plotfig 1.0.0__tar.gz → 1.2.1__tar.gz
This diff represents the content of publicly available package versions that have been released to one of the supported registries. The information contained in this diff is provided for informational purposes only and reflects changes between package versions as they appear in their respective public registries.
- plotfig-1.2.1/.github/workflows/SyncToGitee.yml +18 -0
- plotfig-1.2.1/.github/workflows/ci.yml +29 -0
- plotfig-1.2.1/.gitignore +19 -0
- plotfig-1.2.1/.python-version +1 -0
- plotfig-1.2.1/CHANGELOG.md +43 -0
- {plotfig-1.0.0 → plotfig-1.2.1}/PKG-INFO +52 -60
- {plotfig-1.0.0 → plotfig-1.2.1}/README.md +7 -14
- plotfig-1.2.1/docs/api/index.md +18 -0
- plotfig-1.2.1/docs/assets/plotfig.png +0 -0
- plotfig-1.2.1/docs/index.md +15 -0
- plotfig-1.2.1/docs/installation.md +37 -0
- plotfig-1.2.1/docs/usage/brain_connectivity.md +63 -0
- plotfig-1.2.1/docs/usage/brain_connectivity_files/human.gif +0 -0
- plotfig-1.2.1/docs/usage/brain_surface.md +2014 -0
- plotfig-1.2.1/docs/usage/brain_surface_files/brain_surface_10_0.png +0 -0
- plotfig-1.2.1/docs/usage/brain_surface_files/brain_surface_10_1.png +0 -0
- plotfig-1.2.1/docs/usage/brain_surface_files/brain_surface_11_0.png +0 -0
- plotfig-1.2.1/docs/usage/brain_surface_files/brain_surface_12_0.png +0 -0
- plotfig-1.2.1/docs/usage/brain_surface_files/brain_surface_13_0.png +0 -0
- plotfig-1.2.1/docs/usage/brain_surface_files/brain_surface_13_1.png +0 -0
- plotfig-1.2.1/docs/usage/brain_surface_files/brain_surface_14_0.png +0 -0
- plotfig-1.2.1/docs/usage/brain_surface_files/brain_surface_14_1.png +0 -0
- plotfig-1.2.1/docs/usage/brain_surface_files/brain_surface_15_0.png +0 -0
- plotfig-1.2.1/docs/usage/brain_surface_files/brain_surface_16_0.png +0 -0
- plotfig-1.2.1/docs/usage/brain_surface_files/brain_surface_17_0.png +0 -0
- plotfig-1.2.1/docs/usage/brain_surface_files/brain_surface_18_0.png +0 -0
- plotfig-1.2.1/docs/usage/brain_surface_files/brain_surface_19_0.png +0 -0
- plotfig-1.2.1/docs/usage/brain_surface_files/brain_surface_20_0.png +0 -0
- plotfig-1.2.1/docs/usage/brain_surface_files/brain_surface_21_0.png +0 -0
- plotfig-1.2.1/docs/usage/brain_surface_files/brain_surface_23_0.png +0 -0
- plotfig-1.2.1/docs/usage/brain_surface_files/brain_surface_24_0.png +0 -0
- plotfig-1.2.1/docs/usage/brain_surface_files/brain_surface_25_0.png +0 -0
- plotfig-1.2.1/docs/usage/brain_surface_files/brain_surface_4_0.png +0 -0
- plotfig-1.2.1/docs/usage/brain_surface_files/brain_surface_5_0.png +0 -0
- plotfig-1.2.1/docs/usage/brain_surface_files/brain_surface_6_0.png +0 -0
- plotfig-1.2.1/docs/usage/brain_surface_files/brain_surface_6_1.png +0 -0
- plotfig-1.2.1/docs/usage/brain_surface_files/brain_surface_7_0.png +0 -0
- plotfig-1.2.1/docs/usage/brain_surface_files/brain_surface_7_1.png +0 -0
- plotfig-1.2.1/docs/usage/brain_surface_files/brain_surface_8_0.png +0 -0
- plotfig-1.2.1/docs/usage/brain_surface_files/brain_surface_8_1.png +0 -0
- plotfig-1.2.1/docs/usage/brain_surface_files/brain_surface_9_0.png +0 -0
- plotfig-1.2.1/docs/usage/brain_surface_files/brain_surface_9_1.png +0 -0
- plotfig-1.2.1/docs/usage/circos.md +56 -0
- plotfig-1.2.1/docs/usage/circos_files/circos_1_0.png +0 -0
- plotfig-1.2.1/docs/usage/circos_files/circos_1_1.png +0 -0
- plotfig-1.2.1/docs/usage/circos_files/circos_2_0.png +0 -0
- plotfig-1.2.1/docs/usage/correlation.md +102 -0
- plotfig-1.2.1/docs/usage/correlation_files/correlation_2_0.png +0 -0
- plotfig-1.2.1/docs/usage/correlation_files/correlation_5_0.png +0 -0
- plotfig-1.2.1/docs/usage/correlation_files/correlation_7_0.png +0 -0
- plotfig-1.2.1/docs/usage/matrix.md +66 -0
- plotfig-1.2.1/docs/usage/matrix_files/matrix_2_1.png +0 -0
