ms2rescore 3.2.0.dev2__tar.gz → 3.2.0.post1__tar.gz
This diff represents the content of publicly available package versions that have been released to one of the supported registries. The information contained in this diff is provided for informational purposes only and reflects changes between package versions as they appear in their respective public registries.
- ms2rescore-3.2.0.post1/PKG-INFO +358 -0
- ms2rescore-3.2.0.post1/ms2rescore/__init__.py +34 -0
- {ms2rescore-3.2.0.dev2 → ms2rescore-3.2.0.post1}/ms2rescore/__main__.py +24 -7
- ms2rescore-3.2.0.post1/ms2rescore/_version.py +135 -0
- {ms2rescore-3.2.0.dev2 → ms2rescore-3.2.0.post1}/ms2rescore/config_parser.py +33 -1
- {ms2rescore-3.2.0.dev2 → ms2rescore-3.2.0.post1}/ms2rescore/core.py +9 -1
- {ms2rescore-3.2.0.dev2 → ms2rescore-3.2.0.post1}/ms2rescore/feature_generators/__init__.py +3 -2
- {ms2rescore-3.2.0.dev2 → ms2rescore-3.2.0.post1}/ms2rescore/feature_generators/base.py +1 -1
- {ms2rescore-3.2.0.dev2 → ms2rescore-3.2.0.post1}/ms2rescore/feature_generators/deeplc.py +8 -6
- {ms2rescore-3.2.0.dev2 → ms2rescore-3.2.0.post1}/ms2rescore/gui/app.py +60 -5
- {ms2rescore-3.2.0.dev2 → ms2rescore-3.2.0.post1}/ms2rescore/gui/function2ctk.py +43 -10
- {ms2rescore-3.2.0.dev2 → ms2rescore-3.2.0.post1}/ms2rescore/package_data/config_schema.json +8 -3
- {ms2rescore-3.2.0.dev2 → ms2rescore-3.2.0.post1}/ms2rescore/parse_psms.py +28 -9
- ms2rescore-3.2.0.post1/ms2rescore/parse_spectra.py +236 -0
- {ms2rescore-3.2.0.dev2 → ms2rescore-3.2.0.post1}/ms2rescore/report/generate.py +5 -3
- {ms2rescore-3.2.0.dev2 → ms2rescore-3.2.0.post1}/ms2rescore/utils.py +2 -1
- {ms2rescore-3.2.0.dev2 → ms2rescore-3.2.0.post1}/pyproject.toml +16 -8
- ms2rescore-3.2.0.dev2/PKG-INFO +0 -173
- ms2rescore-3.2.0.dev2/ms2rescore/__init__.py +0 -20
- ms2rescore-3.2.0.dev2/ms2rescore/parse_spectra.py +0 -152
- {ms2rescore-3.2.0.dev2 → ms2rescore-3.2.0.post1}/LICENSE +0 -0
- {ms2rescore-3.2.0.dev2 → ms2rescore-3.2.0.post1}/README.md +0 -0
- {ms2rescore-3.2.0.dev2 → ms2rescore-3.2.0.post1}/ms2rescore/exceptions.py +0 -0
- {ms2rescore-3.2.0.dev2 → ms2rescore-3.2.0.post1}/ms2rescore/feature_generators/basic.py +0 -0
- {ms2rescore-3.2.0.dev2 → ms2rescore-3.2.0.post1}/ms2rescore/feature_generators/im2deep.py +0 -0
- {ms2rescore-3.2.0.dev2 → ms2rescore-3.2.0.post1}/ms2rescore/feature_generators/ionmob.py +0 -0
- {ms2rescore-3.2.0.dev2 → ms2rescore-3.2.0.post1}/ms2rescore/feature_generators/maxquant.py +0 -0
- {ms2rescore-3.2.0.dev2 → ms2rescore-3.2.0.post1}/ms2rescore/feature_generators/ms2pip.py +0 -0
- {ms2rescore-3.2.0.dev2 → ms2rescore-3.2.0.post1}/ms2rescore/gui/__init__.py +0 -0
- {ms2rescore-3.2.0.dev2 → ms2rescore-3.2.0.post1}/ms2rescore/gui/__main__.py +0 -0
- {ms2rescore-3.2.0.dev2 → ms2rescore-3.2.0.post1}/ms2rescore/gui/widgets.py +0 -0
- {ms2rescore-3.2.0.dev2 → ms2rescore-3.2.0.post1}/ms2rescore/package_data/__init__.py +0 -0
- {ms2rescore-3.2.0.dev2 → ms2rescore-3.2.0.post1}/ms2rescore/package_data/config_default.json +0 -0
- {ms2rescore-3.2.0.dev2 → ms2rescore-3.2.0.post1}/ms2rescore/package_data/config_default_tims.json +0 -0
- {ms2rescore-3.2.0.dev2 → ms2rescore-3.2.0.post1}/ms2rescore/package_data/img/__init__.py +0 -0
- {ms2rescore-3.2.0.dev2 → ms2rescore-3.2.0.post1}/ms2rescore/package_data/img/comments_icon_black.png +0 -0
- {ms2rescore-3.2.0.dev2 → ms2rescore-3.2.0.post1}/ms2rescore/package_data/img/comments_icon_white.png +0 -0
- {ms2rescore-3.2.0.dev2 → ms2rescore-3.2.0.post1}/ms2rescore/package_data/img/config_icon.png +0 -0
- {ms2rescore-3.2.0.dev2 → ms2rescore-3.2.0.post1}/ms2rescore/package_data/img/docs_icon_black.png +0 -0
- {ms2rescore-3.2.0.dev2 → ms2rescore-3.2.0.post1}/ms2rescore/package_data/img/docs_icon_white.png +0 -0
- {ms2rescore-3.2.0.dev2 → ms2rescore-3.2.0.post1}/ms2rescore/package_data/img/github_icon_black.png +0 -0
- {ms2rescore-3.2.0.dev2 → ms2rescore-3.2.0.post1}/ms2rescore/package_data/img/github_icon_white.png +0 -0
- {ms2rescore-3.2.0.dev2 → ms2rescore-3.2.0.post1}/ms2rescore/package_data/img/ms2rescore_logo.png +0 -0
