
- File: `qc.json`
|
|
94
|
+
- Provides overall metrics comparing the MAPPING consensus and the integrated consensus:
|
|
95
|
+
- `MAPPING_COVERAGE` → % of genome covered in MAPPING.
|
|
96
|
+
- `FINAL_COVERAGE` → % of genome covered in integrated consensus.
|
|
97
|
+
- `MAPPING_SUBSTITUTIONS` → substitutions vs. reference in MAPPING.
|
|
98
|
+
- `FINAL_SUBSTITUTIONS` → substitutions vs. reference in integrated consensus.
|
|
99
|
+
- `EXPECTED_SUBSTITUTIONS` → expected number of substitutions, extrapolated from mapping.
|
|
100
|
+
- `OBS-EXP_SUBSTITUTIONS` → difference between observed and expected substitutions.
|
|
101
|
+
- `N_INSERTIONS` → number of insertions added back.
|
|
102
|
+
- `TOTAL_INSERTIONS_LENGTH` → total inserted length.
|
|
103
|
+
- `INSERTIONS` → list of insertions with their coordinates.
|
|
104
|
+
|
|
105
|
+
---
|
|
106
|
+
|
|
107
|
+
### ▶️ Example run
|
|
108
|
+
|
|
109
|
+
```bash
|
|
110
|
+
python integrate_consensus.py \
|
|
111
|
+
--input /path/to/<sample_name>.aln \
|
|
112
|
+
--ref RSV_BD \
|
|
113
|
+
--prior /path/to/RSVBD_win100_ovlp50_priors.parquet \
|
|
114
|
+
--output_dir results
|
|
115
|
+
```
|
|
116
|
+
----
|
|
117
|
+
|
|
118
|
+
This will generate:
|
|
119
|
+
|
|
120
|
+
- `results/<sample_name>-INTEGRATED.fasta`
|
|
121
|
+
- `results/windows_trace.csv`
|
|
122
|
+
- `results/qc.json`
|
|
123
|
+
|
|
124
|
+
---
|
|
125
|
+
|
|
126
|
+
## 🛠 Script: build_priors.py
|
|
127
|
+
|
|
128
|
+
This script creates **empirical priors** (overlapped windows) from a large multiple sequence alignment.
|
|
129
|
+
These priors are later used by `integrate_consensus.py` to evaluate windows.
|
|
130
|
+
|
|
131
|
+
### 🔑 Inputs
|
|
132
|
+
|
|
133
|
+
- `-i / --input` → aligned FASTA file with multiple sequences.
|
|
134
|
+
- `-r / --ref` → ID of the reference sequence.
|
|
135
|
+
- `-o / --output` → output file (`.parquet`).
|
|
136
|
+
- `--win` → window size (default: 100).
|
|
137
|
+
- `--overlap` → overlap size (default: 10).
|
|
138
|
+
|
|
139
|
+
### ▶️ Example run
|
|
140
|
+
|
|
141
|
+
```bash
|
|
142
|
+
python build_priors.py \
|
|
143
|
+
-i alignment.fasta \
|
|
144
|
+
-r ReferenceID \
|
|
145
|
+
-o priors.parquet \
|
|
146
|
+
--win 100 \
|
|
147
|
+
--overlap 10
|
|
148
|
+
```
|
|
149
|
+
----
|
|
150
|
+
|
|
151
|
+
### 📦 Output
|
|
152
|
+
|
|
153
|
+
A `.parquet` file with one row per window, containing:
|
|
154
|
+
|
|
155
|
+
- `start`, `end` → window coordinates.
|
|
156
|
+
- `nLL_p95`, `nLL_p99` → empirical thresholds.
|
|
157
|
+
- `profile` → base probability distributions for each position in the window.
|
|
158
|
+
|
|
159
|
+
## 🧮 Methodology (build_priors.py)
|
|
160
|
+
|
|
161
|
+
### 1. Probability distributions per position
|
|
162
|
+
|
|
163
|
+
For each window of size `W` bases (e.g., `W = 100`), and for each position `j` within that window, we compute the probability of observing each nucleotide:
|
|
164
|
+
|
|
165
|
+
=\frac{c_j(b)+\alpha}{\sum_{x\in\{A,C,G,T\}}(c_j(x)+\alpha)})
|
|
166
|
+
|
|
167
|
+
Where:
|
|
168
|
+
- ) = number of sequences with base 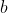 at position 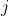.