- plotfig-1.2.1/docs/usage/matrix_files/matrix_5_1.png +0 -0
- plotfig-1.2.1/docs/usage/multi_groups.md +115 -0
- plotfig-1.2.1/docs/usage/multi_groups_files/multi_groups_2_0.png +0 -0
- plotfig-1.2.1/docs/usage/multi_groups_files/multi_groups_5_0.png +0 -0
- plotfig-1.2.1/docs/usage/multi_groups_files/multi_groups_8_0.png +0 -0
- plotfig-1.2.1/docs/usage/single_group.md +676 -0
- plotfig-1.2.1/docs/usage/single_group_files/single_group_11_0.png +0 -0
- plotfig-1.2.1/docs/usage/single_group_files/single_group_12_0.png +0 -0
- plotfig-1.2.1/docs/usage/single_group_files/single_group_13_0.png +0 -0
- plotfig-1.2.1/docs/usage/single_group_files/single_group_14_0.png +0 -0
- plotfig-1.2.1/docs/usage/single_group_files/single_group_16_0.png +0 -0
- plotfig-1.2.1/docs/usage/single_group_files/single_group_17_0.png +0 -0
- plotfig-1.2.1/docs/usage/single_group_files/single_group_19_0.png +0 -0
- plotfig-1.2.1/docs/usage/single_group_files/single_group_20_0.png +0 -0
- plotfig-1.2.1/docs/usage/single_group_files/single_group_21_0.png +0 -0
- plotfig-1.2.1/docs/usage/single_group_files/single_group_22_0.png +0 -0
- plotfig-1.2.1/docs/usage/single_group_files/single_group_23_0.png +0 -0
- plotfig-1.2.1/docs/usage/single_group_files/single_group_24_0.png +0 -0
- plotfig-1.2.1/docs/usage/single_group_files/single_group_25_0.png +0 -0
- plotfig-1.2.1/docs/usage/single_group_files/single_group_26_0.png +0 -0
- plotfig-1.2.1/docs/usage/single_group_files/single_group_27_0.png +0 -0
- plotfig-1.2.1/docs/usage/single_group_files/single_group_28_0.png +0 -0
- plotfig-1.2.1/docs/usage/single_group_files/single_group_2_0.png +0 -0
- plotfig-1.2.1/docs/usage/single_group_files/single_group_30_0.png +0 -0
- plotfig-1.2.1/docs/usage/single_group_files/single_group_31_0.png +0 -0
- plotfig-1.2.1/docs/usage/single_group_files/single_group_33_0.png +0 -0
- plotfig-1.2.1/docs/usage/single_group_files/single_group_34_0.png +0 -0
- plotfig-1.2.1/docs/usage/single_group_files/single_group_35_0.png +0 -0
- plotfig-1.2.1/docs/usage/single_group_files/single_group_36_0.png +0 -0
- plotfig-1.2.1/docs/usage/single_group_files/single_group_39_0.png +0 -0
- plotfig-1.2.1/docs/usage/single_group_files/single_group_3_0.png +0 -0
- plotfig-1.2.1/docs/usage/single_group_files/single_group_5_0.png +0 -0
- plotfig-1.2.1/docs/usage/single_group_files/single_group_6_0.png +0 -0
- plotfig-1.2.1/docs/usage/single_group_files/single_group_6_1.png +0 -0
- plotfig-1.2.1/docs/usage/single_group_files/single_group_7_0.png +0 -0
- plotfig-1.2.1/docs/usage/single_group_files/single_group_8_0.png +0 -0
- plotfig-1.2.1/mkdocs.yml +146 -0
- {plotfig-1.0.0 → plotfig-1.2.1}/pyproject.toml +5 -16
- {plotfig-1.0.0 → plotfig-1.2.1}/src/plotfig/__init__.py +2 -0
- {plotfig-1.0.0 → plotfig-1.2.1}/src/plotfig/bar.py +295 -62
- {plotfig-1.0.0 → plotfig-1.2.1}/src/plotfig/correlation.py +13 -2
- {plotfig-1.0.0 → plotfig-1.2.1}/src/plotfig/matrix.py +1 -1
- plotfig-1.2.1/uv.lock +1320 -0
- plotfig-1.0.0/plotfig.egg-info/PKG-INFO +0 -60
- plotfig-1.0.0/plotfig.egg-info/SOURCES.txt +0 -81
- plotfig-1.0.0/plotfig.egg-info/dependency_links.txt +0 -1
- plotfig-1.0.0/plotfig.egg-info/requires.txt +0 -8
- plotfig-1.0.0/plotfig.egg-info/top_level.txt +0 -1
- plotfig-1.0.0/setup.cfg +0 -4
- {plotfig-1.0.0 → plotfig-1.2.1}/LICENSE +0 -0
- {plotfig-1.0.0 → plotfig-1.2.1}/src/plotfig/brain_connection.py +0 -0
- {plotfig-1.0.0 → plotfig-1.2.1}/src/plotfig/brain_surface.py +0 -0
- {plotfig-1.0.0 → plotfig-1.2.1}/src/plotfig/circos.py +0 -0
- {plotfig-1.0.0 → plotfig-1.2.1}/src/plotfig/data/atlas_tables/chimpanzee_bna.csv +0 -0
- {plotfig-1.0.0 → plotfig-1.2.1}/src/plotfig/data/atlas_tables/human_bna.csv +0 -0
