plotfig 1.18.0__tar.gz → 1.18.1__tar.gz
This diff represents the content of publicly available package versions that have been released to one of the supported registries. The information contained in this diff is provided for informational purposes only and reflects changes between package versions as they appear in their respective public registries.
- {plotfig-1.18.0 → plotfig-1.18.1}/.github/workflows/docs.yml +8 -6
- plotfig-1.18.1/.release-please-manifest.json +1 -0
- {plotfig-1.18.0 → plotfig-1.18.1}/CHANGELOG.md +7 -0
- {plotfig-1.18.0 → plotfig-1.18.1}/PKG-INFO +3 -5
- {plotfig-1.18.0 → plotfig-1.18.1}/README.md +2 -4
- {plotfig-1.18.0 → plotfig-1.18.1}/README_zh.md +1 -4
- {plotfig-1.18.0 → plotfig-1.18.1}/docs/en/index.md +2 -2
- {plotfig-1.18.0 → plotfig-1.18.1}/docs/en/usage/brain_connectivity.md +1 -1
- {plotfig-1.18.0 → plotfig-1.18.1}/docs/en/usage/brain_surface.md +14 -14
- {plotfig-1.18.0 → plotfig-1.18.1}/docs/en/usage/circos.md +7 -7
- {plotfig-1.18.0 → plotfig-1.18.1}/docs/en/usage/correlation.md +3 -3
- {plotfig-1.18.0 → plotfig-1.18.1}/docs/en/usage/matrix.md +2 -2
- {plotfig-1.18.0 → plotfig-1.18.1}/docs/en/usage/multi_groups.md +4 -4
- {plotfig-1.18.0 → plotfig-1.18.1}/docs/en/usage/single_group.md +17 -17
- plotfig-1.18.1/docs/zh/assets/atlas_csv/chimpanzee_bna.csv +201 -0
- plotfig-1.18.1/docs/zh/assets/atlas_csv/human_bna.csv +211 -0
- plotfig-1.18.1/docs/zh/assets/atlas_csv/human_glasser.csv +361 -0
- plotfig-1.18.1/docs/zh/assets/atlas_csv/macaque_bna.csv +249 -0
- plotfig-1.18.1/docs/zh/assets/atlas_csv/macaque_charm4.csv +113 -0
- plotfig-1.18.1/docs/zh/assets/atlas_csv/macaque_charm5.csv +177 -0
- plotfig-1.18.1/docs/zh/assets/atlas_csv/macaque_charm6.csv +269 -0
- plotfig-1.18.1/docs/zh/assets/atlas_csv/macaque_d99.csv +299 -0
- plotfig-1.18.1/docs/zh/assets/plotfig.png +0 -0
- plotfig-1.18.1/docs/zh/assets/usage/brain_connectivity_files/output.gif +0 -0
- plotfig-1.18.1/docs/zh/assets/usage/brain_surface_files/brain_surface_10_0.png +0 -0
- plotfig-1.18.1/docs/zh/assets/usage/brain_surface_files/brain_surface_11_0.png +0 -0
- plotfig-1.18.1/docs/zh/assets/usage/brain_surface_files/brain_surface_12_0.png +0 -0
- plotfig-1.18.1/docs/zh/assets/usage/brain_surface_files/brain_surface_13_0.png +0 -0
- plotfig-1.18.1/docs/zh/assets/usage/brain_surface_files/brain_surface_15_0.png +0 -0
- plotfig-1.18.1/docs/zh/assets/usage/brain_surface_files/brain_surface_16_0.png +0 -0
- plotfig-1.18.1/docs/zh/assets/usage/brain_surface_files/brain_surface_4_0.png +0 -0
- plotfig-1.18.1/docs/zh/assets/usage/brain_surface_files/brain_surface_5_0.png +0 -0
- plotfig-1.18.1/docs/zh/assets/usage/brain_surface_files/brain_surface_6_0.png +0 -0
- plotfig-1.18.1/docs/zh/assets/usage/brain_surface_files/brain_surface_8_0.png +0 -0
- plotfig-1.18.1/docs/zh/assets/usage/brain_surface_files/brain_surface_9_0.png +0 -0
- plotfig-1.18.1/docs/zh/assets/usage/circos_files/circos_11_0.png +0 -0
- plotfig-1.18.1/docs/zh/assets/usage/circos_files/circos_13_1.png +0 -0
- plotfig-1.18.1/docs/zh/assets/usage/circos_files/circos_1_0.png +0 -0
- plotfig-1.18.1/docs/zh/assets/usage/circos_files/circos_3_0.png +0 -0
- plotfig-1.18.1/docs/zh/assets/usage/circos_files/circos_5_0.png +0 -0
- plotfig-1.18.1/docs/zh/assets/usage/circos_files/circos_7_0.png +0 -0
- plotfig-1.18.1/docs/zh/assets/usage/circos_files/circos_9_0.png +0 -0
- plotfig-1.18.1/docs/zh/assets/usage/correlation_files/correlation_10_0.png +0 -0
- plotfig-1.18.1/docs/zh/assets/usage/correlation_files/correlation_4_0.png +0 -0
- plotfig-1.18.1/docs/zh/assets/usage/correlation_files/correlation_7_0.png +0 -0
- plotfig-1.18.1/docs/zh/assets/usage/matrix_files/matrix_2_1.png +0 -0
- plotfig-1.18.1/docs/zh/assets/usage/matrix_files/matrix_3_0.png +0 -0
- plotfig-1.18.1/docs/zh/assets/usage/matrix_files/matrix_4_0.png +0 -0
- plotfig-1.18.1/docs/zh/assets/usage/matrix_files/matrix_5_1.png +0 -0
- plotfig-1.18.1/docs/zh/assets/usage/matrix_files/matrix_6_0.png +0 -0
- plotfig-1.18.1/docs/zh/assets/usage/matrix_files/matrix_7_0.png +0 -0
- plotfig-1.18.1/docs/zh/assets/usage/multi_groups_files/multi_groups_2_0.png +0 -0
- plotfig-1.18.1/docs/zh/assets/usage/multi_groups_files/multi_groups_3_0.png +0 -0
- plotfig-1.18.1/docs/zh/assets/usage/multi_groups_files/multi_groups_5_0.png +0 -0
- plotfig-1.18.1/docs/zh/assets/usage/multi_groups_files/multi_groups_6_0.png +0 -0
- plotfig-1.18.1/docs/zh/assets/usage/multi_groups_files/multi_groups_8_0.png +0 -0
- plotfig-1.18.1/docs/zh/assets/usage/multi_groups_files/multi_groups_9_0.png +0 -0
- plotfig-1.18.1/docs/zh/assets/usage/single_group_files/single_group_12_0.png +0 -0
- plotfig-1.18.1/docs/zh/assets/usage/single_group_files/single_group_14_0.png +0 -0
- plotfig-1.18.1/docs/zh/assets/usage/single_group_files/single_group_17_0.png +0 -0
- plotfig-1.18.1/docs/zh/assets/usage/single_group_files/single_group_20_0.png +0 -0
- plotfig-1.18.1/docs/zh/assets/usage/single_group_files/single_group_22_0.png +0 -0
- plotfig-1.18.1/docs/zh/assets/usage/single_group_files/single_group_24_0.png +0 -0
- plotfig-1.18.1/docs/zh/assets/usage/single_group_files/single_group_26_0.png +0 -0
- plotfig-1.18.1/docs/zh/assets/usage/single_group_files/single_group_28_0.png +0 -0
- plotfig-1.18.1/docs/zh/assets/usage/single_group_files/single_group_31_0.png +0 -0
- plotfig-1.18.1/docs/zh/assets/usage/single_group_files/single_group_34_0.png +0 -0
- plotfig-1.18.1/docs/zh/assets/usage/single_group_files/single_group_36_0.png +0 -0
- plotfig-1.18.1/docs/zh/assets/usage/single_group_files/single_group_38_0.png +0 -0
- plotfig-1.18.1/docs/zh/assets/usage/single_group_files/single_group_39_0.png +0 -0
- plotfig-1.18.1/docs/zh/assets/usage/single_group_files/single_group_3_0.png +0 -0
- plotfig-1.18.1/docs/zh/assets/usage/single_group_files/single_group_41_0.png +0 -0
- plotfig-1.18.1/docs/zh/assets/usage/single_group_files/single_group_6_0.png +0 -0
- plotfig-1.18.1/docs/zh/assets/usage/single_group_files/single_group_8_0.png +0 -0
- {plotfig-1.18.0 → plotfig-1.18.1}/docs/zh/index.md +1 -1
- {plotfig-1.18.0 → plotfig-1.18.1}/docs/zh/usage/brain_connectivity.md +1 -1
- {plotfig-1.18.0 → plotfig-1.18.1}/docs/zh/usage/brain_surface.md +14 -14
- {plotfig-1.18.0 → plotfig-1.18.1}/docs/zh/usage/circos.md +215 -216
- {plotfig-1.18.0 → plotfig-1.18.1}/docs/zh/usage/correlation.md +3 -3
- {plotfig-1.18.0 → plotfig-1.18.1}/docs/zh/usage/matrix.md +56 -57
- {plotfig-1.18.0 → plotfig-1.18.1}/docs/zh/usage/multi_groups.md +4 -4
- {plotfig-1.18.0 → plotfig-1.18.1}/docs/zh/usage/single_group.md +17 -17
- {plotfig-1.18.0 → plotfig-1.18.1}/pyproject.toml +2 -2
- {plotfig-1.18.0 → plotfig-1.18.1}/src/plotfig/single_bar.py +24 -16
- {plotfig-1.18.0 → plotfig-1.18.1}/uv.lock +79 -24
- plotfig-1.18.1/zensical.toml +133 -0
- plotfig-1.18.0/zensical.toml → plotfig-1.18.1/zensical.zh.toml +27 -43
- plotfig-1.18.0/.release-please-manifest.json +0 -1
- {plotfig-1.18.0 → plotfig-1.18.1}/.github/workflows/dependency_review.yml +0 -0