- {ms2rescore-3.2.0.dev2 → ms2rescore-3.2.0.post1}/ms2rescore/package_data/img/program_icon.ico +0 -0
- {ms2rescore-3.2.0.dev2 → ms2rescore-3.2.0.post1}/ms2rescore/report/__init__.py +0 -0
- {ms2rescore-3.2.0.dev2 → ms2rescore-3.2.0.post1}/ms2rescore/report/__main__.py +0 -0
- {ms2rescore-3.2.0.dev2 → ms2rescore-3.2.0.post1}/ms2rescore/report/charts.py +0 -0
- {ms2rescore-3.2.0.dev2 → ms2rescore-3.2.0.post1}/ms2rescore/report/templates/__init__.py +0 -0
- {ms2rescore-3.2.0.dev2 → ms2rescore-3.2.0.post1}/ms2rescore/report/templates/about.html +0 -0
- {ms2rescore-3.2.0.dev2 → ms2rescore-3.2.0.post1}/ms2rescore/report/templates/base.html +0 -0
- {ms2rescore-3.2.0.dev2 → ms2rescore-3.2.0.post1}/ms2rescore/report/templates/config.html +0 -0
- {ms2rescore-3.2.0.dev2 → ms2rescore-3.2.0.post1}/ms2rescore/report/templates/features.html +0 -0
- {ms2rescore-3.2.0.dev2 → ms2rescore-3.2.0.post1}/ms2rescore/report/templates/log.html +0 -0
- {ms2rescore-3.2.0.dev2 → ms2rescore-3.2.0.post1}/ms2rescore/report/templates/metadata.html +0 -0
- {ms2rescore-3.2.0.dev2 → ms2rescore-3.2.0.post1}/ms2rescore/report/templates/overview.html +0 -0
- {ms2rescore-3.2.0.dev2 → ms2rescore-3.2.0.post1}/ms2rescore/report/templates/stats-card.html +0 -0
- {ms2rescore-3.2.0.dev2 → ms2rescore-3.2.0.post1}/ms2rescore/report/templates/style.html +0 -0
- {ms2rescore-3.2.0.dev2 → ms2rescore-3.2.0.post1}/ms2rescore/report/templates/target-decoy.html +0 -0
- {ms2rescore-3.2.0.dev2 → ms2rescore-3.2.0.post1}/ms2rescore/report/templates/texts.toml +0 -0
- {ms2rescore-3.2.0.dev2 → ms2rescore-3.2.0.post1}/ms2rescore/report/utils.py +0 -0
- {ms2rescore-3.2.0.dev2 → ms2rescore-3.2.0.post1}/ms2rescore/rescoring_engines/__init__.py +0 -0
- {ms2rescore-3.2.0.dev2 → ms2rescore-3.2.0.post1}/ms2rescore/rescoring_engines/mokapot.py +0 -0
- {ms2rescore-3.2.0.dev2 → ms2rescore-3.2.0.post1}/ms2rescore/rescoring_engines/percolator.py +0 -0
|
@@ -0,0 +1,358 @@
|
|
|
1
|
+
Metadata-Version: 2.3
|
|
2
|
+
Name: ms2rescore
|
|
3
|
+
Version: 3.2.0.post1
|
|
4
|
+
Summary: Modular and user-friendly platform for AI-assisted rescoring of peptide identifications.
|
|
5
|
+
Keywords: MS2Rescore,MS2PIP,DeepLC,Percolator,proteomics,mass spectrometry,peptide identification,rescoring,machine learning
|
|
6
|
+
Author: Ralf Gabriels, Arthur Declercq, Ana Sílvia C. Silva, Robbin Bouwmeester, Louise Buur
|
|
7
|
+
Author-email: Ralf Gabriels <ralf@gabriels.dev>, Arthur Declercq <arthur.declercq@ugent.be>
|
|
8
|
+
License: Apache License
|
|
9
|
+
Version 2.0, January 2004
|
|
10
|
+
http://www.apache.org/licenses/
|
|
11
|
+
|
|
12
|
+
TERMS AND CONDITIONS FOR USE, REPRODUCTION, AND DISTRIBUTION
|
|
13
|
+
|
|
14
|
+
1. Definitions.
|
|
15
|
+
|
|
16
|
+
"License" shall mean the terms and conditions for use, reproduction,
|
|
17
|
+
and distribution as defined by Sections 1 through 9 of this document.
|
|
18
|
+
|
|
19
|
+
"Licensor" shall mean the copyright owner or entity authorized by
|
|
20
|
+
the copyright owner that is granting the License.
|
|
21
|
+
|
|
22
|
+
"Legal Entity" shall mean the union of the acting entity and all
|
|
23
|
+
other entities that control, are controlled by, or are under common
|
|
24
|
+
control with that entity. For the purposes of this definition,
|
|
25
|
+
"control" means (i) the power, direct or indirect, to cause the
|
|
26
|
+
direction or management of such entity, whether by contract or
|
|
27
|
+
otherwise, or (ii) ownership of fifty percent (50%) or more of the
|
|
28
|
+
outstanding shares, or (iii) beneficial ownership of such entity.
|
|
29
|
+
|
|
30
|
+
"You" (or "Your") shall mean an individual or Legal Entity
|
|
31
|
+
exercising permissions granted by this License.
|
|
32
|
+
|
|
33
|
+
"Source" form shall mean the preferred form for making modifications,
|
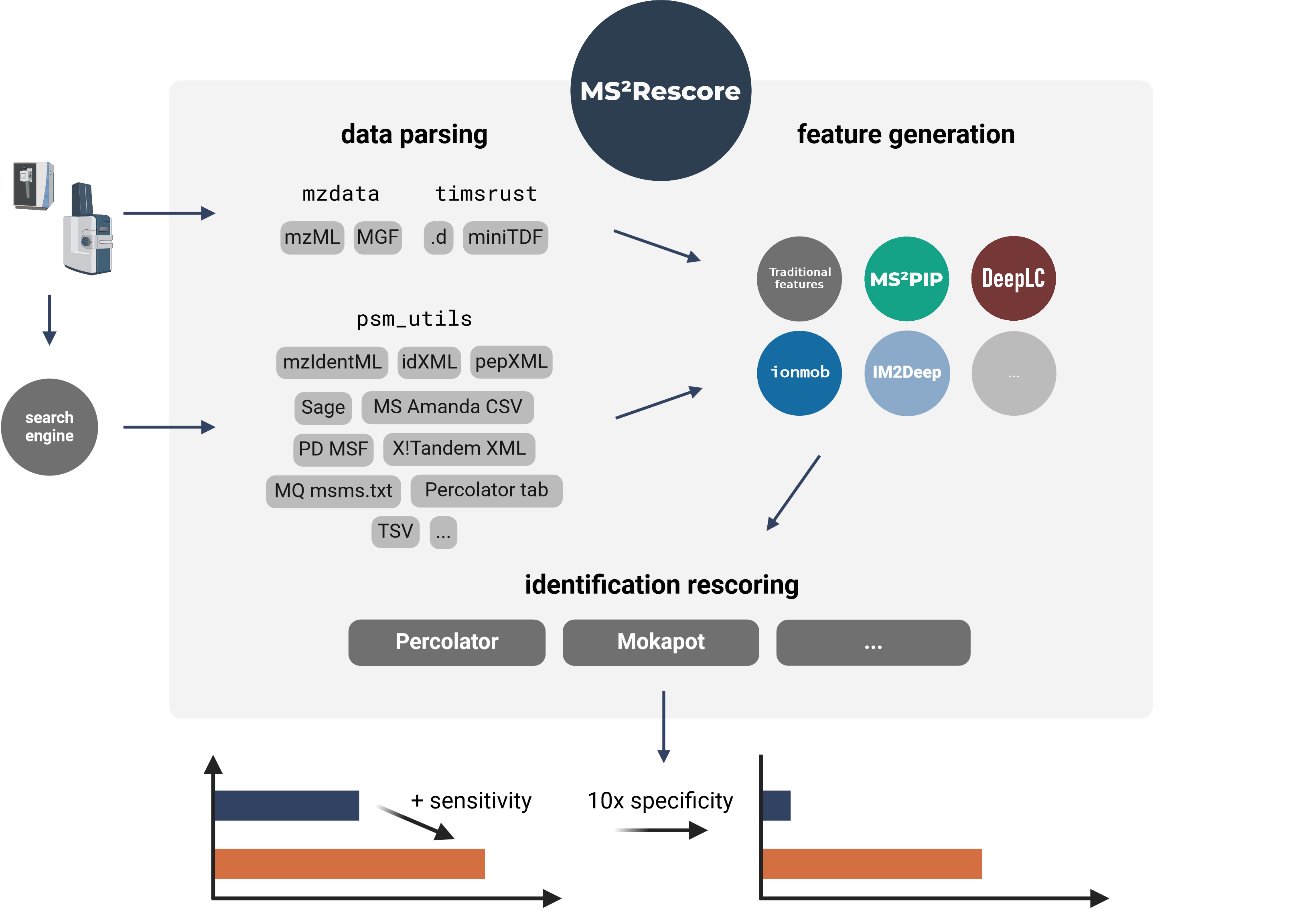
|
34
|
+
including but not limited to software source code, documentation
|
|
35
|
+
source, and configuration files.