- {plotfig-1.0.0 → plotfig-1.2.1}/src/plotfig/data/atlas_tables/human_glasser.csv +0 -0
- {plotfig-1.0.0 → plotfig-1.2.1}/src/plotfig/data/atlas_tables/macaque_bna.csv +0 -0
- {plotfig-1.0.0 → plotfig-1.2.1}/src/plotfig/data/atlas_tables/macaque_charm5.csv +0 -0
- {plotfig-1.0.0 → plotfig-1.2.1}/src/plotfig/data/atlas_tables/macaque_charm5_add_13_sgms.csv +0 -0
- {plotfig-1.0.0 → plotfig-1.2.1}/src/plotfig/data/atlas_tables/macaque_charm6.csv +0 -0
- {plotfig-1.0.0 → plotfig-1.2.1}/src/plotfig/data/atlas_tables/macaque_d99.csv +0 -0
- {plotfig-1.0.0 → plotfig-1.2.1}/src/plotfig/data/neurodata/atlases/chimpanzee_BNA/ChimpBNA.L.32k_fs_LR.label.gii +0 -0
- {plotfig-1.0.0 → plotfig-1.2.1}/src/plotfig/data/neurodata/atlases/chimpanzee_BNA/ChimpBNA.R.32k_fs_LR.label.gii +0 -0
- {plotfig-1.0.0 → plotfig-1.2.1}/src/plotfig/data/neurodata/atlases/human_BNA/fsaverage.L.BNA.32k_fs_LR.label.gii +0 -0
- {plotfig-1.0.0 → plotfig-1.2.1}/src/plotfig/data/neurodata/atlases/human_BNA/fsaverage.R.BNA.32k_fs_LR.label.gii +0 -0
- {plotfig-1.0.0 → plotfig-1.2.1}/src/plotfig/data/neurodata/atlases/human_Glasser/fsaverage.L.Glasser.32k_fs_LR.label.gii +0 -0
- {plotfig-1.0.0 → plotfig-1.2.1}/src/plotfig/data/neurodata/atlases/human_Glasser/fsaverage.R.Glasser.32k_fs_LR.label.gii +0 -0
- {plotfig-1.0.0 → plotfig-1.2.1}/src/plotfig/data/neurodata/atlases/macaque_BNA/MBNA_124_32k_L.label.gii +0 -0
- {plotfig-1.0.0 → plotfig-1.2.1}/src/plotfig/data/neurodata/atlases/macaque_BNA/MBNA_124_32k_R.label.gii +0 -0
- {plotfig-1.0.0 → plotfig-1.2.1}/src/plotfig/data/neurodata/atlases/macaque_CHARM5/L.charm5.label.gii +0 -0
- {plotfig-1.0.0 → plotfig-1.2.1}/src/plotfig/data/neurodata/atlases/macaque_CHARM5/R.charm5.label.gii +0 -0
- {plotfig-1.0.0 → plotfig-1.2.1}/src/plotfig/data/neurodata/atlases/macaque_CHARM6/L.charm6.label.gii +0 -0
- {plotfig-1.0.0 → plotfig-1.2.1}/src/plotfig/data/neurodata/atlases/macaque_CHARM6/R.charm6.label.gii +0 -0
- {plotfig-1.0.0 → plotfig-1.2.1}/src/plotfig/data/neurodata/atlases/macaque_D99/L.d99.label.gii +0 -0
- {plotfig-1.0.0 → plotfig-1.2.1}/src/plotfig/data/neurodata/atlases/macaque_D99/R.d99.label.gii +0 -0
- {plotfig-1.0.0 → plotfig-1.2.1}/src/plotfig/data/neurodata/surfaces/chimpanzee_BNA/ChimpYerkes29_v1.2.L.midthickness.32k_fs_LR.surf.gii +0 -0
- {plotfig-1.0.0 → plotfig-1.2.1}/src/plotfig/data/neurodata/surfaces/chimpanzee_BNA/ChimpYerkes29_v1.2.L.veryinflated.32k_fs_LR.surf.gii +0 -0
- {plotfig-1.0.0 → plotfig-1.2.1}/src/plotfig/data/neurodata/surfaces/chimpanzee_BNA/ChimpYerkes29_v1.2.R.midthickness.32k_fs_LR.surf.gii +0 -0
- {plotfig-1.0.0 → plotfig-1.2.1}/src/plotfig/data/neurodata/surfaces/chimpanzee_BNA/ChimpYerkes29_v1.2.R.veryinflated.32k_fs_LR.surf.gii +0 -0
- {plotfig-1.0.0 → plotfig-1.2.1}/src/plotfig/data/neurodata/surfaces/human_fsLR/README.md +0 -0
- {plotfig-1.0.0 → plotfig-1.2.1}/src/plotfig/data/neurodata/surfaces/human_fsLR/tpl-fsLR_den-32k_hemi-L_desc-nomedialwall_dparc.label.gii +0 -0
- {plotfig-1.0.0 → plotfig-1.2.1}/src/plotfig/data/neurodata/surfaces/human_fsLR/tpl-fsLR_den-32k_hemi-L_desc-sulc_midthickness.shape.gii +0 -0
- {plotfig-1.0.0 → plotfig-1.2.1}/src/plotfig/data/neurodata/surfaces/human_fsLR/tpl-fsLR_den-32k_hemi-L_desc-vaavg_midthickness.shape.gii +0 -0
- {plotfig-1.0.0 → plotfig-1.2.1}/src/plotfig/data/neurodata/surfaces/human_fsLR/tpl-fsLR_den-32k_hemi-L_inflated.surf.gii +0 -0
- {plotfig-1.0.0 → plotfig-1.2.1}/src/plotfig/data/neurodata/surfaces/human_fsLR/tpl-fsLR_den-32k_hemi-L_midthickness.surf.gii +0 -0
- {plotfig-1.0.0 → plotfig-1.2.1}/src/plotfig/data/neurodata/surfaces/human_fsLR/tpl-fsLR_den-32k_hemi-L_sphere.surf.gii +0 -0
- {plotfig-1.0.0 → plotfig-1.2.1}/src/plotfig/data/neurodata/surfaces/human_fsLR/tpl-fsLR_den-32k_hemi-L_veryinflated.surf.gii +0 -0