- {plotfig-1.18.0 → plotfig-1.18.1}/.github/workflows/python_publish.yml +0 -0
- {plotfig-1.18.0 → plotfig-1.18.1}/.github/workflows/release-please.yml +0 -0
- {plotfig-1.18.0 → plotfig-1.18.1}/.github/workflows/sync_to_gitee.yml +0 -0
- {plotfig-1.18.0 → plotfig-1.18.1}/.gitignore +0 -0
- {plotfig-1.18.0 → plotfig-1.18.1}/.python-version +0 -0
- {plotfig-1.18.0 → plotfig-1.18.1}/LICENSE +0 -0
- {plotfig-1.18.0 → plotfig-1.18.1}/docs/en/api/index.md +0 -0
- {plotfig-1.18.0/docs → plotfig-1.18.1/docs/en}/assets/atlas_csv/chimpanzee_bna.csv +0 -0
- {plotfig-1.18.0/docs → plotfig-1.18.1/docs/en}/assets/atlas_csv/human_bna.csv +0 -0
- {plotfig-1.18.0/docs → plotfig-1.18.1/docs/en}/assets/atlas_csv/human_glasser.csv +0 -0
- {plotfig-1.18.0/docs → plotfig-1.18.1/docs/en}/assets/atlas_csv/macaque_bna.csv +0 -0
- {plotfig-1.18.0/docs → plotfig-1.18.1/docs/en}/assets/atlas_csv/macaque_charm4.csv +0 -0
- {plotfig-1.18.0/docs → plotfig-1.18.1/docs/en}/assets/atlas_csv/macaque_charm5.csv +0 -0
- {plotfig-1.18.0/docs → plotfig-1.18.1/docs/en}/assets/atlas_csv/macaque_charm6.csv +0 -0
- {plotfig-1.18.0/docs → plotfig-1.18.1/docs/en}/assets/atlas_csv/macaque_d99.csv +0 -0
- {plotfig-1.18.0/docs → plotfig-1.18.1/docs/en}/assets/plotfig.png +0 -0
- {plotfig-1.18.0/docs → plotfig-1.18.1/docs/en}/assets/usage/brain_connectivity_files/output.gif +0 -0
- {plotfig-1.18.0/docs → plotfig-1.18.1/docs/en}/assets/usage/brain_surface_files/brain_surface_10_0.png +0 -0
- {plotfig-1.18.0/docs → plotfig-1.18.1/docs/en}/assets/usage/brain_surface_files/brain_surface_11_0.png +0 -0
- {plotfig-1.18.0/docs → plotfig-1.18.1/docs/en}/assets/usage/brain_surface_files/brain_surface_12_0.png +0 -0
- {plotfig-1.18.0/docs → plotfig-1.18.1/docs/en}/assets/usage/brain_surface_files/brain_surface_13_0.png +0 -0
- {plotfig-1.18.0/docs → plotfig-1.18.1/docs/en}/assets/usage/brain_surface_files/brain_surface_15_0.png +0 -0
- {plotfig-1.18.0/docs → plotfig-1.18.1/docs/en}/assets/usage/brain_surface_files/brain_surface_16_0.png +0 -0
- {plotfig-1.18.0/docs → plotfig-1.18.1/docs/en}/assets/usage/brain_surface_files/brain_surface_4_0.png +0 -0
- {plotfig-1.18.0/docs → plotfig-1.18.1/docs/en}/assets/usage/brain_surface_files/brain_surface_5_0.png +0 -0
- {plotfig-1.18.0/docs → plotfig-1.18.1/docs/en}/assets/usage/brain_surface_files/brain_surface_6_0.png +0 -0
- {plotfig-1.18.0/docs → plotfig-1.18.1/docs/en}/assets/usage/brain_surface_files/brain_surface_8_0.png +0 -0
- {plotfig-1.18.0/docs → plotfig-1.18.1/docs/en}/assets/usage/brain_surface_files/brain_surface_9_0.png +0 -0
- {plotfig-1.18.0/docs → plotfig-1.18.1/docs/en}/assets/usage/circos_files/circos_11_0.png +0 -0
- {plotfig-1.18.0/docs → plotfig-1.18.1/docs/en}/assets/usage/circos_files/circos_13_1.png +0 -0
- {plotfig-1.18.0/docs → plotfig-1.18.1/docs/en}/assets/usage/circos_files/circos_1_0.png +0 -0
- {plotfig-1.18.0/docs → plotfig-1.18.1/docs/en}/assets/usage/circos_files/circos_3_0.png +0 -0
- {plotfig-1.18.0/docs → plotfig-1.18.1/docs/en}/assets/usage/circos_files/circos_5_0.png +0 -0
- {plotfig-1.18.0/docs → plotfig-1.18.1/docs/en}/assets/usage/circos_files/circos_7_0.png +0 -0
- {plotfig-1.18.0/docs → plotfig-1.18.1/docs/en}/assets/usage/circos_files/circos_9_0.png +0 -0
- {plotfig-1.18.0/docs → plotfig-1.18.1/docs/en}/assets/usage/correlation_files/correlation_10_0.png +0 -0
- {plotfig-1.18.0/docs → plotfig-1.18.1/docs/en}/assets/usage/correlation_files/correlation_4_0.png +0 -0
- {plotfig-1.18.0/docs → plotfig-1.18.1/docs/en}/assets/usage/correlation_files/correlation_7_0.png +0 -0
- {plotfig-1.18.0/docs → plotfig-1.18.1/docs/en}/assets/usage/matrix_files/matrix_2_1.png +0 -0
- {plotfig-1.18.0/docs → plotfig-1.18.1/docs/en}/assets/usage/matrix_files/matrix_3_0.png +0 -0
- {plotfig-1.18.0/docs → plotfig-1.18.1/docs/en}/assets/usage/matrix_files/matrix_4_0.png +0 -0
- {plotfig-1.18.0/docs → plotfig-1.18.1/docs/en}/assets/usage/matrix_files/matrix_5_1.png +0 -0
- {plotfig-1.18.0/docs → plotfig-1.18.1/docs/en}/assets/usage/matrix_files/matrix_6_0.png +0 -0
- {plotfig-1.18.0/docs → plotfig-1.18.1/docs/en}/assets/usage/matrix_files/matrix_7_0.png +0 -0
- {plotfig-1.18.0/docs → plotfig-1.18.1/docs/en}/assets/usage/multi_groups_files/multi_groups_2_0.png +0 -0
- {plotfig-1.18.0/docs → plotfig-1.18.1/docs/en}/assets/usage/multi_groups_files/multi_groups_3_0.png +0 -0
- {plotfig-1.18.0/docs → plotfig-1.18.1/docs/en}/assets/usage/multi_groups_files/multi_groups_5_0.png +0 -0
- {plotfig-1.18.0/docs → plotfig-1.18.1/docs/en}/assets/usage/multi_groups_files/multi_groups_6_0.png +0 -0
- {plotfig-1.18.0/docs → plotfig-1.18.1/docs/en}/assets/usage/multi_groups_files/multi_groups_8_0.png +0 -0
- {plotfig-1.18.0/docs → plotfig-1.18.1/docs/en}/assets/usage/multi_groups_files/multi_groups_9_0.png +0 -0
- {plotfig-1.18.0/docs → plotfig-1.18.1/docs/en}/assets/usage/single_group_files/single_group_12_0.png +0 -0
- {plotfig-1.18.0/docs → plotfig-1.18.1/docs/en}/assets/usage/single_group_files/single_group_14_0.png +0 -0
- {plotfig-1.18.0/docs → plotfig-1.18.1/docs/en}/assets/usage/single_group_files/single_group_17_0.png +0 -0
- {plotfig-1.18.0/docs → plotfig-1.18.1/docs/en}/assets/usage/single_group_files/single_group_20_0.png +0 -0
- {plotfig-1.18.0/docs → plotfig-1.18.1/docs/en}/assets/usage/single_group_files/single_group_22_0.png +0 -0
- {plotfig-1.18.0/docs → plotfig-1.18.1/docs/en}/assets/usage/single_group_files/single_group_24_0.png +0 -0
- {plotfig-1.18.0/docs → plotfig-1.18.1/docs/en}/assets/usage/single_group_files/single_group_26_0.png +0 -0
- {plotfig-1.18.0/docs → plotfig-1.18.1/docs/en}/assets/usage/single_group_files/single_group_28_0.png +0 -0
- {plotfig-1.18.0/docs → plotfig-1.18.1/docs/en}/assets/usage/single_group_files/single_group_31_0.png +0 -0
- {plotfig-1.18.0/docs → plotfig-1.18.1/docs/en}/assets/usage/single_group_files/single_group_34_0.png +0 -0
- {plotfig-1.18.0/docs → plotfig-1.18.1/docs/en}/assets/usage/single_group_files/single_group_36_0.png +0 -0
- {plotfig-1.18.0/docs → plotfig-1.18.1/docs/en}/assets/usage/single_group_files/single_group_38_0.png +0 -0
- {plotfig-1.18.0/docs → plotfig-1.18.1/docs/en}/assets/usage/single_group_files/single_group_39_0.png +0 -0
- {plotfig-1.18.0/docs → plotfig-1.18.1/docs/en}/assets/usage/single_group_files/single_group_3_0.png +0 -0
- {plotfig-1.18.0/docs → plotfig-1.18.1/docs/en}/assets/usage/single_group_files/single_group_41_0.png +0 -0
- {plotfig-1.18.0/docs → plotfig-1.18.1/docs/en}/assets/usage/single_group_files/single_group_6_0.png +0 -0
- {plotfig-1.18.0/docs → plotfig-1.18.1/docs/en}/assets/usage/single_group_files/single_group_8_0.png +0 -0
- {plotfig-1.18.0 → plotfig-1.18.1}/docs/en/changelog.md +0 -0
- {plotfig-1.18.0 → plotfig-1.18.1}/docs/en/installation.md +0 -0
- {plotfig-1.18.0 → plotfig-1.18.1}/docs/zh/api/index.md +0 -0
- {plotfig-1.18.0 → plotfig-1.18.1}/docs/zh/changelog.md +0 -0
- {plotfig-1.18.0 → plotfig-1.18.1}/docs/zh/installation.md +0 -0