|
|
36
|
+
|
|
37
|
+
"Object" form shall mean any form resulting from mechanical
|
|
38
|
+
transformation or translation of a Source form, including but
|
|
39
|
+
not limited to compiled object code, generated documentation,
|
|
40
|
+
and conversions to other media types.
|
|
41
|
+
|
|
42
|
+
"Work" shall mean the work of authorship, whether in Source or
|
|
43
|
+
Object form, made available under the License, as indicated by a
|
|
44
|
+
copyright notice that is included in or attached to the work
|
|
45
|
+
(an example is provided in the Appendix below).
|
|
46
|
+
|
|
47
|
+
"Derivative Works" shall mean any work, whether in Source or Object
|
|
48
|
+
form, that is based on (or derived from) the Work and for which the
|
|
49
|
+
editorial revisions, annotations, elaborations, or other modifications
|
|
50
|
+
represent, as a whole, an original work of authorship. For the purposes
|
|
51
|
+
of this License, Derivative Works shall not include works that remain
|
|
52
|
+
separable from, or merely link (or bind by name) to the interfaces of,
|
|
53
|
+
the Work and Derivative Works thereof.
|
|
54
|
+
|
|
55
|
+
"Contribution" shall mean any work of authorship, including
|
|
56
|
+
the original version of the Work and any modifications or additions
|
|
57
|
+
to that Work or Derivative Works thereof, that is intentionally
|
|
58
|
+
submitted to Licensor for inclusion in the Work by the copyright owner
|
|
59
|
+
or by an individual or Legal Entity authorized to submit on behalf of
|
|
60
|
+
the copyright owner. For the purposes of this definition, "submitted"
|
|
61
|
+
means any form of electronic, verbal, or written communication sent
|
|
62
|
+
to the Licensor or its representatives, including but not limited to
|
|
63
|
+
communication on electronic mailing lists, source code control systems,
|
|
64
|
+
and issue tracking systems that are managed by, or on behalf of, the
|
|
65
|
+
Licensor for the purpose of discussing and improving the Work, but
|
|
66
|
+
excluding communication that is conspicuously marked or otherwise
|
|
67
|
+
designated in writing by the copyright owner as "Not a Contribution."
|
|
68
|
+
|
|
69
|
+
"Contributor" shall mean Licensor and any individual or Legal Entity
|
|
70
|
+
on behalf of whom a Contribution has been received by Licensor and
|
|
71
|
+
subsequently incorporated within the Work.
|
|
72
|
+
|
|
73
|
+
2. Grant of Copyright License. Subject to the terms and conditions of
|
|
74
|
+
this License, each Contributor hereby grants to You a perpetual,
|
|
75
|
+
worldwide, non-exclusive, no-charge, royalty-free, irrevocable
|
|
76
|
+
copyright license to reproduce, prepare Derivative Works of,
|
|
77
|
+
publicly display, publicly perform, sublicense, and distribute the
|
|
78
|
+
Work and such Derivative Works in Source or Object form.
|
|
79
|
+
|
|
80
|
+
3. Grant of Patent License. Subject to the terms and conditions of
|
|
81
|
+
this License, each Contributor hereby grants to You a perpetual,
|
|
82
|
+
worldwide, non-exclusive, no-charge, royalty-free, irrevocable
|
|
83
|
+
(except as stated in this section) patent license to make, have made,
|
|
84
|
+
use, offer to sell, sell, import, and otherwise transfer the Work,
|
|
85
|
+
where such license applies only to those patent claims licensable
|
|
86
|
+
by such Contributor that are necessarily infringed by their
|
|
87
|
+
Contribution(s) alone or by combination of their Contribution(s)
|
|
88
|
+
with the Work to which such Contribution(s) was submitted. If You
|
|
89
|
+
institute patent litigation against any entity (including a
|
|
90
|
+
cross-claim or counterclaim in a lawsuit) alleging that the Work
|
|
91
|
+
or a Contribution incorporated within the Work constitutes direct
|
|
92
|
+
or contributory patent infringement, then any patent licenses
|
|
93
|
+
granted to You under this License for that Work shall terminate
|
|
94
|
+
as of the date such litigation is filed.
|
|
95
|
+
|
|
96
|
+
4. Redistribution. You may reproduce and distribute copies of the
|
|
97
|
+
Work or Derivative Works thereof in any medium, with or without
|
|
98
|
+
modifications, and in Source or Object form, provided that You
|
|
99
|
+
meet the following conditions:
|
|
100
|
+
|
|
101
|
+
(a) You must give any other recipients of the Work or
|
|
102
|
+
Derivative Works a copy of this License; and
|
|
103
|
+
|
|
104
|
+
(b) You must cause any modified files to carry prominent notices
|
|
105
|
+
stating that You changed the files; and
|
|
106
|
+
|
|
107
|
+
(c) You must retain, in the Source form of any Derivative Works
|
|
108
|
+
that You distribute, all copyright, patent, trademark, and
|
|
109
|
+
attribution notices from the Source form of the Work,
|
|
110
|
+
excluding those notices that do not pertain to any part of
|
|
111
|
+
the Derivative Works; and
|
|
112
|
+
|
|
113
|
+
(d) If the Work includes a "NOTICE" text file as part of its
|
|
114
|
+
distribution, then any Derivative Works that You distribute must
|
|
115
|
+
include a readable copy of the attribution notices contained
|
|
116
|
+
within such NOTICE file, excluding those notices that do not
|
|
117
|
+
pertain to any part of the Derivative Works, in at least one
|
|
118
|
+
of the following places: within a NOTICE text file distributed
|
|
119
|
+
as part of the Derivative Works; within the Source form or
|
|
120
|
+
documentation, if provided along with the Derivative Works; or,
|
|
121
|
+
within a display generated by the Derivative Works, if and
|
|
122
|
+
wherever such third-party notices normally appear. The contents
|
|
123
|
+
of the NOTICE file are for informational purposes only and
|
|
124
|
+
do not modify the License. You may add Your own attribution
|
|
125
|
+
notices within Derivative Works that You distribute, alongside
|
|
126
|
+
or as an addendum to the NOTICE text from the Work, provided
|
|
127
|
+
that such additional attribution notices cannot be construed
|
|
128
|
+
as modifying the License.