- {plotfig-1.0.0 → plotfig-1.2.1}/src/plotfig/data/neurodata/surfaces/human_fsLR/tpl-fsLR_den-32k_hemi-R_desc-nomedialwall_dparc.label.gii +0 -0
- {plotfig-1.0.0 → plotfig-1.2.1}/src/plotfig/data/neurodata/surfaces/human_fsLR/tpl-fsLR_den-32k_hemi-R_desc-sulc_midthickness.shape.gii +0 -0
- {plotfig-1.0.0 → plotfig-1.2.1}/src/plotfig/data/neurodata/surfaces/human_fsLR/tpl-fsLR_den-32k_hemi-R_desc-vaavg_midthickness.shape.gii +0 -0
- {plotfig-1.0.0 → plotfig-1.2.1}/src/plotfig/data/neurodata/surfaces/human_fsLR/tpl-fsLR_den-32k_hemi-R_inflated.surf.gii +0 -0
- {plotfig-1.0.0 → plotfig-1.2.1}/src/plotfig/data/neurodata/surfaces/human_fsLR/tpl-fsLR_den-32k_hemi-R_midthickness.surf.gii +0 -0
- {plotfig-1.0.0 → plotfig-1.2.1}/src/plotfig/data/neurodata/surfaces/human_fsLR/tpl-fsLR_den-32k_hemi-R_sphere.surf.gii +0 -0
- {plotfig-1.0.0 → plotfig-1.2.1}/src/plotfig/data/neurodata/surfaces/human_fsLR/tpl-fsLR_den-32k_hemi-R_veryinflated.surf.gii +0 -0
- {plotfig-1.0.0 → plotfig-1.2.1}/src/plotfig/data/neurodata/surfaces/human_fsLR/tpl-fsLR_space-fsaverage_den-32k_hemi-L_sphere.surf.gii +0 -0
- {plotfig-1.0.0 → plotfig-1.2.1}/src/plotfig/data/neurodata/surfaces/human_fsLR/tpl-fsLR_space-fsaverage_den-32k_hemi-R_sphere.surf.gii +0 -0
- {plotfig-1.0.0 → plotfig-1.2.1}/src/plotfig/data/neurodata/surfaces/macaque_BNA/civm.L.inflated.32k_fs_LR.surf.gii +0 -0
- {plotfig-1.0.0 → plotfig-1.2.1}/src/plotfig/data/neurodata/surfaces/macaque_BNA/civm.L.midthickness.32k_fs_LR.surf.gii +0 -0
- {plotfig-1.0.0 → plotfig-1.2.1}/src/plotfig/data/neurodata/surfaces/macaque_BNA/civm.L.pial.32k_fs_LR.surf.gii +0 -0
- {plotfig-1.0.0 → plotfig-1.2.1}/src/plotfig/data/neurodata/surfaces/macaque_BNA/civm.L.veryinflated.32k_fs_LR.surf.gii +0 -0
- {plotfig-1.0.0 → plotfig-1.2.1}/src/plotfig/data/neurodata/surfaces/macaque_BNA/civm.L.white.32k_fs_LR.surf.gii +0 -0
- {plotfig-1.0.0 → plotfig-1.2.1}/src/plotfig/data/neurodata/surfaces/macaque_BNA/civm.R.inflated.32k_fs_LR.surf.gii +0 -0
- {plotfig-1.0.0 → plotfig-1.2.1}/src/plotfig/data/neurodata/surfaces/macaque_BNA/civm.R.midthickness.32k_fs_LR.surf.gii +0 -0
- {plotfig-1.0.0 → plotfig-1.2.1}/src/plotfig/data/neurodata/surfaces/macaque_BNA/civm.R.pial.32k_fs_LR.surf.gii +0 -0
- {plotfig-1.0.0 → plotfig-1.2.1}/src/plotfig/data/neurodata/surfaces/macaque_BNA/civm.R.veryinflated.32k_fs_LR.surf.gii +0 -0
- {plotfig-1.0.0 → plotfig-1.2.1}/src/plotfig/data/neurodata/surfaces/macaque_BNA/civm.R.white.32k_fs_LR.surf.gii +0 -0
- {plotfig-1.0.0 → plotfig-1.2.1}/src/plotfig/data/neurodata/surfaces/macaque_NMT2/L.gray_surface.inf_300.surf.gii +0 -0
- {plotfig-1.0.0 → plotfig-1.2.1}/src/plotfig/data/neurodata/surfaces/macaque_NMT2/L.gray_surface.surf.gii +0 -0
- {plotfig-1.0.0 → plotfig-1.2.1}/src/plotfig/data/neurodata/surfaces/macaque_NMT2/L.mid_surface.inf_300.surf.gii +0 -0
- {plotfig-1.0.0 → plotfig-1.2.1}/src/plotfig/data/neurodata/surfaces/macaque_NMT2/L.mid_surface.surf.gii +0 -0
- {plotfig-1.0.0 → plotfig-1.2.1}/src/plotfig/data/neurodata/surfaces/macaque_NMT2/L.white_surface.inf_300.surf.gii +0 -0
- {plotfig-1.0.0 → plotfig-1.2.1}/src/plotfig/data/neurodata/surfaces/macaque_NMT2/L.white_surface.surf.gii +0 -0
- {plotfig-1.0.0 → plotfig-1.2.1}/src/plotfig/data/neurodata/surfaces/macaque_NMT2/R.gray_surface.inf_300.surf.gii +0 -0
- {plotfig-1.0.0 → plotfig-1.2.1}/src/plotfig/data/neurodata/surfaces/macaque_NMT2/R.gray_surface.surf.gii +0 -0
- {plotfig-1.0.0 → plotfig-1.2.1}/src/plotfig/data/neurodata/surfaces/macaque_NMT2/R.mid_surface.inf_300.surf.gii +0 -0
- {plotfig-1.0.0 → plotfig-1.2.1}/src/plotfig/data/neurodata/surfaces/macaque_NMT2/R.mid_surface.surf.gii +0 -0