- {plotfig-1.18.0 → plotfig-1.18.1}/notebooks/brain_connectivity.ipynb +0 -0
- {plotfig-1.18.0 → plotfig-1.18.1}/notebooks/brain_surface.ipynb +0 -0
- {plotfig-1.18.0 → plotfig-1.18.1}/notebooks/circos.ipynb +0 -0
- {plotfig-1.18.0 → plotfig-1.18.1}/notebooks/correlation.ipynb +0 -0
- {plotfig-1.18.0 → plotfig-1.18.1}/notebooks/matrix.ipynb +0 -0
- {plotfig-1.18.0 → plotfig-1.18.1}/notebooks/multi_groups.ipynb +0 -0
- {plotfig-1.18.0 → plotfig-1.18.1}/notebooks/single_group.ipynb +0 -0
- {plotfig-1.18.0 → plotfig-1.18.1}/release-please-config.json +0 -0
- {plotfig-1.18.0 → plotfig-1.18.1}/src/plotfig/__init__.py +0 -0
- {plotfig-1.18.0 → plotfig-1.18.1}/src/plotfig/brain_connection.py +0 -0
- {plotfig-1.18.0 → plotfig-1.18.1}/src/plotfig/brain_surface.py +0 -0
- {plotfig-1.18.0 → plotfig-1.18.1}/src/plotfig/circos.py +0 -0
- {plotfig-1.18.0 → plotfig-1.18.1}/src/plotfig/correlation.py +0 -0
- {plotfig-1.18.0 → plotfig-1.18.1}/src/plotfig/data/atlas_tables/chimpanzee_bna.csv +0 -0
- {plotfig-1.18.0 → plotfig-1.18.1}/src/plotfig/data/atlas_tables/human_bna.csv +0 -0
- {plotfig-1.18.0 → plotfig-1.18.1}/src/plotfig/data/atlas_tables/human_glasser.csv +0 -0
- {plotfig-1.18.0 → plotfig-1.18.1}/src/plotfig/data/atlas_tables/macaque_bna.csv +0 -0
- {plotfig-1.18.0 → plotfig-1.18.1}/src/plotfig/data/atlas_tables/macaque_charm4.csv +0 -0
- {plotfig-1.18.0 → plotfig-1.18.1}/src/plotfig/data/atlas_tables/macaque_charm5.csv +0 -0
- {plotfig-1.18.0 → plotfig-1.18.1}/src/plotfig/data/atlas_tables/macaque_charm5_add_13_sgms.csv +0 -0
- {plotfig-1.18.0 → plotfig-1.18.1}/src/plotfig/data/atlas_tables/macaque_charm6.csv +0 -0
- {plotfig-1.18.0 → plotfig-1.18.1}/src/plotfig/data/atlas_tables/macaque_d99.csv +0 -0
- {plotfig-1.18.0 → plotfig-1.18.1}/src/plotfig/data/neurodata/atlases/chimpanzee_BNA/ChimpBNA.L.32k_fs_LR.label.gii +0 -0
- {plotfig-1.18.0 → plotfig-1.18.1}/src/plotfig/data/neurodata/atlases/chimpanzee_BNA/ChimpBNA.R.32k_fs_LR.label.gii +0 -0
- {plotfig-1.18.0 → plotfig-1.18.1}/src/plotfig/data/neurodata/atlases/human_BNA/fsaverage.L.BNA.32k_fs_LR.label.gii +0 -0
- {plotfig-1.18.0 → plotfig-1.18.1}/src/plotfig/data/neurodata/atlases/human_BNA/fsaverage.R.BNA.32k_fs_LR.label.gii +0 -0
- {plotfig-1.18.0 → plotfig-1.18.1}/src/plotfig/data/neurodata/atlases/human_Glasser/fsaverage.L.Glasser.32k_fs_LR.label.gii +0 -0
- {plotfig-1.18.0 → plotfig-1.18.1}/src/plotfig/data/neurodata/atlases/human_Glasser/fsaverage.R.Glasser.32k_fs_LR.label.gii +0 -0
- {plotfig-1.18.0 → plotfig-1.18.1}/src/plotfig/data/neurodata/atlases/macaque_BNA/MBNA_124_32k_L.label.gii +0 -0
- {plotfig-1.18.0 → plotfig-1.18.1}/src/plotfig/data/neurodata/atlases/macaque_BNA/MBNA_124_32k_R.label.gii +0 -0
- {plotfig-1.18.0 → plotfig-1.18.1}/src/plotfig/data/neurodata/atlases/macaque_CHARM4/L.charm4.label.gii +0 -0
- {plotfig-1.18.0 → plotfig-1.18.1}/src/plotfig/data/neurodata/atlases/macaque_CHARM4/R.charm4.label.gii +0 -0
- {plotfig-1.18.0 → plotfig-1.18.1}/src/plotfig/data/neurodata/atlases/macaque_CHARM5/L.charm5.label.gii +0 -0
- {plotfig-1.18.0 → plotfig-1.18.1}/src/plotfig/data/neurodata/atlases/macaque_CHARM5/R.charm5.label.gii +0 -0
- {plotfig-1.18.0 → plotfig-1.18.1}/src/plotfig/data/neurodata/atlases/macaque_CHARM6/L.charm6.label.gii +0 -0
- {plotfig-1.18.0 → plotfig-1.18.1}/src/plotfig/data/neurodata/atlases/macaque_CHARM6/R.charm6.label.gii +0 -0
- {plotfig-1.18.0 → plotfig-1.18.1}/src/plotfig/data/neurodata/atlases/macaque_D99/L.d99.label.gii +0 -0
- {plotfig-1.18.0 → plotfig-1.18.1}/src/plotfig/data/neurodata/atlases/macaque_D99/R.d99.label.gii +0 -0
- {plotfig-1.18.0 → plotfig-1.18.1}/src/plotfig/data/neurodata/surfaces/chimpanzee_BNA/ChimpYerkes29_v1.2.L.midthickness.32k_fs_LR.surf.gii +0 -0
- {plotfig-1.18.0 → plotfig-1.18.1}/src/plotfig/data/neurodata/surfaces/chimpanzee_BNA/ChimpYerkes29_v1.2.L.veryinflated.32k_fs_LR.surf.gii +0 -0
- {plotfig-1.18.0 → plotfig-1.18.1}/src/plotfig/data/neurodata/surfaces/chimpanzee_BNA/ChimpYerkes29_v1.2.R.midthickness.32k_fs_LR.surf.gii +0 -0
- {plotfig-1.18.0 → plotfig-1.18.1}/src/plotfig/data/neurodata/surfaces/chimpanzee_BNA/ChimpYerkes29_v1.2.R.veryinflated.32k_fs_LR.surf.gii +0 -0
- {plotfig-1.18.0 → plotfig-1.18.1}/src/plotfig/data/neurodata/surfaces/human_fsLR/100206.L.sulc.32k_fs_LR.shape.gii +0 -0
- {plotfig-1.18.0 → plotfig-1.18.1}/src/plotfig/data/neurodata/surfaces/human_fsLR/100206.R.sulc.32k_fs_LR.shape.gii +0 -0
- {plotfig-1.18.0 → plotfig-1.18.1}/src/plotfig/data/neurodata/surfaces/human_fsLR/README.md +0 -0
- {plotfig-1.18.0 → plotfig-1.18.1}/src/plotfig/data/neurodata/surfaces/human_fsLR/tpl-fsLR_den-32k_hemi-L_desc-nomedialwall_dparc.label.gii +0 -0
- {plotfig-1.18.0 → plotfig-1.18.1}/src/plotfig/data/neurodata/surfaces/human_fsLR/tpl-fsLR_den-32k_hemi-L_desc-sulc_midthickness.shape.gii +0 -0
- {plotfig-1.18.0 → plotfig-1.18.1}/src/plotfig/data/neurodata/surfaces/human_fsLR/tpl-fsLR_den-32k_hemi-L_desc-vaavg_midthickness.shape.gii +0 -0
- {plotfig-1.18.0 → plotfig-1.18.1}/src/plotfig/data/neurodata/surfaces/human_fsLR/tpl-fsLR_den-32k_hemi-L_flat.surf.gii +0 -0
- {plotfig-1.18.0 → plotfig-1.18.1}/src/plotfig/data/neurodata/surfaces/human_fsLR/tpl-fsLR_den-32k_hemi-L_inflated.surf.gii +0 -0
- {plotfig-1.18.0 → plotfig-1.18.1}/src/plotfig/data/neurodata/surfaces/human_fsLR/tpl-fsLR_den-32k_hemi-L_midthickness.surf.gii +0 -0
- {plotfig-1.18.0 → plotfig-1.18.1}/src/plotfig/data/neurodata/surfaces/human_fsLR/tpl-fsLR_den-32k_hemi-L_sphere.surf.gii +0 -0
- {plotfig-1.18.0 → plotfig-1.18.1}/src/plotfig/data/neurodata/surfaces/human_fsLR/tpl-fsLR_den-32k_hemi-L_veryinflated.surf.gii +0 -0
- {plotfig-1.18.0 → plotfig-1.18.1}/src/plotfig/data/neurodata/surfaces/human_fsLR/tpl-fsLR_den-32k_hemi-R_desc-nomedialwall_dparc.label.gii +0 -0
- {plotfig-1.18.0 → plotfig-1.18.1}/src/plotfig/data/neurodata/surfaces/human_fsLR/tpl-fsLR_den-32k_hemi-R_desc-sulc_midthickness.shape.gii +0 -0
- {plotfig-1.18.0 → plotfig-1.18.1}/src/plotfig/data/neurodata/surfaces/human_fsLR/tpl-fsLR_den-32k_hemi-R_desc-vaavg_midthickness.shape.gii +0 -0
- {plotfig-1.18.0 → plotfig-1.18.1}/src/plotfig/data/neurodata/surfaces/human_fsLR/tpl-fsLR_den-32k_hemi-R_flat.surf.gii +0 -0
- {plotfig-1.18.0 → plotfig-1.18.1}/src/plotfig/data/neurodata/surfaces/human_fsLR/tpl-fsLR_den-32k_hemi-R_inflated.surf.gii +0 -0
- {plotfig-1.18.0 → plotfig-1.18.1}/src/plotfig/data/neurodata/surfaces/human_fsLR/tpl-fsLR_den-32k_hemi-R_midthickness.surf.gii +0 -0
- {plotfig-1.18.0 → plotfig-1.18.1}/src/plotfig/data/neurodata/surfaces/human_fsLR/tpl-fsLR_den-32k_hemi-R_sphere.surf.gii +0 -0
- {plotfig-1.18.0 → plotfig-1.18.1}/src/plotfig/data/neurodata/surfaces/human_fsLR/tpl-fsLR_den-32k_hemi-R_veryinflated.surf.gii +0 -0
- {plotfig-1.18.0 → plotfig-1.18.1}/src/plotfig/data/neurodata/surfaces/human_fsLR/tpl-fsLR_space-fsaverage_den-32k_hemi-L_sphere.surf.gii +0 -0
- {plotfig-1.18.0 → plotfig-1.18.1}/src/plotfig/data/neurodata/surfaces/human_fsLR/tpl-fsLR_space-fsaverage_den-32k_hemi-R_sphere.surf.gii +0 -0