|
|
129
|
+
|
|
130
|
+
You may add Your own copyright statement to Your modifications and
|
|
131
|
+
may provide additional or different license terms and conditions
|
|
132
|
+
for use, reproduction, or distribution of Your modifications, or
|
|
133
|
+
for any such Derivative Works as a whole, provided Your use,
|
|
134
|
+
reproduction, and distribution of the Work otherwise complies with
|
|
135
|
+
the conditions stated in this License.
|
|
136
|
+
|
|
137
|
+
5. Submission of Contributions. Unless You explicitly state otherwise,
|
|
138
|
+
any Contribution intentionally submitted for inclusion in the Work
|
|
139
|
+
by You to the Licensor shall be under the terms and conditions of
|
|
140
|
+
this License, without any additional terms or conditions.
|
|
141
|
+
Notwithstanding the above, nothing herein shall supersede or modify
|
|
142
|
+
the terms of any separate license agreement you may have executed
|
|
143
|
+
with Licensor regarding such Contributions.
|
|
144
|
+
|
|
145
|
+
6. Trademarks. This License does not grant permission to use the trade
|
|
146
|
+
names, trademarks, service marks, or product names of the Licensor,
|
|
147
|
+
except as required for reasonable and customary use in describing the
|
|
148
|
+
origin of the Work and reproducing the content of the NOTICE file.
|
|
149
|
+
|
|
150
|
+
7. Disclaimer of Warranty. Unless required by applicable law or
|
|
151
|
+
agreed to in writing, Licensor provides the Work (and each
|
|
152
|
+
Contributor provides its Contributions) on an "AS IS" BASIS,
|
|
153
|
+
WITHOUT WARRANTIES OR CONDITIONS OF ANY KIND, either express or
|
|
154
|
+
implied, including, without limitation, any warranties or conditions
|
|
155
|
+
of TITLE, NON-INFRINGEMENT, MERCHANTABILITY, or FITNESS FOR A
|
|
156
|
+
PARTICULAR PURPOSE. You are solely responsible for determining the
|
|
157
|
+
appropriateness of using or redistributing the Work and assume any
|
|
158
|
+
risks associated with Your exercise of permissions under this License.
|
|
159
|
+
|
|
160
|
+
8. Limitation of Liability. In no event and under no legal theory,
|
|
161
|
+
whether in tort (including negligence), contract, or otherwise,
|
|
162
|
+
unless required by applicable law (such as deliberate and grossly
|
|
163
|
+
negligent acts) or agreed to in writing, shall any Contributor be
|
|
164
|
+
liable to You for damages, including any direct, indirect, special,
|
|
165
|
+
incidental, or consequential damages of any character arising as a
|
|
166
|
+
result of this License or out of the use or inability to use the
|
|
167
|
+
Work (including but not limited to damages for loss of goodwill,
|
|
168
|
+
work stoppage, computer failure or malfunction, or any and all
|
|
169
|
+
other commercial damages or losses), even if such Contributor
|
|
170
|
+
has been advised of the possibility of such damages.
|
|
171
|
+
|
|
172
|
+
9. Accepting Warranty or Additional Liability. While redistributing
|
|
173
|
+
the Work or Derivative Works thereof, You may choose to offer,
|
|
174
|
+
and charge a fee for, acceptance of support, warranty, indemnity,
|
|
175
|
+
or other liability obligations and/or rights consistent with this
|
|
176
|
+
License. However, in accepting such obligations, You may act only
|
|
177
|
+
on Your own behalf and on Your sole responsibility, not on behalf
|
|
178
|
+
of any other Contributor, and only if You agree to indemnify,
|
|
179
|
+
defend, and hold each Contributor harmless for any liability
|
|
180
|
+
incurred by, or claims asserted against, such Contributor by reason
|
|
181
|
+
of your accepting any such warranty or additional liability.
|
|
182
|
+
|
|
183
|
+
END OF TERMS AND CONDITIONS
|
|
184
|
+
|
|
185
|
+
APPENDIX: How to apply the Apache License to your work.
|
|
186
|
+
|
|
187
|
+
To apply the Apache License to your work, attach the following
|
|
188
|
+
boilerplate notice, with the fields enclosed by brackets "[]"
|
|
189
|
+
replaced with your own identifying information. (Don't include
|
|
190
|
+
the brackets!) The text should be enclosed in the appropriate
|
|
191
|
+
comment syntax for the file format. We also recommend that a
|
|
192
|
+
file or class name and description of purpose be included on the
|
|
193
|
+
same "printed page" as the copyright notice for easier
|
|
194
|
+
identification within third-party archives.
|
|
195
|
+
|
|
196
|
+
Copyright [yyyy] [name of copyright owner]
|
|
197
|
+
|
|
198
|
+
Licensed under the Apache License, Version 2.0 (the "License");
|
|
199
|
+
you may not use this file except in compliance with the License.
|
|
200
|
+
You may obtain a copy of the License at
|
|
201
|
+
|
|
202
|
+
http://www.apache.org/licenses/LICENSE-2.0
|
|
203
|
+
|
|
204
|
+
Unless required by applicable law or agreed to in writing, software
|
|
205
|
+
distributed under the License is distributed on an "AS IS" BASIS,
|
|
206
|
+
WITHOUT WARRANTIES OR CONDITIONS OF ANY KIND, either express or implied.
|
|
207
|
+
See the License for the specific language governing permissions and
|
|
208
|
+
limitations under the License.