- {plotfig-1.0.0 → plotfig-1.2.1}/src/plotfig/data/neurodata/surfaces/macaque_NMT2/R.white_surface.inf_300.surf.gii +0 -0
- {plotfig-1.0.0 → plotfig-1.2.1}/src/plotfig/data/neurodata/surfaces/macaque_NMT2/R.white_surface.surf.gii +0 -0
- {plotfig-1.0.0 → plotfig-1.2.1}/src/plotfig/data/neurodata/volumes/macaque_NMT2/CHARM5_add_13_sgms_asym.nii.gz +0 -0
|
@@ -0,0 +1,18 @@
|
|
|
1
|
+
name: Sync To Gitee
|
|
2
|
+
|
|
3
|
+
on: [ push, delete, create ]
|
|
4
|
+
|
|
5
|
+
jobs:
|
|
6
|
+
build:
|
|
7
|
+
runs-on: ubuntu-latest
|
|
8
|
+
steps:
|
|
9
|
+
- name: Sync to Gitee
|
|
10
|
+
uses: wearerequired/git-mirror-action@master
|
|
11
|
+
env:
|
|
12
|
+
# 注意在 Settings->Secrets 配置 GITEE_RSA_PRIVATE_KEY
|
|
13
|
+
SSH_PRIVATE_KEY: ${{ secrets.GITEE_RSA_PRIVATE_KEY }}
|
|
14
|
+
with:
|
|
15
|
+
# 注意替换为你的 GitHub 源仓库地址
|
|
16
|
+
source-repo: git@github.com:RicardoRyn/Plot_figure.git
|
|
17
|
+
# 注意替换为你的 Gitee 目标仓库地址
|
|
18
|
+
destination-repo: git@gitee.com:RicardoRyn/Plot_figure.git
|
|
@@ -0,0 +1,29 @@
|
|
|
1
|
+
name: ci
|
|
2
|
+
on:
|
|
3
|
+
push:
|
|
4
|
+
branches:
|
|
5
|
+
- main
|
|
6
|
+
permissions:
|
|
7
|
+
contents: write
|
|
8
|
+
jobs:
|
|
9
|
+
deploy:
|
|
10
|
+
runs-on: ubuntu-latest
|
|
11
|
+
steps:
|
|
12
|
+
- uses: actions/checkout@v4
|
|
13
|
+
- name: Configure Git Credentials
|
|
14
|
+
run: |
|
|
15
|
+
git config user.name github-actions[bot]
|
|
16
|
+
git config user.email 41898282+github-actions[bot]@users.noreply.github.com
|
|
17
|
+
- uses: actions/setup-python@v5
|
|
18
|
+
with:
|
|
19
|
+
python-version: 3.x
|
|
20
|
+
- run: echo "cache_id=$(date --utc '+%V')" >> $GITHUB_ENV
|
|
21
|
+
- uses: actions/cache@v4
|
|
22
|
+
with:
|
|
23
|
+
key: mkdocs-material-${{ env.cache_id }}
|
|
24
|
+
path: .cache
|
|
25
|
+
restore-keys: |
|
|
26
|
+
mkdocs-material-
|
|
27
|
+
- run: pip install uv
|
|
28
|
+
- run: uv sync --only-dev
|
|
29
|
+
- run: uv run mkdocs gh-deploy --force
|
plotfig-1.2.1/.gitignore
ADDED
|
@@ -0,0 +1 @@
|
|
|
1
|
+
3.11
|
|
@@ -0,0 +1,43 @@
|
|
|
1
|
+
## v1.2.1 (2025-07-24)
|
|
2
|
+
|
|
3
|
+
### Fix
|
|
4
|
+
|
|
5
|
+
- **bar**: rename `y_lim_range` to `y_lim` in `plot_one_group_bar_figure`
|
|
6
|
+
|
|
7
|
+
## v1.2.0 (2025-07-24)
|
|
8
|
+
|
|
9
|
+
### Feat
|
|
10
|
+
|
|
11
|
+
- **violin**: add function to plot single-group violin fig
|
|
12
|
+
|
|
13
|
+
### Fix
|
|
14
|
+
|
|
15
|
+
- **matrix**: changed return value to None
|
|
16
|
+
|
|
17
|
+
## v1.1.0 (2025-07-21)
|
|
18
|
+
|
|
19
|
+
### Feat
|
|
20
|
+
|
|
21
|
+
- **corr**: allow hexbin to show dense scatter points in correlation plot
|
|
22
|
+
- **bar**: support gradient color bars and now can change border color
|
|
23
|
+
|
|
24
|
+
## v1.0.0 (2025-07-03)
|
|
25
|
+
|
|
26
|
+
### Feat
|
|
27
|
+
|
|
28
|
+
- **bar**: support plotting single-group bar charts with statistical tests
|
|
29
|
+
- **bar**: support plotting multi-group bars charts
|
|
30
|
+
- **corr**: support combined sactter and line correlation plots
|
|
31
|
+
- **matrix**: support plotting matrix plots (i.e. heatmaps)
|
|
32
|
+
- **surface**: support brain region plots for human, chimpanzee and macaque
|
|
33
|
+
- **circos**: support brain connectivity circos plots
|
|
34
|
+
- **connection**: support glass brain connectivity plots
|
|
35
|
+
|
|
36
|
+
### Fix
|
|
37
|
+
|
|
38
|
+
- **surface**: fix bug where function did not retrun fig only
|
|
39
|
+
- **surface**: fix bug where brain region with zero values were not displayed
|
|
40
|
+
|
|
41
|
+
### Refactor
|
|
42
|
+
|
|
43
|
+
- **src**: refactor code for more readability and maintainability
|
|
@@ -1,60 +1,52 @@
|
|
|
1
|
-
Metadata-Version: 2.4
|
|
2
|
-
Name: plotfig
|
|
3
|
-
Version: 1.