- {plotfig-1.18.0 → plotfig-1.18.1}/src/plotfig/data/neurodata/surfaces/macaque_BNA/README.md +0 -0
- {plotfig-1.18.0 → plotfig-1.18.1}/src/plotfig/data/neurodata/surfaces/macaque_BNA/SC_06018.L.sulc.32k_fs_LR.shape.gii +0 -0
- {plotfig-1.18.0 → plotfig-1.18.1}/src/plotfig/data/neurodata/surfaces/macaque_BNA/SC_06018.R.sulc.32k_fs_LR.shape.gii +0 -0
- {plotfig-1.18.0 → plotfig-1.18.1}/src/plotfig/data/neurodata/surfaces/macaque_BNA/civm.L.flat.32k_fs_LR.surf.gii +0 -0
- {plotfig-1.18.0 → plotfig-1.18.1}/src/plotfig/data/neurodata/surfaces/macaque_BNA/civm.L.inflated.32k_fs_LR.surf.gii +0 -0
- {plotfig-1.18.0 → plotfig-1.18.1}/src/plotfig/data/neurodata/surfaces/macaque_BNA/civm.L.midthickness.32k_fs_LR.surf.gii +0 -0
- {plotfig-1.18.0 → plotfig-1.18.1}/src/plotfig/data/neurodata/surfaces/macaque_BNA/civm.L.pial.32k_fs_LR.surf.gii +0 -0
- {plotfig-1.18.0 → plotfig-1.18.1}/src/plotfig/data/neurodata/surfaces/macaque_BNA/civm.L.veryinflated.32k_fs_LR.surf.gii +0 -0
- {plotfig-1.18.0 → plotfig-1.18.1}/src/plotfig/data/neurodata/surfaces/macaque_BNA/civm.L.white.32k_fs_LR.surf.gii +0 -0
- {plotfig-1.18.0 → plotfig-1.18.1}/src/plotfig/data/neurodata/surfaces/macaque_BNA/civm.R.flat.32k_fs_LR.surf.gii +0 -0
- {plotfig-1.18.0 → plotfig-1.18.1}/src/plotfig/data/neurodata/surfaces/macaque_BNA/civm.R.inflated.32k_fs_LR.surf.gii +0 -0
- {plotfig-1.18.0 → plotfig-1.18.1}/src/plotfig/data/neurodata/surfaces/macaque_BNA/civm.R.midthickness.32k_fs_LR.surf.gii +0 -0
- {plotfig-1.18.0 → plotfig-1.18.1}/src/plotfig/data/neurodata/surfaces/macaque_BNA/civm.R.pial.32k_fs_LR.surf.gii +0 -0
- {plotfig-1.18.0 → plotfig-1.18.1}/src/plotfig/data/neurodata/surfaces/macaque_BNA/civm.R.veryinflated.32k_fs_LR.surf.gii +0 -0
- {plotfig-1.18.0 → plotfig-1.18.1}/src/plotfig/data/neurodata/surfaces/macaque_BNA/civm.R.white.32k_fs_LR.surf.gii +0 -0
- {plotfig-1.18.0 → plotfig-1.18.1}/src/plotfig/data/neurodata/surfaces/macaque_NMT2/L.gray_surface.inf_300.surf.gii +0 -0
- {plotfig-1.18.0 → plotfig-1.18.1}/src/plotfig/data/neurodata/surfaces/macaque_NMT2/L.gray_surface.surf.gii +0 -0
- {plotfig-1.18.0 → plotfig-1.18.1}/src/plotfig/data/neurodata/surfaces/macaque_NMT2/L.mid_surface.inf_300.surf.gii +0 -0
- {plotfig-1.18.0 → plotfig-1.18.1}/src/plotfig/data/neurodata/surfaces/macaque_NMT2/L.mid_surface.surf.gii +0 -0
- {plotfig-1.18.0 → plotfig-1.18.1}/src/plotfig/data/neurodata/surfaces/macaque_NMT2/L.white_surface.inf_300.surf.gii +0 -0
- {plotfig-1.18.0 → plotfig-1.18.1}/src/plotfig/data/neurodata/surfaces/macaque_NMT2/L.white_surface.surf.gii +0 -0
- {plotfig-1.18.0 → plotfig-1.18.1}/src/plotfig/data/neurodata/surfaces/macaque_NMT2/R.gray_surface.inf_300.surf.gii +0 -0
- {plotfig-1.18.0 → plotfig-1.18.1}/src/plotfig/data/neurodata/surfaces/macaque_NMT2/R.gray_surface.surf.gii +0 -0
- {plotfig-1.18.0 → plotfig-1.18.1}/src/plotfig/data/neurodata/surfaces/macaque_NMT2/R.mid_surface.inf_300.surf.gii +0 -0
- {plotfig-1.18.0 → plotfig-1.18.1}/src/plotfig/data/neurodata/surfaces/macaque_NMT2/R.mid_surface.surf.gii +0 -0
- {plotfig-1.18.0 → plotfig-1.18.1}/src/plotfig/data/neurodata/surfaces/macaque_NMT2/R.white_surface.inf_300.surf.gii +0 -0
- {plotfig-1.18.0 → plotfig-1.18.1}/src/plotfig/data/neurodata/surfaces/macaque_NMT2/R.white_surface.surf.gii +0 -0
- {plotfig-1.18.0 → plotfig-1.18.1}/src/plotfig/data/neurodata/volumes/macaque_NMT2/CHARM5_add_13_sgms_asym.nii.gz +0 -0
- {plotfig-1.18.0 → plotfig-1.18.1}/src/plotfig/externals/surfplot/__init__.py +0 -0
- {plotfig-1.18.0 → plotfig-1.18.1}/src/plotfig/externals/surfplot/data/lh_default_mode_example.tsv +0 -0
- {plotfig-1.18.0 → plotfig-1.18.1}/src/plotfig/externals/surfplot/data/lh_frontoparietal_example.tsv +0 -0
- {plotfig-1.18.0 → plotfig-1.18.1}/src/plotfig/externals/surfplot/data/medwall.tsv +0 -0
- {plotfig-1.18.0 → plotfig-1.18.1}/src/plotfig/externals/surfplot/data/rh_default_mode_example.tsv +0 -0
- {plotfig-1.18.0 → plotfig-1.18.1}/src/plotfig/externals/surfplot/data/rh_frontoparietal_example.tsv +0 -0
- {plotfig-1.18.0 → plotfig-1.18.1}/src/plotfig/externals/surfplot/datasets.py +0 -0
- {plotfig-1.18.0 → plotfig-1.18.1}/src/plotfig/externals/surfplot/plotting.py +0 -0
- {plotfig-1.18.0 → plotfig-1.18.1}/src/plotfig/externals/surfplot/surf.py +0 -0
- {plotfig-1.18.0 → plotfig-1.18.1}/src/plotfig/externals/surfplot/utils.py +0 -0
- {plotfig-1.18.0 → plotfig-1.18.1}/src/plotfig/matrix.py +0 -0
- {plotfig-1.18.0 → plotfig-1.18.1}/src/plotfig/multi_bars.py +0 -0
- {plotfig-1.18.0 → plotfig-1.18.1}/src/plotfig/utils/__init__.py +0 -0
- {plotfig-1.18.0 → plotfig-1.18.1}/src/plotfig/utils/bar.py +0 -0
- {plotfig-1.18.0 → plotfig-1.18.1}/src/plotfig/utils/color.py +0 -0
- {plotfig-1.18.0 → plotfig-1.18.1}/src/plotfig/utils/matrix.py +0 -0
- {plotfig-1.18.0 → plotfig-1.18.1}/tests/plotfig/test_brain_surface.py +0 -0
- {plotfig-1.18.0 → plotfig-1.18.1}/tests/plotfig/test_single_bar.py +0 -0
|
@@ -15,15 +15,17 @@ jobs:
|
|
|
15
15
|
url: ${{ steps.deployment.outputs.page_url }}
|
|
16
16
|
runs-on: ubuntu-latest
|
|
17
17
|
steps:
|
|
18
|
-
- uses: actions/configure-pages@
|
|
19
|
-
- uses: actions/checkout@
|
|
18
|
+
- uses: actions/configure-pages@v6
|
|
19
|
+
- uses: actions/checkout@v6
|
|
20
20
|
- run: cp CHANGELOG.md docs/zh/changelog.md && cp CHANGELOG.md docs/en/changelog.md
|
|
21
|
-
- uses: actions/setup-python@
|
|
21
|
+
- uses: actions/setup-python@v6
|
|
22
22
|
with:
|
|
23
23
|
python-version: 3.x
|
|
24
24
|
- run: pip install zensical mkdocstrings-python
|
|
25
|
-
- run: zensical build --clean
|
|
26
|
-
-
|
|
25
|
+
- run: zensical build --config-file zensical.toml --clean
|
|
26
|
+
- run: zensical build --config-file zensical.zh.toml
|
|
27
|
+
- uses: actions/upload-pages-artifact@v5
|
|
27
28
|
with:
|
|
28
29
|
path: site
|
|
29
|
-
- uses: actions/deploy-pages@
|
|
30
|
+
- uses: actions/deploy-pages@v5
|
|
31
|
+
id: deployment
|
|
@@ -0,0 +1 @@
|
|
|
1
|
+
{".":"1.18.1"}
|
|
@@ -1,5 +1,12 @@
|
|
|
1
1
|
# Changelog
|
|
2
2
|
|
|
3
|
+
## [1.18.1](https://github.com/RicardoRyn/plotfig/compare/v1.18.0...v1.18.1) (2026-05-07)
|
|
4
|
+
|
|
5
|
+
|
|
6
|
+
### Code Refactoring ♻️
|
|
7
|
+
|
|
8
|
+
* **plotfig:** improve type hints for test_method and x_label_ha parameters ([d6fe349](https://github.com/RicardoRyn/plotfig/commit/d6fe349644a92c7ce89b980c7c67d36fc71ac2e3))
|
|
9
|
+
|
|
3
10
|
## [1.18.0](https://github.com/RicardoRyn/plotfig/compare/v1.17.0...v1.18.0) (2026-05-04)
|
|
4
11
|
|
|
5
12
|
|
|
@@ -1,6 +1,6 @@
|
|
|
1
1
|
Metadata-Version: 2.4
|
|
2
2
|
Name: plotfig
|
|
3
|
-
Version: 1.18.
|
|
3
|
+
Version: 1.18.1
|
|
4
4
|
Summary: Scientific plotting package for Cognitive neuroscience
|
|
5
5
|
Author-email: Ricardo Ryn <ricardoRyn1317@gmail.com>
|
|
6
6
|
License-File: LICENSE
|
|
@@ -31,7 +31,7 @@ Description-Content-Type: text/markdown
|
|
|
31
31
|
|
|
32
32
|
A Python visualization library designed for cognitive neuroscience research, providing efficient, easy-to-use, and beautiful plotting tools.
|
|
33
33
|
|
|
34
|
-