|
|
169
|
+
- 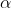 = pseudocount (Laplace smoothing, default ) to avoid zero probabilities.
|
|
170
|
+
- Bases `N` are ignored in the counts.
|
|
171
|
+
|
|
172
|
+
This gives a **per-position categorical distribution**.
|
|
173
|
+
|
|
174
|
+
---
|
|
175
|
+
|
|
176
|
+
### 2. Log-likelihood of a sequence in a window
|
|
177
|
+
|
|
178
|
+
Given a query sequence 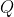, we compute its probability under the window profile.
|
|
179
|
+
For each valid (non-`N`) position 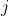 with observed base :
|
|
180
|
+
|
|
181
|
+
=\sum_{j=1}^{W}\log%20P_j(q_j))
|
|
182
|
+
|
|
183
|
+
The **normalized negative log-likelihood (nLL)** is:
|
|
184
|
+
|
|
185
|
+
=-\frac{1}{N_{\text{valid}}}\sum_{j=1}^{W}\log%20P_j(q_j))
|
|
186
|
+
|
|
187
|
+
Where:
|
|
188
|
+
-  = number of positions in the window where 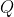 has a non-`N` base.
|
|
189
|
+
|
|
190
|
+
Smaller nLL values indicate sequences more likely under the empirical profile.
|
|
191
|
+
|
|
192
|
+
|
|
193
|
+
### 3. Empirical priors
|
|
194
|
+
|
|
195
|
+
To characterize "normal variation" for each window:
|
|
196
|
+
|
|
197
|
+
1. Score **all sequences** from the alignment against the window profile.
|
|
198
|
+
2. Collect the distribution of nLL values.
|
|
199
|
+
3. Extract percentiles (e.g., 95th and 99th) to serve as thresholds.
|
|
200
|
+
|
|
201
|
+
Thus, for each window we store:
|
|
202
|
+
- The **distribution (profile)**.
|
|
203
|
+
- Empirical thresholds: `nLL_p95` and `nLL_p99`.
|
|
204
|
+
|
|
205
|
+
A new sequence can later be compared:
|
|
206
|
+
- If `nLL < nLL_p95` → typical.
|
|
207
|
+
- If `nLL > nLL_p99` → unusually variable, possibly unreliable region.
|
|
208
|
+
|
|
209
|
+
---
|
|
210
|
+
|
|
211
|
+
---
|
|
212
|
+
|
|
213
|
+
## � Supporting utils
|
|
214
|
+
|
|
215
|
+
Several utility scripts provide reusable functions for both processes:
|
|
216
|
+
|
|
217
|
+
- **utils.py** → basic alignment and scoring functions:
|
|
218
|
+
- `load_alignment`, `extract_ref_positions`, `sliding_windows`, `score_window`.
|
|
219
|
+
|
|
220
|
+
- **utils_integrate_consensus.py** → additional helpers for consensus integration:
|
|
221
|
+
- missingness and fragmentation counts,
|
|
222
|
+
- insertion handling,
|
|
223
|
+
- QC calculations,
|
|
224
|
+
- consensus merging,
|
|
225
|
+
- window evaluation wrapper.
|
|
226
|
+
|
|
227
|
+
These modular functions keep the pipeline clean and reusable.