|
|
209
|
+
Classifier: Intended Audience :: Science/Research
|
|
210
|
+
Classifier: License :: OSI Approved :: Apache Software License
|
|
211
|
+
Classifier: Operating System :: OS Independent
|
|
212
|
+
Classifier: Programming Language :: Python :: 3 :: Only
|
|
213
|
+
Classifier: Topic :: Scientific/Engineering :: Bio-Informatics
|
|
214
|
+
Classifier: Development Status :: 5 - Production/Stable
|
|
215
|
+
Requires-Dist: cascade-config>=0.4.0
|
|
216
|
+
Requires-Dist: click>=7
|
|
217
|
+
Requires-Dist: customtkinter>=5,<6
|
|
218
|
+
Requires-Dist: deeplc>=3.1
|
|
219
|
+
Requires-Dist: deeplcretrainer
|
|
220
|
+
Requires-Dist: im2deep>=0.3.1
|
|
221
|
+
Requires-Dist: jinja2>=3
|
|
222
|
+
Requires-Dist: lxml>=4.5
|
|
223
|
+
Requires-Dist: mokapot==0.10
|
|
224
|
+
Requires-Dist: ms2pip>=4.0.0
|
|
225
|
+
Requires-Dist: ms2rescore-rs>=0.4.3
|
|
226
|
+
Requires-Dist: numpy>=1.25
|
|
227
|
+
Requires-Dist: packaging>=25.0
|
|
228
|
+
Requires-Dist: pandas>=1
|
|
229
|
+
Requires-Dist: plotly>=5
|
|
230
|
+
Requires-Dist: psm-utils>=1.1
|
|
231
|
+
Requires-Dist: pyteomics>=4.7.2
|
|
232
|
+
Requires-Dist: rich>=12
|
|
233
|
+
Requires-Dist: tomli>=2 ; python_full_version < '3.11'
|
|
234
|
+
Requires-Dist: pyopenms>=3.3 ; extra == 'idxml'
|
|
235
|
+
Requires-Dist: ionmob>=0.2 ; extra == 'ionmob'
|
|
236
|
+
Requires-Dist: tensorflow ; extra == 'ionmob'
|
|
237
|
+
Requires-Python: >=3.10
|
|
238
|
+
Project-URL: CompOmics, https://www.compomics.com
|
|
239
|
+
Project-URL: GitHub, https://github.com/compomics/ms2rescore
|
|
240
|
+
Project-URL: PyPi, https://pypi.org/project/ms2rescore/
|
|
241
|
+
Project-URL: ReadTheDocs, https://ms2rescore.readthedocs.io
|
|
242
|
+
Provides-Extra: idxml
|
|
243
|
+
Provides-Extra: ionmob
|
|
244
|
+
Description-Content-Type: text/markdown
|
|
245
|
+
|
|
246
|
+
<img src="https://github.com/compomics/ms2rescore/raw/main/img/ms2rescore_logo.png" width="150" height="150" alt="MS²Rescore"/>
|
|
247
|
+
<br/><br/>
|
|
248
|
+
|
|
249
|
+
[](https://github.com/compomics/ms2rescore/releases)
|
|
250
|
+
[](https://pypi.org/project/ms2rescore/)
|
|
251
|
+
[](https://github.com/compomics/ms2rescore/actions/)
|
|
252
|
+
[](https://github.com/compomics/ms2rescore/issues)
|
|
253
|
+
[](https://www.apache.org/licenses/LICENSE-2.0)
|
|
254
|
+
[](https://github.com/compomics/ms2rescore/commits/)
|
|
255
|
+
|
|
256
|
+
Modular and user-friendly platform for AI-assisted rescoring of peptide identifications
|
|
257
|
+
|
|
258
|
+
## About MS²Rescore
|
|
259
|
+
|
|
260
|
+
MS²Rescore performs ultra-sensitive peptide identification rescoring with LC-MS predictors such as
|
|
261
|
+
[MS²PIP][ms2pip] and [DeepLC][deeplc], and with ML-driven rescoring engines
|
|
262
|
+
[Percolator][percolator] or [Mokapot][mokapot]. This results in more confident peptide
|
|
263
|
+
identifications, which allows you to get **more peptide IDs** at the same false discovery rate
|
|
264
|
+
(FDR) threshold, or to set a **more stringent FDR threshold** while still retaining a similar
|
|
265
|
+
number of peptide IDs. MS²Rescore is **ideal for challenging proteomics identification workflows**,
|
|
266
|
+
such as proteogenomics, metaproteomics, or immunopeptidomics.
|
|
267
|
+
|
|
268
|
+
|
|
269
|
+
|
|
270
|
+
MS²Rescore can read peptide identifications in any format supported by [psm_utils][psm_utils]
|
|
271
|
+
(see [Supported file formats][file-formats]) and has been tested with various search engines output
|
|
272
|
+
files:
|
|
273
|
+
|
|
274
|
+
- [MS Amanda](http://ms.imp.ac.at/?goto=msamanda) `.csv`
|
|
275
|
+
- [Sage](https://github.com/lazear/sage) `.sage.tsv`
|
|
276
|
+
- [PeptideShaker](https://compomics.github.io/projects/peptide-shaker.html) `.mzid`
|
|
277
|
+
- [ProteomeDiscoverer](#)`.msf`
|
|
278
|
+
- [MSGFPlus](https://omics.pnl.gov/software/ms-gf) `.mzid`
|
|
279
|
+
- [Mascot](https://www.matrixscience.com/) `.mzid`
|
|
280
|
+
- [MaxQuant](https://www.maxquant.org/) `msms.txt`
|
|
281
|
+
- [X!Tandem](https://www.thegpm.org/tandem/) `.xml`
|
|
282
|
+
- [PEAKS](https://www.bioinfor.com/peaksdb/) `.mzid`
|
|
283
|
+
|
|
284
|
+
MS²Rescore is available as a [desktop application][desktop], a [command line tool][cli], and a
|
|
285
|
+
[modular Python API][python-package].
|
|
286
|
+
|
|
287
|
+
## TIMS²Rescore: Direct support for DDA-PASEF data
|
|
288
|
+
|
|
289
|
+
MS²Rescore v3.1+ includes TIMS²Rescore, a usage mode with specialized default configurations for
|
|
290
|
+
DDA-PASEF data from timsTOF instruments. TIMS²Rescore makes use of new MS²PIP prediction models for
|
|
291
|
+
timsTOF fragmentation and IM2Deep for ion mobility separation. Bruker .d and miniTDF spectrum
|
|
292
|
+
files are directly supported through the [timsrust](https://github.com/MannLabs/timsrust) library.
|
|
293
|
+
|
|
294
|
+
Checkout our [paper](https://doi.org/10.1021/acs.jproteome.4c00609) for more information and the
|
|
295
|
+
[TIMS²Rescore documentation][tims2rescore] to get started.
|
|
296
|
+
|
|
297
|
+
## Citing
|
|
298
|
+
|
|
299
|
+
**Latest MS²Rescore publication:**
|
|
300
|
+
|
|
301
|
+
> **MS²Rescore 3.0 is a modular, flexible, and user-friendly platform to boost peptide identifications, as showcased with MS Amanda 3.0.**
|
|
302
|
+
> Louise Marie Buur*, Arthur Declercq*, Marina Strobl, Robbin Bouwmeester, Sven Degroeve, Lennart Martens, Viktoria Dorfer*, and Ralf Gabriels*.
|
|
303
|
+
> _Journal of Proteome Research_ (2024) [doi:10.1021/acs.jproteome.3c00785](https://doi.org/10.1021/acs.jproteome.3c00785) <br/> \*contributed equally <span class="__dimensions_badge_embed__" data-doi="10.1021/acs.jproteome.3c00785" data-hide-zero-citations="true" data-style="small_rectangle"></span>
|
|
304
|
+
|
|
305
|
+
**MS²Rescore for immunopeptidomics:**
|
|
306
|
+
|
|
307
|
+
> **MS²Rescore: Data-driven rescoring dramatically boosts immunopeptide identification rates.**
|
|
308
|
+
> Arthur Declercq, Robbin Bouwmeester, Aurélie Hirschler, Christine Carapito, Sven Degroeve, Lennart Martens, and Ralf Gabriels.