|
|
34
|
+

|
|
35
35
|
|
|
36
36
|
</div>
|
|
37
37
|
|
|
@@ -72,10 +72,8 @@ pip install plotfig
|
|
|
72
72
|
|
|
73
73
|
## Documentation
|
|
74
74
|
|
|
75
|
-
For detailed documentation and usage examples, please visit the [plotfig documentation](https://ricardoryn.github.io/plotfig/
|
|
75
|
+
For detailed documentation and usage examples, please visit the [plotfig documentation](https://ricardoryn.github.io/plotfig/).
|
|
76
76
|
|
|
77
77
|
## Contributing
|
|
78
78
|
|
|
79
79
|
We welcome Issues and PRs! Whether it's bug reports, feature suggestions, or documentation improvements, please feel free to open an [Issue](https://github.com/RicardoRyn/plotfig/issues).
|
|
80
|
-
|
|
81
|
-
For contribution guidelines, please see the [contribution guide](https://ricardoryn.github.io/plotfig/).
|
|
@@ -10,7 +10,7 @@
|
|
|
10
10
|
|
|
11
11
|
A Python visualization library designed for cognitive neuroscience research, providing efficient, easy-to-use, and beautiful plotting tools.
|
|
12
12
|
|
|
13
|
-

|
|
13
|
+

|
|
14
14
|
|
|
15
15
|
</div>
|
|
16
16
|
|
|
@@ -51,10 +51,8 @@ pip install plotfig
|
|
|
51
51
|
|
|
52
52
|
## Documentation
|
|
53
53
|
|
|
54
|
-
For detailed documentation and usage examples, please visit the [plotfig documentation](https://ricardoryn.github.io/plotfig/
|
|
54
|
+
For detailed documentation and usage examples, please visit the [plotfig documentation](https://ricardoryn.github.io/plotfig/).
|
|
55
55
|
|
|
56
56
|
## Contributing
|
|
57
57
|
|
|
58
58
|
We welcome Issues and PRs! Whether it's bug reports, feature suggestions, or documentation improvements, please feel free to open an [Issue](https://github.com/RicardoRyn/plotfig/issues).
|
|
59
|
-
|
|
60
|
-
For contribution guidelines, please see the [contribution guide](https://ricardoryn.github.io/plotfig/).
|
|
@@ -10,7 +10,7 @@
|
|
|
10
10
|
|
|
11
11
|
一个专为认知神经科学研究设计的 Python 可视化库,提供高效、易用且美观的绘图工具。
|
|
12
12
|
|
|
13
|
-

|
|
13
|
+

|
|
14
14
|
|
|
15
15
|
</div>
|
|
16
16
|
|
|
@@ -56,6 +56,3 @@ pip install plotfig
|
|
|
56
56
|
## 贡献
|
|
57
57
|
|
|
58
58
|
欢迎提交 Issue 或 PR!无论是 Bug 报告、功能建议还是文档改进,都非常欢迎在 [Issue](https://github.com/RicardoRyn/plotfig/issues) 中提出。
|
|
59
|
-
|
|
60
|
-
开发贡献流程请参见[贡献指南](https://ricardoryn.github.io/plotfig/)。
|
|
61
|
-
|
|
@@ -3,7 +3,7 @@
|
|
|
3
3
|
`plotfig` is a Python library designed specifically for scientific data visualization, dedicated to providing efficient, easy-to-use, and beautiful plotting tools for cognitive neuroscience researchers.
|
|
4
4
|
This project is developed based on mainstream visualization libraries in the industry—`matplotlib`, `surfplot`, and `plotly`, integrating their powerful features to meet the complex plotting needs in neuroscience and brain connectomics across various scenarios.
|
|
5
5
|
|
|
6
|
-

|
|
7
7
|
|
|
8
8
|
## Project Structure
|
|
9
9
|
|
|
@@ -27,4 +27,4 @@ The project adopts a modular design, containing the following main functional mo
|
|
|
27
27
|
Fun fact: All elements of a figure[^1].
|
|
28
28
|
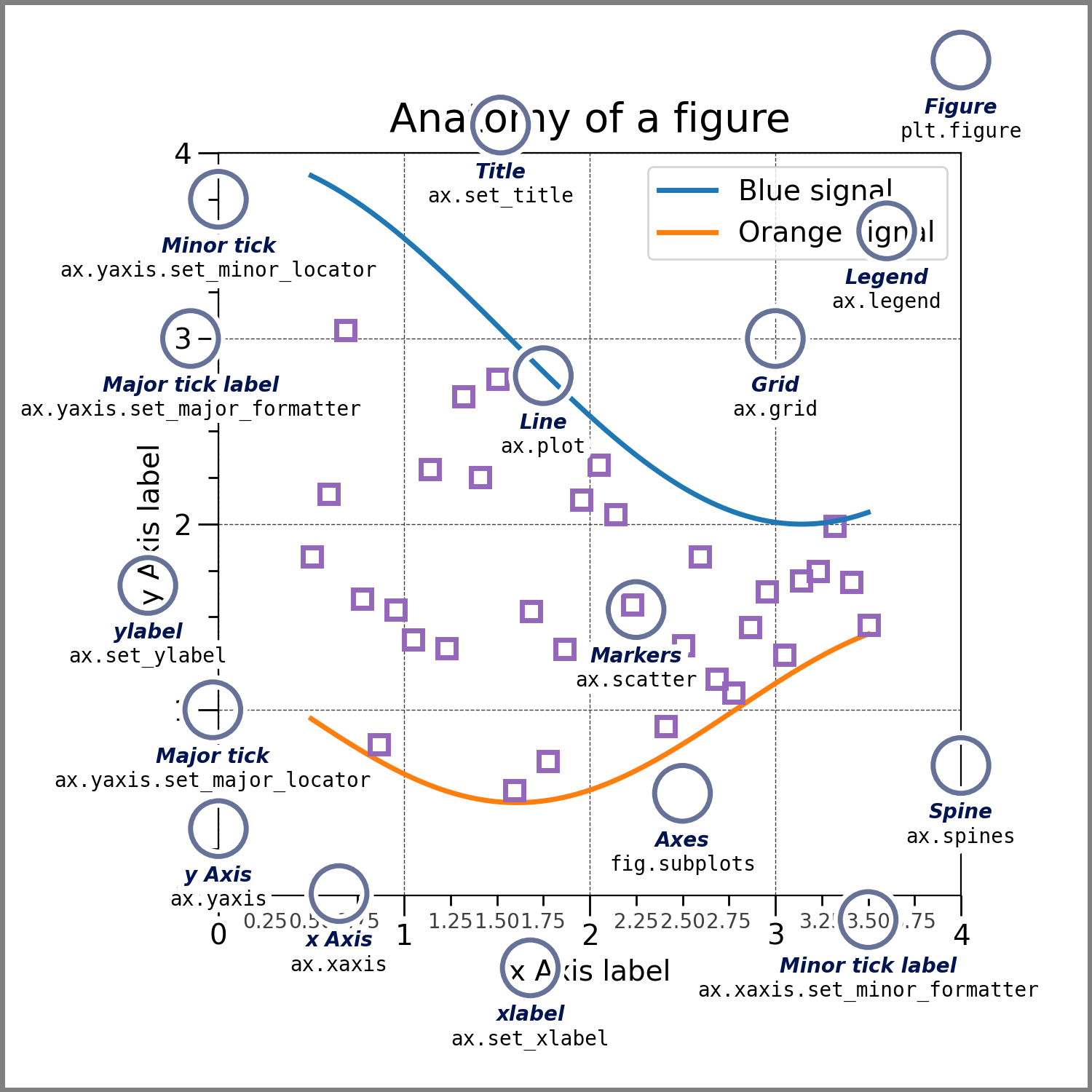
|
|
29
29
|
|
|
30
|
-
[^1]: [Quick start guide of matplotlib.](https://matplotlib.org/stable/tutorials/introductory/usage.html#parts-of-a-figure)
|
|
30
|
+
[^1]: [Quick start guide of matplotlib.](https://matplotlib.org/stable/tutorials/introductory/usage.html#parts-of-a-figure)
|
|
@@ -35,7 +35,7 @@ fig = plot_brain_connection_figure(
|
|
|
35
35
|
|
|
36
36
|
## Result Display
|
|
37
37
|
|
|
38
|
-