|
|
4
|
-
Summary: Scientific plotting package for Cognitive neuroscience
|
|
5
|
-
Author-email: Ricardo Ryn <ricardoRyn1317@gmail.com>
|
|
6
|
-
|
|
7
|
-
|
|
8
|
-
|
|
9
|
-
|
|
10
|
-
Requires-Dist:
|
|
11
|
-
Requires-Dist:
|
|
12
|
-
Requires-Dist:
|
|
13
|
-
Requires-Dist:
|
|
14
|
-
Requires-Dist:
|
|
15
|
-
Requires-Dist:
|
|
16
|
-
Requires-Dist:
|
|
17
|
-
|
|
18
|
-
|
|
19
|
-
|
|
20
|
-
|
|
21
|
-
|
|
22
|
-
`
|
|
23
|
-
|
|
24
|
-
|
|
25
|
-
|
|
26
|
-
|
|
27
|
-
|
|
28
|
-
|
|
29
|
-
|
|
30
|
-
|
|
31
|
-
|
|
32
|
-
|
|
33
|
-
|
|
34
|
-
|
|
35
|
-
1.
|
|
36
|
-
1.
|
|
37
|
-
1.
|
|
38
|
-
1.
|
|
39
|
-
1.
|
|
40
|
-
|
|
41
|
-
|
|
42
|
-
|
|
43
|
-
|
|
44
|
-
|
|
45
|
-
|
|
46
|
-
|
|
47
|
-
|
|
48
|
-
|
|
49
|
-
|
|
50
|
-
|
|
51
|
-
|
|
52
|
-
|
|
53
|
-
|
|
54
|
-
写堆函数来画图,
|
|
55
|
-
|
|
56
|
-
文档一行无。
|
|
57
|
-
|
|
58
|
-
最自嗨!
|
|
59
|
-
|
|
60
|
-
> 🌟 “调包侠再得一分!”🐶
|
|
1
|
+
Metadata-Version: 2.4
|
|
2
|
+
Name: plotfig
|
|
3
|
+
Version: 1.2.1
|
|
4
|
+
Summary: Scientific plotting package for Cognitive neuroscience
|
|
5
|
+
Author-email: Ricardo Ryn <ricardoRyn1317@gmail.com>
|
|
6
|
+
License-File: LICENSE
|
|
7
|
+
Keywords: neuroscience,plotting,visualization
|
|
8
|
+
Requires-Python: >=3.11
|
|
9
|
+
Requires-Dist: matplotlib>=3.10.1
|
|
10
|
+
Requires-Dist: mne-connectivity>=0.7.0
|
|
11
|
+
Requires-Dist: nibabel>=5.3.2
|
|
12
|
+
Requires-Dist: numpy>=2.2.4
|
|
13
|
+
Requires-Dist: pandas>=2.2.3
|
|
14
|
+
Requires-Dist: plotly>=6.0.1
|
|
15
|
+
Requires-Dist: scipy>=1.15.2
|
|
16
|
+
Requires-Dist: surfplot>=0.2.0
|
|
17
|
+
Description-Content-Type: text/markdown
|
|
18
|
+
|
|
19
|
+
# 简介
|
|
20
|
+
|
|
21
|
+
`plotfig` 是一个面向认知神经科学研究者的 Python 绘图工具包。
|
|
22
|
+
封装了 `matplotlib` 的各种 API,简化了复杂绘图流程。
|
|
23
|
+
可扩展,兼容 `matplotlib` 与 `seaborn`。
|
|
24
|
+
|
|
25
|
+
希望它能让你专注于数据本身而不是琐碎的图形参数调试🥵。
|
|
26
|
+
|
|
27
|
+
[使用教程](https://ricardoryn.github.io/plotfig/)。
|
|
28
|
+
|
|
29
|
+

|
|
30
|
+
|
|
31
|
+
# 功能
|
|
32
|
+
|
|
33
|
+
画图种类:
|
|
34
|
+
1. 单组柱状图
|
|
35
|
+
1. 多组柱状图
|
|
36
|
+
1. 相关点线图
|
|
37
|
+
1. 矩阵图/热图
|
|
38
|
+
1. 图集脑区图
|
|
39
|
+
1. 连线图/circos图
|
|
40
|
+
1. 玻璃脑连接图
|
|
41
|
+
|
|
42
|
+
# 叠甲
|
|
43
|
+
|
|
44
|
+
闲来无事做,
|
|
45
|
+
|
|
46
|
+
写堆函数来画图,
|
|
47
|
+
|
|
48
|
+
文档一行无。
|
|
49
|
+
|
|
50
|
+
最自嗨!
|
|
51
|
+
|
|
52
|
+
> 🌟 “调包侠再得一分!”🐶
|
|
@@ -13,20 +13,13 @@
|
|
|
13
13
|
# 功能
|
|
14
14
|
|
|
15
15
|
画图种类:
|
|
16
|
-
1.
|
|
16
|
+
1. 单组柱状图
|
|
17
17
|
1. 多组柱状图
|
|
18
|
-
1.
|
|
19
|
-
1.
|
|
20
|
-
1.
|
|
21
|
-
|
|
22
|
-
|
|
23
|
-
1. 猕猴CHARM 5-level脑区图
|
|
24
|
-
1. 猕猴CHARM 6-level脑区图
|
|
25
|
-
1. 猕猴BNA脑区图
|
|
26
|
-
1. 连线图(circos图)
|
|
27
|
-
1. 对称circos图
|
|
28
|
-
1. 不对称circos图
|
|
29
|
-
1. 脑连接图
|
|
18
|
+
1. 相关点线图
|
|
19
|
+
1. 矩阵图/热图
|
|
20
|
+
1. 图集脑区图
|
|
21
|
+
1. 连线图/circos图
|
|
22
|
+
1. 玻璃脑连接图
|
|
30
23
|
|
|
31
24
|
# 叠甲
|
|
32
25
|
|
|
@@ -38,4 +31,4 @@
|
|
|
38
31
|
|
|
39
32
|
最自嗨!