|
|
309
|
+
> _Molecular & Cellular Proteomics_ (2021) [doi:10.1016/j.mcpro.2022.100266](https://doi.org/10.1016/j.mcpro.2022.100266) <span class="__dimensions_badge_embed__" data-doi="10.1016/j.mcpro.2022.100266" data-hide-zero-citations="true" data-style="small_rectangle"></span>
|
|
310
|
+
|
|
311
|
+
**MS²Rescore for timsTOF DDA-PASEF data:**
|
|
312
|
+
|
|
313
|
+
> **TIMS²Rescore: A DDA-PASEF optimized data-driven rescoring pipeline based on MS²Rescore.**
|
|
314
|
+
> Arthur Declercq*, Robbe Devreese*, Jonas Scheid, Caroline Jachmann, Tim Van Den Bossche, Annica Preikschat, David Gomez-Zepeda, Jeewan Babu Rijal, Aurélie Hirschler, Jonathan R Krieger, Tharan Srikumar, George Rosenberger, Dennis Trede, Christine Carapito, Stefan Tenzer, Juliane S Walz, Sven Degroeve, Robbin Bouwmeester, Lennart Martens, and Ralf Gabriels.
|
|
315
|
+
> _Journal of Proteome Research_ (2025) [doi:10.1021/acs.jproteome.4c00609](https://doi.org/10.1021/acs.jproteome.4c00609) <span class="__dimensions_badge_embed__" data-doi="10.1021/acs.jproteome.4c00609" data-hide-zero-citations="true" data-style="small_rectangle"></span>
|
|
316
|
+
|
|
317
|
+
**Original publication describing the concept of rescoring with predicted spectra:**
|
|
318
|
+
|
|
319
|
+
> **Accurate peptide fragmentation predictions allow data driven approaches to replace and improve upon proteomics search engine scoring functions.**
|
|
320
|
+
> Ana S C Silva, Robbin Bouwmeester, Lennart Martens, and Sven Degroeve.
|
|
321
|
+
> _Bioinformatics_ (2019) [doi:10.1093/bioinformatics/btz383](https://doi.org/10.1093/bioinformatics/btz383) <span class="__dimensions_badge_embed__" data-doi="10.1093/bioinformatics/btz383" data-hide-zero-citations="true" data-style="small_rectangle"></span>
|
|
322
|
+
|
|
323
|
+
To replicate the experiments described in this article, check out the
|
|
324
|
+
[publication branch][publication-branch] of the repository.
|
|
325
|
+
|
|
326
|
+
## Getting started
|
|
327
|
+
|
|
328
|
+
The desktop application can be installed on Windows with a [one-click installer][desktop-installer].
|
|
329
|
+
The Python package and command line interface can be installed with `pip`, `conda`, or `docker`.
|
|
330
|
+
Check out the [full documentation][docs] to get started.
|
|
331
|
+
|
|
332
|
+
## Questions or issues?
|
|
333
|
+
|
|
334
|
+
Have questions on how to apply MS²Rescore on your data? Or ran into issues while using MS²Rescore?
|
|
335
|
+
Post your questions on the [GitHub Discussions][discussions] forum and we are happy to help!
|
|
336
|
+
|
|
337
|
+
## How to contribute
|
|
338
|
+
|
|
339
|
+
Bugs, questions or suggestions? Feel free to post an issue in the [issue tracker][issues] or to
|
|
340
|
+
make a [pull request][pr]!
|
|
341
|
+
|
|
342
|
+
[docs]: https://ms2rescore.readthedocs.io/
|
|
343
|
+
[issues]: https://github.com/compomics/ms2rescore/issues/
|
|
344
|
+
[discussions]: https://github.com/compomics/ms2rescore/discussions/
|
|
345
|
+
[pr]: https://github.com/compomics/ms2rescore/pulls/
|
|
346
|
+
[desktop]: https://ms2rescore.readthedocs.io/en/stable/gui/
|
|
347
|
+
[desktop-installer]: https://github.com/compomics/ms2rescore/releases/latest
|
|
348
|
+
[cli]: https://ms2rescore.readthedocs.io/en/stable/cli/
|
|
349
|
+
[python-package]: https://ms2rescore.readthedocs.io/en/stable/api/ms2rescore/
|
|
350
|
+
[docker]: https://ms2rescore.readthedocs.io/en/stable/installation#docker-container
|
|
351
|
+
[publication-branch]: https://github.com/compomics/ms2rescore/tree/pub
|
|
352
|
+
[ms2pip]: https://github.com/compomics/ms2pip
|
|
353
|
+
[deeplc]: https://github.com/compomics/deeplc
|
|
354
|
+
[percolator]: https://github.com/percolator/percolator/
|
|
355
|
+
[mokapot]: https://mokapot.readthedocs.io/
|
|
356
|
+
[psm_utils]: https://github.com/compomics/psm_utils
|
|
357
|
+
[file-formats]: https://psm-utils.readthedocs.io/en/stable/#supported-file-formats
|
|
358
|
+
[tims2rescore]: https://ms2rescore.readthedocs.io/en/stable/userguide/tims2Rescore
|
|
@@ -0,0 +1,34 @@
|
|
|
1
|
+
"""Modular and user-friendly platform for AI-assisted rescoring of peptide identifications ."""