|
|
39
39
|
|
|
40
40
|
The HTML file can be interactively viewed in a browser. You can manually take screenshots, or use the following command to batch generate multi-view images.
|
|
41
41
|
|
|
@@ -7,14 +7,14 @@ This type of chart is commonly used to display various brain region metrics in n
|
|
|
7
7
|
The `plot_brain_surface_figure` function is developed based on the `surfplot` library, providing a unified and simplified interface for plotting brain surface maps for human, macaque, and chimpanzee brains.
|
|
8
8
|
Currently supports multiple brain atlases including:
|
|
9
9
|
|
|
10
|
-
1. Human Glasser (HCP-MMP) atlas[^1]. [Atlas CSV file](
|
|
11
|
-
1. Human BNA atlas[^2]. [Atlas CSV file](
|
|
12
|
-
1. Chimpanzee BNA atlas[^3]. [Atlas CSV file](
|
|
13
|
-
1. Macaque CHARM 4-level[^4]. [Atlas CSV file](
|
|
14
|
-
1. Macaque CHARM 5-level[^4]. [Atlas CSV file](
|
|
15
|
-
1. Macaque CHARM 6-level[^4]. [Atlas CSV file](
|
|
16
|
-
1. Macaque BNA atlas[^5]. [Atlas CSV file](
|
|
17
|
-
1. Macaque D99 atlas[^6]. [Atlas CSV file](
|
|
10
|
+
1. Human Glasser (HCP-MMP) atlas[^1]. [Atlas CSV file](../assets/atlas_csv/human_glasser.csv).
|
|
11
|
+
1. Human BNA atlas[^2]. [Atlas CSV file](../assets/atlas_csv/human_bna.csv).
|
|
12
|
+
1. Chimpanzee BNA atlas[^3]. [Atlas CSV file](../assets/atlas_csv/chimpanzee_bna.csv).
|
|
13
|
+
1. Macaque CHARM 4-level[^4]. [Atlas CSV file](../assets/atlas_csv/macaque_charm4.csv).
|
|
14
|
+
1. Macaque CHARM 5-level[^4]. [Atlas CSV file](../assets/atlas_csv/macaque_charm5.csv).
|
|
15
|
+
1. Macaque CHARM 6-level[^4]. [Atlas CSV file](../assets/atlas_csv/macaque_charm6.csv).
|
|
16
|
+
1. Macaque BNA atlas[^5]. [Atlas CSV file](../assets/atlas_csv/macaque_bna.csv).
|
|
17
|
+
1. Macaque D99 atlas[^6]. [Atlas CSV file](../assets/atlas_csv/macaque_d99.csv).
|
|
18
18
|
|
|
19
19
|
[^1]:
|
|
20
20
|
Glasser, M. F., Coalson, T. S., Robinson, E. C., Hacker, C. D., Harwell, J., Yacoub, E., Ugurbil, K., Andersson, J., Beckmann, C. F., Jenkinson, M., Smith, S. M., & Van Essen, D. C. (2016). A multi-modal parcellation of human cerebral cortex. Nature, 536(7615), Article 7615. https://doi.org/10.1038/nature18933
|
|
@@ -42,7 +42,7 @@ data = {"lh_V1": 1, "rh_MT": 1.5}
|
|
|
42
42
|
ax = plot_brain_surface_figure(data, species="human", atlas="glasser")
|
|
43
43
|
```
|
|
44
44
|
|
|
45
|
-

|
|
46
46
|
|
|
47
47
|
```python
|
|
48
48
|
from plotfig import plot_brain_surface_figure
|
|
@@ -61,7 +61,7 @@ ax2 = plot_brain_surface_figure(
|
|
|
61
61
|
)
|
|
62
62
|
```
|
|
63
63
|
|
|
64
|
-

|
|
65
65
|
|
|
66
66
|
## Different Surface Files
|
|
67
67
|
|
|
@@ -89,7 +89,7 @@ ax4 = plot_brain_surface_figure(plot_data, surf="sphere", ax=axes[1,0], title_na
|
|
|
89
89
|
ax5 = plot_brain_surface_figure(plot_data, surf="flat", ax=axes[1,1], title_name="flat")
|
|
90
90
|
```
|
|
91
91
|
|
|
92
|
-

|
|
93
93
|
|
|
94
94
|
For chimpanzees, the following surface files are provided:
|
|
95
95
|
|
|
@@ -108,7 +108,7 @@ ax1 = plot_brain_surface_figure(plot_data, species="chimpanzee", atlas="bna", su
|
|
|
108
108
|
ax3 = plot_brain_surface_figure(plot_data, species="chimpanzee", atlas="bna", surf="midthickness", ax=axes[1], title_name="midthickness")
|
|
109
109
|
```
|
|
110
110
|
|
|
111
|
-

|
|
112
112
|
|
|
113
113
|
For macaques, the following surface files are provided:
|
|
114
114
|
|
|
@@ -135,7 +135,7 @@ ax5 = plot_brain_surface_figure(plot_data, species="macaque", atlas="charm5", su
|
|
|
135
135
|
ax6 = plot_brain_surface_figure(plot_data, species="macaque", atlas="charm5", surf="flat", ax=axes[1,2], title_name="flat")
|
|
136
136
|
```
|
|
137
137
|
|
|
138
|
-

|
|
139
139
|
|
|
140
140
|
## More Settings
|
|
141
141
|
|
|
@@ -159,4 +159,4 @@ ax = plot_brain_surface_figure(
|
|
|
159
159
|
)
|
|
160
160
|
```
|
|
161
161
|
|
|
162
|
-

|
|
@@ -19,7 +19,7 @@ fig = plot_circos_figure(connectome)
|
|
|
19
19
|
# fig.savefig("./figures/circos1.png")
|
|
20
20
|
```
|
|
21
21
|
|
|
22
|
-

|
|
23
23
|
|
|
24
24
|
## Parameter Settings
|
|
25
25
|
|
|
@@ -55,7 +55,7 @@ fig = plot_circos_figure(
|
|
|
55
55
|
# fig.savefig("./figures/circos.png")
|
|
56
56
|
```
|
|
57
57
|
|
|
58
|
-

|
|
59
59
|
|
|
60
60
|
### Combining with Other Plots
|
|
61
61
|
|
|
@@ -81,7 +81,7 @@ ax2 = plot_circos_figure(connectome, ax=ax2)
|
|
|
81
81
|
# fig.savefig("./figures/circos.png")
|
|
82
82
|
```
|
|
83
83
|
|
|
84
|
-

|
|
85
85
|
|
|
86
86
|
### Symmetric and Asymmetric Circos Plots
|
|
87
87
|
|
|
@@ -105,7 +105,7 @@ ax2 = plot_circos_figure(connectome, symmetric=False, ax=ax2)
|
|
|
105
105
|
# fig.savefig("./figures/circos.png")
|
|
106
106
|
```
|
|
107
107
|
|
|
108
|
-

|
|
109
109
|
|
|
110
110
|
### Edge Colors
|
|
111
111
|
|
|
@@ -131,7 +131,7 @@ ax3 = plot_circos_figure(connectome, ax=ax3, edge_color="blue")
|
|
|
131
131
|
# fig.savefig("./figures/circos.png")
|
|
132
132
|
```
|
|
133
133
|
|
|
134
|
-