|
|
40
33
|
|
|
41
|
-
> 🌟 “调包侠再得一分!”🐶
|
|
34
|
+
> 🌟 “调包侠再得一分!”🐶
|
|
@@ -0,0 +1,18 @@
|
|
|
1
|
+
# API
|
|
2
|
+
|
|
3
|
+
::: plotfig.bar
|
|
4
|
+
options:
|
|
5
|
+
members:
|
|
6
|
+
- plot_one_group_bar_figure
|
|
7
|
+
- plot_one_group_violin_figure
|
|
8
|
+
- plot_multi_group_bar_figure
|
|
9
|
+
|
|
10
|
+
::: plotfig.correlation
|
|
11
|
+
|
|
12
|
+
::: plotfig.matrix
|
|
13
|
+
|
|
14
|
+
::: plotfig.circos
|
|
15
|
+
|
|
16
|
+
::: plotfig.brain_surface
|
|
17
|
+
|
|
18
|
+
::: plotfig.brain_connection
|
|
Binary file
|
|
@@ -0,0 +1,15 @@
|
|
|
1
|
+
# plotfig
|
|
2
|
+
|
|
3
|
+
`plotfig` 是一个面向认知神经科学研究者的 Python 绘图工具包。
|
|
4
|
+
封装了 `matplotlib` 的各种 API,简化了复杂绘图流程。
|
|
5
|
+
可扩展,兼容 `matplotlib` 与 `seaborn`。
|
|
6
|
+
|
|
7
|
+
希望它能让你专注于数据本身而不是琐碎的图形参数调试🥵。
|
|
8
|
+
|
|
9
|
+

|
|
10
|
+
|
|
11
|
+
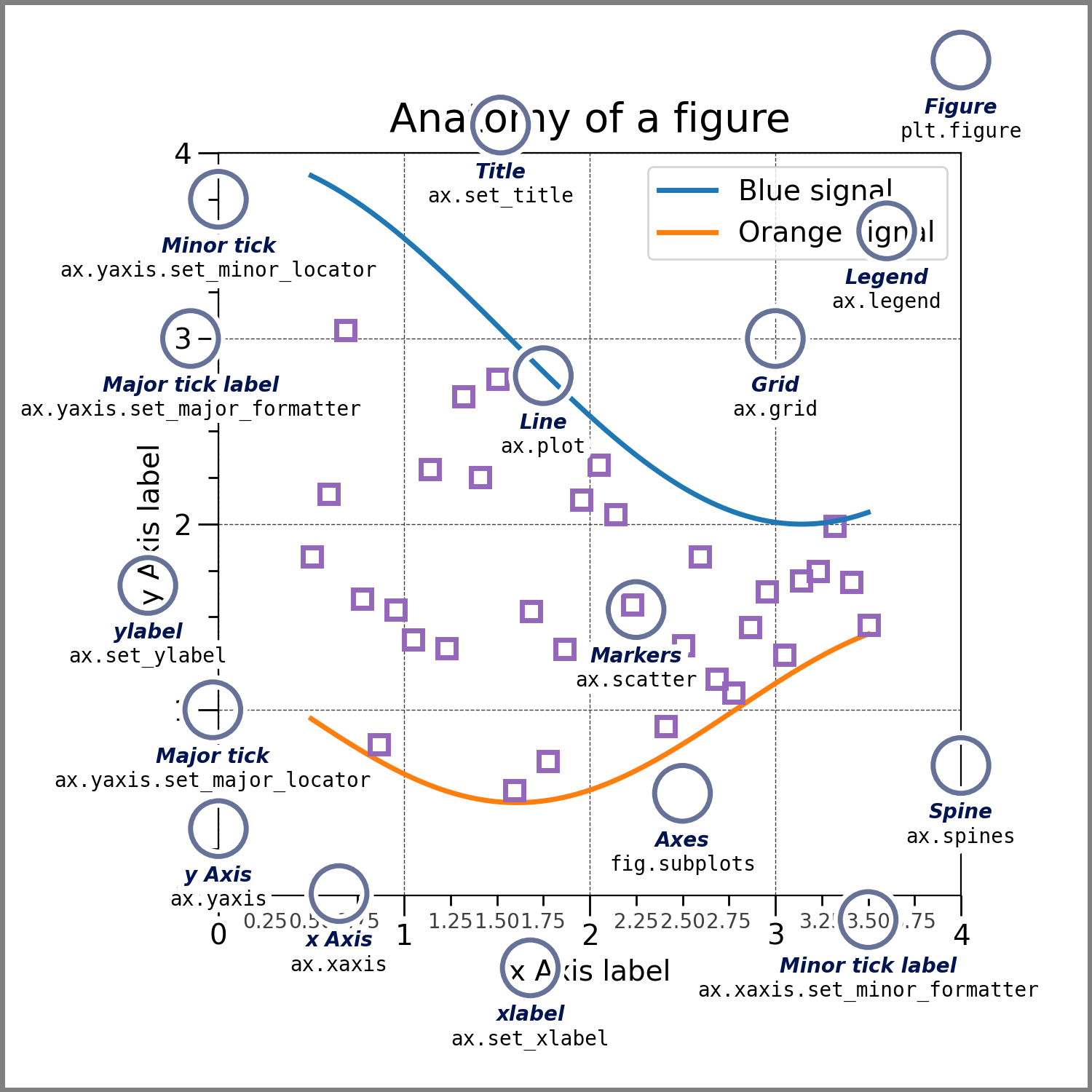
烫知识:一张图上的所有元素[^1]。
|
|
12
|
+
|
|
13
|
+
|
|
14
|
+
[^1]:
|
|
15
|
+
[Quick start guide of matplotlib.](https://matplotlib.org/stable/tutorials/introductory/usage.html#parts-of-a-figure)
|
|
@@ -0,0 +1,37 @@
|
|
|
1
|
+
# 安装
|
|
2
|
+
|
|
3
|
+
`plotfig` 支持通过 `pip` 或源码安装,要求 Python 3.11 及以上版本。
|
|
4
|
+
|
|
5
|
+
---
|
|
6
|
+
|
|
7
|
+
## 📦 使用 pip 安装 <small>(推荐)</small>
|
|
8
|
+
|
|
9
|
+
```bash
|
|
10
|
+
pip install plotfig
|
|
11
|
+
```
|
|
12
|
+
|
|
13
|
+
---
|
|
14
|
+
|
|
15
|
+
## 🛠 从 GitHub 源码安装 <small>(开发版)</small>
|
|
16
|
+
|
|
17
|
+
```bash
|
|
18
|
+
git clone https://github.com/RicardoRyn/plotfig.git
|
|
19
|
+
cd plotfig
|
|
20
|
+
pip install -e .