|
|
2
|
+
|
|
3
|
+
# __version__ is provided by the small helper module _version which prefers
|
|
4
|
+
# installed distribution metadata and falls back to reading pyproject.toml.
|
|
5
|
+
# The single source of truth for the version is pyproject.toml.
|
|
6
|
+
|
|
7
|
+
__all__ = [
|
|
8
|
+
"parse_configurations",
|
|
9
|
+
"rescore",
|
|
10
|
+
]
|
|
11
|
+
|
|
12
|
+
from warnings import filterwarnings
|
|
13
|
+
|
|
14
|
+
# mzmlb is not used, so hdf5plugin is not needed
|
|
15
|
+
filterwarnings(
|
|
16
|
+
"ignore",
|
|
17
|
+
message="hdf5plugin is missing",
|
|
18
|
+
category=UserWarning,
|
|
19
|
+
module="psims.mzmlb",
|
|
20
|
+
)
|
|
21
|
+
|
|
22
|
+
# Attempt to suppress pyopenms warning about OPENMS_DATA_PATH already set
|
|
23
|
+
filterwarnings(
|
|
24
|
+
"ignore",
|
|
25
|
+
message="Warning: OPENMS_DATA_PATH environment variable already exists.*",
|
|
26
|
+
category=UserWarning,
|
|
27
|
+
module="pyopenms",
|
|
28
|
+
)
|
|
29
|
+
|
|
30
|
+
from ms2rescore._version import get_version # noqa: E402
|
|
31
|
+
from ms2rescore.config_parser import parse_configurations # noqa: E402
|
|
32
|
+
from ms2rescore.core import rescore # noqa: E402
|
|
33
|
+
|
|
34
|
+
__version__ = get_version()
|
|
@@ -14,6 +14,7 @@ from rich.logging import RichHandler
|
|
|
14
14
|
from rich.text import Text
|
|
15
15
|
|
|
16
16
|
from ms2rescore import __version__, package_data
|
|
17
|
+
from ms2rescore._version import check_for_update
|
|
17
18
|
from ms2rescore.config_parser import parse_configurations
|
|
18
19
|
from ms2rescore.core import rescore
|
|
19
20
|
from ms2rescore.exceptions import MS2RescoreConfigurationError
|
|
@@ -148,18 +149,24 @@ def _argument_parser() -> argparse.ArgumentParser:
|
|
|
148
149
|
)
|
|
149
150
|
parser.add_argument(
|
|
150
151
|
"--write-report",
|
|
151
|
-
# metavar="BOOL",
|
|
152
152
|
action="store_true",
|
|
153
|
+
default=None,
|
|
153
154
|
dest="write_report",
|
|
154
|
-
help="boolean to
|
|
155
|
+
help="boolean whether to write an HTML report (default: True)",
|
|
156
|
+
)
|
|
157
|
+
parser.add_argument(
|
|
158
|
+
"--disable-update-check",
|
|
159
|
+
action="store_true",
|
|
160
|
+
default=None,
|
|
161
|
+
dest="disable_update_check",
|
|
162
|
+
help="Disable automatic check for software updates (default: False)",
|
|
155
163
|
)
|
|
156
164
|
parser.add_argument(
|
|
157
165
|
"--profile",
|
|
158
|
-
# metavar="BOOL",
|
|
159
166
|
action="store_true",
|
|
160
|
-
|
|
161
|
-
|
|
162
|
-
help="
|
|
167
|
+
default=None,
|
|
168
|
+
dest="profile",
|
|
169
|
+
help="Enable profiling with cProfile",
|
|
163
170
|
)
|
|
164
171
|
|
|
165
172
|
return parser
|
|
@@ -228,9 +235,19 @@ def main(tims=False):
|
|
|
228
235
|
config["ms2rescore"]["log_level"], config["ms2rescore"]["output_path"] + ".log.txt"
|
|
229
236
|
)
|
|
230
237
|
|
|
238
|
+
# Check for updates
|
|
239
|
+
if config["ms2rescore"]["disable_update_check"] is False:
|
|
240
|
+
update_info = check_for_update()
|
|
241
|
+
if update_info["update_available"]:
|
|
242
|
+
LOGGER.info(
|
|
243
|
+
f"New version of MS²Rescore available: {update_info['latest_version']} "
|
|
244
|
+
f"(you are using {update_info['current_version']})"
|
|
245
|
+
)
|
|
246
|
+
LOGGER.info(f"Download the latest version at: {update_info['html_url']}")
|
|
247
|
+
|
|
231
248
|
# Run MS²Rescore
|
|
232
249
|
try:
|
|
233
|
-
if
|
|
250
|
+
if config["ms2rescore"]["profile"]:
|
|
234
251
|
profiled_rescore = profile(rescore, config["ms2rescore"]["output_path"])
|
|
235
252
|
profiled_rescore(configuration=config)
|
|
236
253
|
else:
|
|
@@ -0,0 +1,135 @@
|
|
|
1
|
+
"""
|
|
2
|
+
Resolve the package version.
|
|
3
|
+
|
|
4
|
+
Prefer installed distribution metadata (importlib.metadata); if the package is not installed, fall
|
|
5
|
+
back to reading ``pyproject.toml`` from the repository root (PEP 621: ``[project].version``).
|
|
6
|
+
"""
|
|
7
|
+
|
|
8
|
+
import importlib.metadata
|
|
9
|
+
import json
|
|
10
|
+
import logging
|
|
11
|
+
from pathlib import Path
|
|
12
|
+
from typing import Dict, Optional, Tuple, Union
|
|
13
|
+
from urllib.error import HTTPError, URLError
|
|
14
|
+
from urllib.request import Request, urlopen
|
|
15
|
+
|
|
16
|
+
from packaging.version import Version
|
|
17
|
+
|
|
18
|
+
try:
|
|
19
|
+
import tomllib as toml # type: ignore
|
|
20
|
+
except ImportError:
|
|
21
|
+
import tomli as toml # type: ignore
|
|
22
|
+
|
|
23
|
+
from ms2rescore.exceptions import MS2RescoreError
|
|
24
|
+
|
|
25
|
+
LOGGER = logging.getLogger(__name__)
|
|
26
|
+
|
|
27
|
+
_GITHUB_REPO = "CompOmics/ms2rescore"
|
|
28
|
+
_GITHUB_TIMEOUT_SECONDS = 2.5
|
|
29
|
+
|
|
30
|
+
|
|
31
|
+
class UpdateCheckError(MS2RescoreError):
|
|
32
|
+
"""An error occurred while checking for software updates."""
|
|
33
|
+
|
|
34
|
+
pass
|
|
35
|
+
|
|
36
|
+
|
|
37
|
+
def _version_from_metadata() -> Optional[Version]:
|
|
38
|
+
try:
|
|
39
|
+
return Version(importlib.metadata.version("ms2rescore"))
|
|
40
|
+
except importlib.metadata.PackageNotFoundError:
|
|
41
|
+
return None
|
|
42
|
+
|
|
43
|
+
|
|
44
|
+
def _version_from_pyproject() -> Optional[Version]:
|
|
45
|
+
pyproject = Path(__file__).resolve().parents[1] / "pyproject.toml"
|
|
46
|
+
if not pyproject.is_file():
|
|
47
|
+
return None
|
|
48
|
+
|
|
49
|
+
try:
|
|
50
|
+
data = toml.loads(pyproject.read_text(encoding="utf-8"))
|
|
51
|
+
except (OSError, ValueError):
|
|
52
|
+
return None
|
|
53
|
+
|
|
54
|
+
project = data.get("project") if isinstance(data, dict) else None
|
|
55
|
+
if isinstance(project, dict):
|
|
56
|
+
ver = project.get("version")
|
|
57
|
+
if isinstance(ver, str):
|
|
58
|
+
return Version(ver)
|
|
59
|
+
return None
|
|
60
|
+
|
|
61
|
+
|
|
62
|
+
def _get_latest_version(timeout_seconds: float) -> Tuple[Version, Optional[str]]:
|
|
63
|
+
"""Check GitHub latest release and return the version string."""
|
|
64
|
+
# Prepare GitHub API request
|
|
65
|
+
url = f"https://api.github.com/repos/{_GITHUB_REPO}/releases/latest"
|
|
66
|
+
user_agent = "ms2rescore/ (+https://github.com/compomics/ms2rescore)"
|
|
67
|
+
req = Request(url, headers={"Accept": "application/vnd.github+json", "User-Agent": user_agent})
|
|
68
|
+
|
|
69
|
+
# Fetch and parse release info
|
|
70
|
+
try:
|
|
71
|
+
with urlopen(req, timeout=timeout_seconds) as resp:
|
|
72
|
+
raw = resp.read()
|
|
73
|
+
data = json.loads(raw.decode("utf-8", errors="replace")) if raw else {}
|
|
74
|
+
except HTTPError as e:
|
|
75
|
+
raise UpdateCheckError(f"HTTP {e.code}") from e
|
|
76
|
+
except URLError as e:
|
|
77
|
+
raise UpdateCheckError("Network unavailable or timed out") from e
|
|
78
|
+
except Exception as e:
|
|
79
|
+
raise UpdateCheckError("Failed to fetch release info") from e
|
|
80
|
+
|
|
81
|
+
tag = data.get("tag_name") or data.get("name") or ""
|
|
82
|
+
if not tag:
|
|
83
|
+
raise UpdateCheckError("Latest release tag not found in API response")
|
|
84
|
+
|
|
85
|
+
# Validate version string
|
|
86
|
+
try:
|
|
87
|
+
latest_version = Version(tag)
|
|
88
|
+
except Exception as e:
|
|
89
|
+
raise UpdateCheckError("Latest release tag is not a valid version") from e
|
|
90
|
+
|
|
91
|
+
return latest_version, data.get("html_url") or data.get("url")
|
|
92
|
+
|
|
93
|
+
|
|
94
|
+
def get_version() -> str:
|
|
95
|
+
"""Return the version of this MS²Rescore installation."""