|
|
228
|
+
|
|
229
|
+
|
|
@@ -0,0 +1,13 @@
|
|
|
1
|
+
"""
|
|
2
|
+
priorcons package
|
|
3
|
+
Author: Germán Vallejo Palma
|
|
4
|
+
Developed at: Instituto de Salud Carlos III - National Centre of Microbiology
|
|
5
|
+
"""
|
|
6
|
+
|
|
7
|
+
__version__ = "0.1.0"
|
|
8
|
+
__author__ = "Germán Vallejo Palma"
|
|
9
|
+
__email__ = "german.vallejo@isciii.es"
|
|
10
|
+
|
|
11
|
+
# Expose key functions/modules for convenience
|
|
12
|
+
from .build_priors import main as build_priors_main
|
|
13
|
+
from .integrate_consensus import main as integrate_consensus_main
|
|
@@ -0,0 +1,39 @@
|
|
|
1
|
+
import argparse
|
|
2
|
+
import sys
|
|
3
|
+
from typing import List
|
|
4
|
+
from .utils import (
|
|
5
|
+
save_priors,
|
|
6
|
+
load_alignment,
|
|
7
|
+
build_window_profiles,
|
|
8
|
+
build_priors
|
|
9
|
+
)
|
|
10
|
+
|
|
11
|
+
def main(argv: List[str] = None) -> int:
|
|
12
|
+
parser = argparse.ArgumentParser(description="Build empirical priors from alignment")
|
|
13
|
+
parser.add_argument("-i", "--input", required=True, help="Input alignment FASTA file")
|
|
14
|
+
parser.add_argument("-r", "--ref", required=True, help="Reference sequence ID")
|
|
15
|
+
parser.add_argument("-o", "--output", required=True, help="Output file (.parquet)")
|
|
16
|
+
parser.add_argument("--win", type=int, default=100, help="Window size (default=100)")
|
|
17
|
+
parser.add_argument("--overlap", type=int, default=50, help="Window overlap (default=50)")
|
|
18
|
+
argv = argv if argv is not None else sys.argv[1:]
|
|
19
|
+
args = parser.parse_args(argv)
|
|
20
|
+
|
|
21
|
+
# Load alignment
|
|
22
|
+
ids, seqs = load_alignment(args.input)
|
|
23
|
+
|
|
24
|
+
# Build window profiles
|
|
25
|
+
profiles, seqs_filtered, ref_seq = build_window_profiles(
|
|
26
|
+
ids, seqs, args.ref, args.win, args.overlap
|
|
27
|
+
)
|
|
28
|
+
|
|
29
|
+
# Compute priors
|
|
30
|
+
df = build_priors(seqs_filtered, profiles)
|
|
31
|
+
|
|
32
|
+
# Save to Parquet
|
|
33
|
+
save_priors(df, args.output)
|
|
34
|
+
|
|
35
|
+
return 0
|
|
36
|
+
|
|
37
|
+
|
|
38
|
+
if __name__ == "__main__":
|
|
39
|
+
main()
|
|
@@ -0,0 +1,49 @@
|
|
|
1
|
+
#!/usr/bin/env python3
|
|
2
|
+
"""
|
|
3
|
+
priorCons CLI
|
|
4
|
+
"""
|
|
5
|
+
import sys
|
|
6
|
+
from . import __version__
|
|
7
|
+
from . import build_priors as bp
|
|
8
|
+
from . import integrate_consensus as ic
|
|
9
|
+
|
|
10
|
+
|
|
11
|
+
def main(argv=None):
|
|
12
|
+
argv = argv if argv is not None else sys.argv[1:]
|
|
13
|
+
|
|
14
|
+
# si el usuario no da subcomando o pide --help general
|
|
15
|
+
if len(argv) == 0 or argv[0] in ("-h", "--help"):
|
|
16
|
+
print(f"""
|
|
17
|
+
priorCons {__version__}
|
|
18
|
+
|
|
19
|
+
Usage:
|
|
20
|
+
priorcons <subcommand> [options]
|
|
21
|
+
|
|
22
|
+
Available subcommands:
|
|
23
|
+
build-priors Build priors parquet file
|
|
24
|
+
integrate-consensus Run consensus integration workflow
|
|
25
|
+
|
|
26
|
+
Use 'priorcons <subcommand> -h' for details on each one.
|
|
27
|
+
""")
|
|
28
|
+
sys.exit(0)
|
|
29
|
+
|
|
30
|
+
if argv[0] in ("--version", "-v", "-V"):
|
|
31
|
+
print(f"priorcons {__version__}")
|
|
32
|
+
sys.exit(0)
|
|
33
|
+
|
|
34
|
+
# delegar completamente al módulo correspondiente
|
|
35
|
+
subcmd = argv[0]
|
|
36
|
+
subargs = argv[1:]
|
|
37
|
+
|
|
38
|
+
if subcmd == "build-priors":
|
|
39
|
+
sys.exit(bp.main(subargs))
|
|
40
|
+
elif subcmd == "integrate-consensus":
|
|
41
|
+
sys.exit(ic.main(subargs))
|
|
42
|
+
else:
|
|
43
|
+
print(f"Unknown command: {subcmd}")
|
|
44
|
+
print("Use 'priorcons --help' for available commands.")
|
|
45
|
+
sys.exit(1)
|
|
46
|
+
|
|
47
|
+
|
|
48
|
+
if __name__ == "__main__":
|
|
49
|
+
main()
|