|
|
135
135
|
|
|
136
136
|
You can also apply Matplotlib's built-in common color maps (Colormap) through the `cmap` parameter.
|
|
137
137
|
|
|
@@ -158,7 +158,7 @@ ax3 = plot_circos_figure(connectome, ax=ax3, cmap="bwr")
|
|
|
158
158
|
# fig.savefig("./figures/circos.png")
|
|
159
159
|
```
|
|
160
160
|
|
|
161
|
-

|
|
162
162
|
|
|
163
163
|
When negative values exist in the connectome data, edge colors cannot be customized, and the system will default to using Matplotlib's `bwr` color map.
|
|
164
164
|
|
|
@@ -177,4 +177,4 @@ fig = plot_circos_figure(connectome)
|
|
|
177
177
|
|
|
178
178
|
2025-09-05 15:09:37.347 | WARNING | plotfig.circos:plot_circos_figure:116 - Due to negative values in connectome, connection colors cannot be customized; positive values will be displayed in red and negative values in blue
|
|
179
179
|
|
|
180
|
-

|
|
@@ -25,7 +25,7 @@ data2 = data1 + np.random.normal(1,50, 100)
|
|
|
25
25
|
ax = plot_correlation_figure(data1,data2)
|
|
26
26
|
```
|
|
27
27
|
|
|
28
|
-

|
|
29
29
|
|
|
30
30
|
## Hexbin Plot
|
|
31
31
|
|
|
@@ -59,7 +59,7 @@ hb = plot_correlation_figure(
|
|
|
59
59
|
cb = fig.colorbar(hb, ax=ax2, label='counts')
|
|
60
60
|
```
|
|
61
61
|
|
|
62
|
-

|
|
63
63
|
|
|
64
64
|
## Parameter Settings
|
|
65
65
|
|
|
@@ -94,4 +94,4 @@ ax = plot_correlation_figure(
|
|
|
94
94
|
)
|
|
95
95
|
```
|
|
96
96
|
|
|
97
|
-

|
|
@@ -18,7 +18,7 @@ data = np.random.rand(10, 19)
|
|
|
18
18
|
ax = plot_matrix_figure(data)
|
|
19
19
|
```
|
|
20
20
|
|
|
21
|
-

|
|
22
22
|
|
|
23
23
|
## Parameter Settings
|
|
24
24
|
|
|
@@ -44,4 +44,4 @@ ax = plot_matrix_figure(
|
|
|
44
44
|
)
|
|
45
45
|
```
|
|
46
46
|
|
|
47
|
-

|
|
@@ -29,14 +29,14 @@ group2_bar3 = np.random.normal(3, 1, 10)
|
|
|
29
29
|
ax = plot_multi_group_bar_figure([[group1_bar1, group1_bar2, group1_bar3], [group2_bar1, group2_bar2, group2_bar3]])
|
|
30
30
|
```
|
|
31
31
|
|
|
32
|
-

|
|
33
33
|
|
|
34
34
|
## Plot Beautification
|
|
35
35
|
|
|
36
36
|
Similar to single-group bar charts, multi-group bar charts also provide a large number of adjustable parameters for flexible control of the chart appearance.
|
|
37
37
|
This section only shows a portion of these parameters.
|
|
38
38
|
|
|
39
|
-
For the complete parameter list, please refer to the API documentation for [`plot_multi_group_bar_figure`](../api/#plotfig.
|
|
39
|
+
For the complete parameter list, please refer to the API documentation for [`plot_multi_group_bar_figure`](../api/#plotfig.multi_bars.plot_multi_group_bar_figure).
|
|
40
40
|
|
|
41
41
|
```python
|
|
42
42
|
import numpy as np
|
|
@@ -69,7 +69,7 @@ ax = plot_multi_group_bar_figure(
|
|
|
69
69
|
)
|
|
70
70
|
```
|
|
71
71
|
|
|
72
|
-

|
|
73
73
|
|
|
74
74
|
## Statistics
|
|
75
75
|
|
|
@@ -111,4 +111,4 @@ ax = plot_multi_group_bar_figure(
|
|
|
111
111
|
)
|
|
112
112
|
```
|
|
113
113
|
|
|
114
|
-

|
|
@@ -24,7 +24,7 @@ ax = plot_one_group_bar_figure([data1, data2, data3])
|
|
|
24
24
|
|
|
25
25
|
|
|
26
26
|
|
|
27
|
-

|
|
28
28
|
|
|
29
29
|
|
|
30
30
|
|
|
@@ -52,7 +52,7 @@ ax2 = plot_one_group_bar_figure([ax2_bar1, ax2_bar2], ax=axes[1])
|
|
|
52
52
|
|
|
53
53
|
|
|
54
54
|
|
|
55
|
-

|
|
56
56
|
|
|
57
57
|
|
|
58
58
|
|
|
@@ -85,7 +85,7 @@ ax4 = plot_one_group_bar_figure([ax4_bar1, ax4_bar2], ax=axes[1,1], labels_name=
|
|
|
85
85
|
|
|
86
86
|
|
|
87
87
|
|
|
88
|
-