|
|
96
|
+
version = _version_from_metadata() or _version_from_pyproject() or "0+unknown"
|
|
97
|
+
return str(version)
|
|
98
|
+
|
|
99
|
+
|
|
100
|
+
def check_for_update(
|
|
101
|
+
timeout_seconds: Optional[float] = None,
|
|
102
|
+
) -> Dict[str, Optional[Union[str, bool]]]:
|
|
103
|
+
"""Check GitHub latest release and report whether an update exists."""
|
|
104
|
+
timeout_seconds = timeout_seconds or _GITHUB_TIMEOUT_SECONDS
|
|
105
|
+
|
|
106
|
+
# Initialize result dictionary
|
|
107
|
+
result: Dict[str, Optional[Union[str, bool]]] = {
|
|
108
|
+
"update_available": False,
|
|
109
|
+
"current_version": None,
|
|
110
|
+
"latest_version": None,
|
|
111
|
+
"html_url": None,
|
|
112
|
+
}
|
|
113
|
+
|
|
114
|
+
# Get current version
|
|
115
|
+
current_version = _version_from_metadata() or _version_from_pyproject()
|
|
116
|
+
if current_version is None:
|
|
117
|
+
LOGGER.warning("Update check failed: Cannot determine current version")
|
|
118
|
+
return result
|
|
119
|
+
result["current_version"] = str(current_version)
|
|
120
|
+
|
|
121
|
+
# Get latest version from GitHub
|
|
122
|
+
try:
|
|
123
|
+
latest_version, html_url = _get_latest_version(timeout_seconds)
|
|
124
|
+
except UpdateCheckError as e:
|
|
125
|
+
LOGGER.warning("Update check failed: %s", e)
|
|
126
|
+
return result
|
|
127
|
+
result["latest_version"] = str(latest_version)
|
|
128
|
+
result["html_url"] = html_url
|
|
129
|
+
|
|
130
|
+
try:
|
|
131
|
+
result["update_available"] = latest_version > current_version
|
|
132
|
+
except Exception:
|
|
133
|
+
# If current_version can't be parsed, don't treat as updateable
|
|
134
|
+
result["update_available"] = False
|
|
135
|
+
return result
|
|
@@ -3,6 +3,7 @@
|
|
|
3
3
|
import importlib.resources
|
|
4
4
|
import json
|
|
5
5
|
import multiprocessing as mp
|
|
6
|
+
import re
|
|
6
7
|
from argparse import Namespace
|
|
7
8
|
from pathlib import Path
|
|
8
9
|
from typing import Dict, List, Union
|
|
@@ -10,7 +11,7 @@ from typing import Dict, List, Union
|
|
|
10
11
|
try:
|
|
11
12
|
import tomllib
|
|
12
13
|
except ImportError:
|
|
13
|
-
import tomli as tomllib
|
|
14
|
+
import tomli as tomllib # type: ignore
|
|
14
15
|
|
|
15
16
|
from cascade_config import CascadeConfig
|
|
16
17
|
|
|
@@ -86,6 +87,36 @@ def _validate_processes(config: Dict) -> Dict:
|
|
|
86
87
|
return config
|
|
87
88
|
|
|
88
89
|
|
|
90
|
+
def _validate_regular_expressions(config: Dict) -> Dict:
|
|
91
|
+
"""Validate regular expressions in configuration."""
|
|
92
|
+
for field in [
|
|
93
|
+
"psm_id_pattern",
|
|
94
|
+
"spectrum_id_pattern",
|
|
95
|
+
"psm_id_rt_pattern",
|
|
96
|
+
"psm_id_im_pattern",
|
|
97
|
+
]:
|
|
98
|
+
if config["ms2rescore"][field]:
|
|
99
|
+
|
|
100
|
+
# Check if valid regex
|
|
101
|
+
try:
|
|
102
|
+
pattern = re.compile(config["ms2rescore"][field])
|
|
103
|
+
except re.error as e:
|
|
104
|
+
raise MS2RescoreConfigurationError(
|
|
105
|
+
f"Invalid regular expression provided for '{field}': {e}"
|
|
106
|
+
) from e
|
|
107
|
+
|
|
108
|
+
# Check if regex has exactly one capturing group
|
|
109
|
+
if pattern.groups != 1:
|
|
110
|
+
raise MS2RescoreConfigurationError(
|
|
111
|
+
f"Regular expression for '{field}' should contain exactly one "
|
|
112
|
+
"capturing group. Please check and try again. "
|
|
113
|
+
"See https://ms2rescore.readthedocs.io/en/stable/userguide/configuration/#mapping-psms-to-spectra "
|
|
114
|
+
"for more information."
|
|
115
|
+
)
|
|
116
|
+
|
|
117
|
+
return config
|
|
118
|
+
|
|
119
|
+
|
|
89
120
|
def parse_configurations(configurations: List[Union[dict, str, Path, Namespace]]) -> Dict:
|
|
90
121
|
"""
|
|
91
122
|
Parse and validate MS²Rescore configuration files and CLI arguments.
|
|
@@ -142,6 +173,7 @@ def parse_configurations(configurations: List[Union[dict, str, Path, Namespace]]
|
|
|
142
173
|
# Validate and infer filenames and number of parallel processes
|
|
143
174
|
config = _validate_filenames(config)
|
|
144
175
|
config = _validate_processes(config)
|
|
176
|
+
config = _validate_regular_expressions(config)
|
|
145
177
|
|
|
146
178
|
# Convert feature_generators and rescoring_engine names to lowercase
|
|
147
179
|
config["ms2rescore"]["feature_generators"] = {
|
|
@@ -61,8 +61,16 @@ def rescore(configuration: Dict, psm_list: Optional[PSMList] = None) -> None:
|
|
|
61
61
|
)
|
|
62
62
|
|
|
63
63
|
# Add missing precursor info from spectrum file if needed
|
|
64
|
+
required_ms_data = {
|
|
65
|
+
ms_data
|
|
66
|
+
for fgen_name in config["feature_generators"].keys()
|
|
67
|
+
for ms_data in FEATURE_GENERATORS[fgen_name].required_ms_data
|
|
68
|
+
}
|
|
64
69
|
available_ms_data = add_precursor_values(
|
|
65
|
-
psm_list,
|
|
70
|
+
psm_list,
|
|
71
|
+
required_ms_data,
|
|
72
|
+
spectrum_path=config["spectrum_path"],
|
|
73
|
+
spectrum_id_pattern=config["spectrum_id_pattern"],
|
|
66
74
|
)
|
|
67
75
|
|
|
68
76
|
# Add rescoring features
|