|
|
21
|
+
```
|
|
22
|
+
|
|
23
|
+
---
|
|
24
|
+
|
|
25
|
+
## 🔧 依赖要求
|
|
26
|
+
|
|
27
|
+
!!! note
|
|
28
|
+
`plotfig` 依赖以下核心库(均会自动安装,但建议提前了解版本要求)。
|
|
29
|
+
|
|
30
|
+
- [matplotlib](https://matplotlib.org/) ≥ 3.10.1
|
|
31
|
+
- [mne-connectivity](https://mne.tools/mne-connectivity/stable/index.html) ≥ 0.7.0
|
|
32
|
+
- [nibabel](https://nipy.org/nibabel/) ≥ 5.3.2
|
|
33
|
+
- [numpy](https://numpy.org/) ≥ 2.2.4
|
|
34
|
+
- [pandas](https://pandas.pydata.org/) ≥ 2.2.3
|
|
35
|
+
- [plotly](https://plotly.com/) ≥ 6.0.1
|
|
36
|
+
- [scipy](https://scipy.org/) ≥ 1.15.2
|
|
37
|
+
- [surfplot](https://github.com/danjgale/surfplot) ≥ 0.2.0
|
|
@@ -0,0 +1,63 @@
|
|
|
1
|
+
# 脑连接图
|
|
2
|
+
|
|
3
|
+
透明的大脑图,可以展示脑区间的连接关系。
|
|
4
|
+
需要准备左右半脑的surface文件、脑区相关的nii.gz文件以及连接矩阵。
|
|
5
|
+
|
|
6
|
+
|
|
7
|
+
```python
|
|
8
|
+
import numpy as np
|
|
9
|
+
from plotfig import *
|
|
10
|
+
|
|
11
|
+
# 生成一个 31x31 的连接矩阵(对称加权矩阵,对角线为0)
|
|
12
|
+
matrix = np.zeros((31, 31))
|
|
13
|
+
matrix[0, 1] = 1
|
|
14
|
+
matrix[0, 2] = 2
|
|
15
|
+
matrix[0, 3] = 3
|
|
16
|
+
matrix[4, 1] = -1
|
|
17
|
+
matrix[4, 2] = -2
|
|
18
|
+
matrix[4, 3] = -3
|
|
19
|
+
matrix = (matrix + matrix.T) / 2 # 矩阵对称
|
|
20
|
+
|
|
21
|
+
connectome = matrix
|
|
22
|
+
|
|
23
|
+
output_file = "./figures/brain_connection.html"
|
|
24
|
+
|
|
25
|
+
lh_surfgii_file = r"e:\6_Self\plot_self_brain_connectivity\103818.L.midthickness.32k_fs_LR.surf.gii"
|
|
26
|
+
rh_surfgii_file = r"e:\6_Self\plot_self_brain_connectivity\103818.R.midthickness.32k_fs_LR.surf.gii"
|
|
27
|
+
niigz_file = rf"e:\6_Self\plot_self_brain_connectivity\human_Self_processing_network.nii.gz"
|
|
28
|
+
|
|
29
|
+
fig = plot_brain_connection_figure(
|
|
30
|
+
connectome,
|
|
31
|
+
lh_surfgii_file=lh_surfgii_file,
|
|
32
|
+
rh_surfgii_file=rh_surfgii_file,
|
|
33
|
+
niigz_file=niigz_file,
|
|
34
|
+
output_file=output_file,
|
|
35
|
+
scale_method="width",
|
|
36
|
+
line_width=10,
|
|
37
|
+
)
|
|
38
|
+
```
|
|
39
|
+
|
|
40
|
+
|
|
41
|
+
|
|
42
|
+

|
|
43
|
+
|
|
44
|
+
|
|
45
|
+
|
|
46
|
+
html文件可以在浏览器中交互。可以手动截图,也可以使用以下命令来生成图片。
|
|
47
|
+
|
|
48
|
+
|
|
49
|
+
```python
|
|
50
|
+
from pathlib import Path
|
|
51
|
+
|
|
52
|
+
|
|
53
|
+
Path(f"./figures/brain_connection").mkdir(parents=True, exist_ok=True) # 新建文件夹保存帧图
|
|
54
|
+
save_brain_connection_frames(fig, output_dir=rf"./figures/brain_connection", n_frames=36)
|
|
55
|
+
```
|
|
56
|
+
|
|
57
|
+
100%|██████████| 36/36 [02:01<00:00, 3.37s/it]
|
|
58
|
+
|
|
59
|
+
保存了 36 张图片在 ./figures/brain_connection
|
|
60
|
+
|
|
61
|
+
|
|
62
|
+
|
|
63
|
+
|
|
Binary file
|