|
|
89
89
|
|
|
90
90
|
|
|
91
91
|
|
|
@@ -97,7 +97,7 @@ We can create a `fig` object externally to flexibly control figure size.
|
|
|
97
97
|
`plotfig` provides rich options for customizing plot styles.
|
|
98
98
|
Below is an example showing some commonly used parameters of `plot_one_group_bar_figure`.
|
|
99
99
|
|
|
100
|
-
For complete parameter descriptions, see the API documentation for [`plot_one_group_bar_figure`](../api
|
|
100
|
+
For complete parameter descriptions, see the API documentation for [`plot_one_group_bar_figure`](../api/#plotfig.single_bar.plot_one_group_bar_figure).
|
|
101
101
|
|
|
102
102
|
|
|
103
103
|
```python
|
|
@@ -129,7 +129,7 @@ ax = plot_one_group_bar_figure(
|
|
|
129
129
|
|
|
130
130
|
|
|
131
131
|
|
|
132
|
-

|
|
133
133
|
|
|
134
134
|
|
|
135
135
|
|
|
@@ -169,7 +169,7 @@ ax = plot_one_group_bar_figure(
|
|
|
169
169
|
|
|
170
170
|
|
|
171
171
|
|
|
172
|
-

|
|
173
173
|
|
|
174
174
|
|
|
175
175
|
|
|
@@ -211,7 +211,7 @@ ax2 = plot_one_group_bar_figure(
|
|
|
211
211
|
|
|
212
212
|
|
|
213
213
|
|
|
214
|
-

|
|
215
215
|
|
|
216
216
|
|
|
217
217
|
|
|
@@ -251,7 +251,7 @@ ax2 = plot_one_group_bar_figure(
|
|
|
251
251
|
|
|
252
252
|
|
|
253
253
|
|
|
254
|
-

|
|
255
255
|
|
|
256
256
|
|
|
257
257
|
|
|
@@ -288,7 +288,7 @@ ax2 = plot_one_group_bar_figure(
|
|
|
288
288
|
|
|
289
289
|
|
|
290
290
|
|
|
291
|
-

|
|
292
292
|
|
|
293
293
|
|
|
294
294
|
|
|
@@ -325,7 +325,7 @@ ax2 = plot_one_group_bar_figure(
|
|
|
325
325
|
|
|
326
326
|
|
|
327
327
|
|
|
328
|
-

|
|
329
329
|
|
|
330
330
|
|
|
331
331
|
|
|
@@ -379,7 +379,7 @@ ax4 = plot_one_group_bar_figure(
|
|
|
379
379
|
|
|
380
380
|
|
|
381
381
|
|
|
382
|
-

|
|
383
383
|
|
|
384
384
|
|
|
385
385
|
|
|
@@ -421,7 +421,7 @@ ax2 = plot_one_group_bar_figure(
|
|
|
421
421
|
|
|
422
422
|
|
|
423
423
|
|
|
424
|
-

|
|
425
425
|
|
|
426
426
|
|
|
427
427
|
|
|
@@ -464,7 +464,7 @@ ax2 = plot_one_group_bar_figure(
|
|
|
464
464
|
|
|
465
465
|
|
|
466
466
|
|
|
467
|
-

|
|
468
468
|
|
|
469
469
|
|
|
470
470
|
|
|
@@ -538,7 +538,7 @@ ax4 = plot_one_group_bar_figure(
|
|
|
538
538
|
|
|
539
539
|
|
|
540
540
|
|
|
541
|
-

|
|
542
542
|
|
|
543
543
|
|
|
544
544
|
|
|
@@ -582,7 +582,7 @@ ax = plot_one_group_bar_figure(
|
|
|
582
582
|
|
|
583
583
|
|
|
584
584
|
|
|
585
|
-

|
|
586
586
|
|
|
587
587
|
|
|
588
588
|
|
|
@@ -656,7 +656,7 @@ ax4 = plot_one_group_bar_figure(
|
|
|
656
656
|
|
|
657
657
|
|
|
658
658
|
|
|
659
|
-

|
|
660
660
|
|
|
661
661
|
|
|
662
662
|
|
|
@@ -710,6 +710,6 @@ ax = plot_one_group_violin_figure(
|
|
|
710
710
|
|
|
711
711
|
|
|
712
712
|
|
|
713
|
-

|
|
714
714
|
|
|
715
715
|
|
|
@@ -0,0 +1,201 @@
|
|
|
1
|
+
Label,ROIs_name
|
|
2
|
+
1,lh_SFG.r
|
|
3
|
+
2,lh_SFG.ri
|
|
4
|
+
3,lh_SFG.ci
|
|
5
|
+
4,lh_SFG.c
|
|
6
|
+
5,lh_SFG.lc
|
|
7
|
+
6,lh_SFG.mc
|
|
8
|
+
7,lh_MFG.r
|
|
9
|
+
8,lh_MFG.ri
|
|
10
|
+
9,lh_MFG.i
|
|
11
|
+
10,lh_MFG.di
|
|
12
|
+
11,lh_MFG.vi
|
|
13
|
+
12,lh_MFG.dc
|
|
14
|
+
13,lh_MFG.vc
|
|
15
|
+
14,lh_IFG.r
|
|
16
|
+
15,lh_IFG.ri
|
|
17
|
+
16,lh_IFG.ci
|
|
18
|
+
17,lh_IFG.dc
|
|
19
|
+
18,lh_IFG.vc
|
|
20
|
+
19,lh_IFG.c
|
|
21
|
+
20,lh_OrG.m
|
|
22
|
+
21,lh_OrG.r
|
|
23
|
+
22,lh_OrG.i
|
|
24
|
+
23,lh_OrG.c
|
|
25
|
+
24,lh_OrG.rl
|
|
26
|
+
25,lh_OrG.cl
|
|
27
|
+
26,lh_PrG.rd
|
|
28
|
+
27,lh_PrG.cd
|
|
29
|
+
28,lh_PrG.ri
|
|
30
|
+
29,lh_PrG.ci
|
|
31
|
+
30,lh_PrG.i
|
|
32
|
+
31,lh_PrG.vi
|
|
33
|
+
32,lh_PrG.v
|
|
34
|
+
33,lh_PCL.r
|
|
35
|
+
34,lh_PCL.c
|
|
36
|
+
35,lh_STG.r
|
|
37
|
+
36,lh_STG.ri
|
|
38
|
+
37,lh_STG.ci
|
|
39
|
+
38,lh_STG.c
|
|
40
|
+
39,lh_STG.dc
|
|
41
|
+
40,lh_STG.vc
|
|
42
|
+
41,lh_MTG.r
|
|
43
|
+
42,lh_MTG.i
|
|
44
|
+
43,lh_MTG.c
|
|
45
|
+
44,lh_MTG.cv
|
|
46
|
+
45,lh_ITG.r
|
|
47
|
+
46,lh_ITG.dr
|
|
48
|
+
47,lh_ITG.vr
|
|
49
|
+
48,lh_ITG.i
|
|
50
|
+
49,lh_ITG.dc
|
|
51
|
+
50,lh_ITG.vc
|
|
52
|
+
51,lh_ITG.c
|
|
53
|
+
52,lh_FuG.r
|
|
54
|
+
53,lh_FuG.ri
|
|
55
|
+
54,lh_FuG.ci
|
|
56
|
+
55,lh_FuG.m
|
|
57
|
+
56,lh_FuG.lc
|
|
58
|
+
57,lh_FuG.mc
|
|
59
|
+
58,lh_PhG.r
|
|
60
|
+
59,lh_PhG.mi
|
|
61
|
+
60,lh_PhG.li
|
|
62
|
+
61,lh_PhG.mc
|
|
63
|
+
62,lh_PhG.lc
|
|
64
|
+
63,lh_SPL.dr
|
|
65
|
+
64,lh_SPL.vr
|
|
66
|
+
65,lh_SPL.i
|
|
67
|
+
66,lh_SPL.ci
|
|
68
|
+
67,lh_SPL.c
|
|
69
|
+
68,lh_IPL.r
|
|
70
|
+
69,lh_IPL.ri
|
|
71
|
+
70,lh_IPL.v
|
|
72
|
+
71,lh_IPL.ci
|
|
73
|
+
72,lh_IPL.c
|
|
74
|
+
73,lh_PrL.r
|
|
75
|
+
74,lh_PrL.c
|
|
76
|
+
75,lh_Pcun.d
|
|
77
|
+
76,lh_Pcun.di
|
|
78
|
+
77,lh_Pcun.vi
|
|
79
|
+
78,lh_Pcun.v
|
|
80
|
+
79,lh_PoG.d
|
|
81
|
+
80,lh_PoG.i
|
|
82
|
+
81,lh_PoG.v
|
|
83
|
+
82,lh_INS.dr
|
|
84
|
+
83,lh_INS.ir
|
|
85
|
+
84,lh_INS.vr
|
|
86
|
+
85,lh_INS.rd
|
|
87
|
+
86,lh_INS.cd
|
|
88
|
+
87,lh_INS.i
|
|
89
|
+
88,lh_INS.v
|
|
90
|
+
89,lh_CG.r
|
|
91
|
+
90,lh_CG.i
|
|
92
|
+
91,lh_CG.dc
|
|
93
|
+
92,lh_CG.vc
|
|
94
|
+
93,lh_MVOcC.rd
|
|
95
|
+
94,lh_MVOcC.cd
|
|
96
|
+
95,lh_MVOcC.rv
|
|
97
|
+
96,lh_MVOcC.cv
|
|
98
|
+
97,lh_LOcC.d
|
|
99
|
+
98,lh_LOcC.i
|
|
100
|
+
99,lh_LOcC.rv
|
|
101
|
+
100,lh_LOcC.cv
|
|
102
|
+
101,rh_SFG.r
|
|
103
|
+
102,rh_SFG.ri
|
|
104
|
+
103,rh_SFG.ci
|
|
105
|
+
104,rh_SFG.c
|
|
106
|
+
105,rh_SFG.lc
|
|
107
|
+
106,rh_SFG.mc
|
|
108
|
+
107,rh_MFG.r
|
|
109
|
+
108,rh_MFG.ri
|
|
110
|
+
109,rh_MFG.i
|
|
111
|
+
110,rh_MFG.di
|
|
112
|
+
111,rh_MFG.vi
|
|
113
|
+
112,rh_MFG.dc
|
|
114
|
+
113,rh_MFG.vc
|
|
115
|
+
114,rh_IFG.r
|
|
116
|
+
115,rh_IFG.ri
|
|
117
|
+
116,rh_IFG.ci
|
|
118
|
+
117,rh_IFG.dc
|
|
119
|
+
118,rh_IFG.vc
|
|
120
|
+
119,rh_IFG.c
|
|
121
|
+
120,rh_OrG.m
|
|
122
|
+
121,rh_OrG.r
|
|
123
|
+
122,rh_OrG.i
|
|
124
|
+
123,rh_OrG.c
|
|
125
|
+
124,rh_OrG.rl
|
|
126
|
+
125,rh_OrG.cl
|
|
127
|
+
126,rh_PrG.rd
|
|
128
|
+
127,rh_PrG.cd
|
|
129
|
+
128,rh_PrG.ri
|
|
130
|
+
129,rh_PrG.ci
|
|
131
|
+
130,rh_PrG.i
|
|
132
|
+
131,rh_PrG.vi
|
|
133
|
+
132,rh_PrG.v
|
|
134
|
+
133,rh_PCL.r
|
|
135
|
+
134,rh_PCL.c
|
|
136
|
+
135,rh_STG.r
|
|
137
|
+
136,rh_STG.ri
|
|
138
|
+
137,rh_STG.ci
|
|
139
|
+
138,rh_STG.c
|
|
140
|
+
139,rh_STG.dc
|
|
141
|
+
140,rh_STG.vc
|
|
142
|
+
141,rh_MTG.r
|
|
143
|
+
142,rh_MTG.i
|
|
144
|
+
143,rh_MTG.c
|
|
145
|
+
144,rh_MTG.cv
|
|
146
|
+
145,rh_ITG.r
|
|
147
|
+
146,rh_ITG.dr
|
|
148
|
+
147,rh_ITG.vr
|
|
149
|
+
148,rh_ITG.i
|
|
150
|
+
149,rh_ITG.dc
|
|
151
|
+
150,rh_ITG.vc
|
|
152
|
+
151,rh_ITG.c
|
|
153
|
+
152,rh_FuG.r
|
|
154
|
+
153,rh_FuG.ri
|
|
155
|
+
154,rh_FuG.ci
|
|
156
|
+
155,rh_FuG.m
|
|
157
|
+
156,rh_FuG.lc
|
|
158
|
+
157,rh_FuG.mc
|
|
159
|
+
158,rh_PhG.r
|
|
160
|
+
159,rh_PhG.mi
|
|
161
|
+
160,rh_PhG.li
|
|
162
|
+
161,rh_PhG.mc
|
|
163
|
+
162,rh_PhG.lc
|
|
164
|
+
163,rh_SPL.dr
|
|
165
|
+
164,rh_SPL.vr
|
|
166
|
+
165,rh_SPL.i
|
|
167
|
+
166,rh_SPL.ci
|
|
168
|
+
167,rh_SPL.c
|
|
169
|
+
168,rh_IPL.r
|
|
170
|
+
169,rh_IPL.ri
|
|
171
|
+
170,rh_IPL.v
|
|
172
|
+
171,rh_IPL.ci
|
|
173
|
+
172,rh_IPL.c
|
|
174
|
+
173,rh_PrL.r
|
|
175
|
+
174,rh_PrL.c
|
|
176
|
+
175,rh_Pcun.d
|
|
177
|
+
176,rh_Pcun.di
|
|
178
|
+
177,rh_Pcun.vi
|
|
179
|
+
178,rh_Pcun.v
|
|
180
|
+
179,rh_PoG.d
|
|
181
|
+
180,rh_PoG.i
|
|
182
|
+
181,rh_PoG.v
|
|
183
|
+
182,rh_INS.dr
|
|
184
|
+
183,rh_INS.ir
|
|
185
|
+
184,rh_INS.vr
|
|
186
|
+
185,rh_INS.rd
|
|
187
|
+
186,rh_INS.cd
|
|
188
|
+
187,rh_INS.i
|
|
189
|
+
188,rh_INS.v
|
|
190
|
+
189,rh_CG.r
|
|
191
|
+
190,rh_CG.i
|
|
192
|
+
191,rh_CG.dc
|
|
193
|
+
192,rh_CG.vc
|
|
194
|
+
193,rh_MVOcC.rd
|
|
195
|
+
194,rh_MVOcC.cd
|
|
196
|
+
195,rh_MVOcC.rv
|
|
197
|
+
196,rh_MVOcC.cv
|
|
198
|
+
197,rh_LOcC.d
|
|
199
|
+
198,rh_LOcC.i
|
|
200
|
+
199,rh_LOcC.rv
|
|
201
|
+
200,rh_LOcC.cv
|