imml 0.1.0__tar.gz
This diff represents the content of publicly available package versions that have been released to one of the supported registries. The information contained in this diff is provided for informational purposes only and reflects changes between package versions as they appear in their respective public registries.
- imml-0.1.0/LICENSE +28 -0
- imml-0.1.0/PKG-INFO +237 -0
- imml-0.1.0/README.md +150 -0
- imml-0.1.0/imml/__init__.py +1 -0
- imml-0.1.0/imml/ampute/__init__.py +2 -0
- imml-0.1.0/imml/ampute/amputer.py +202 -0
- imml-0.1.0/imml/ampute/remove_mods.py +77 -0
- imml-0.1.0/imml/classify/__init__.py +3 -0
- imml-0.1.0/imml/classify/_m3care/__init__.py +4 -0
- imml-0.1.0/imml/classify/_m3care/graph_convolution.py +39 -0
- imml-0.1.0/imml/classify/_m3care/mm_transformer_encoder.py +82 -0
- imml-0.1.0/imml/classify/_m3care/nmt.py +367 -0
- imml-0.1.0/imml/classify/_m3care/utils.py +57 -0
- imml-0.1.0/imml/classify/_muse/__init__.py +5 -0
- imml-0.1.0/imml/classify/_muse/code_encoder.py +212 -0
- imml-0.1.0/imml/classify/_muse/ffn_encoder.py +63 -0
- imml-0.1.0/imml/classify/_muse/gnn.py +236 -0
- imml-0.1.0/imml/classify/_muse/rnn_encoder.py +106 -0
- imml-0.1.0/imml/classify/_muse/text_encoder.py +25 -0
- imml-0.1.0/imml/classify/_ragpt/__init__.py +2 -0
- imml-0.1.0/imml/classify/_ragpt/core_tools.py +5 -0
- imml-0.1.0/imml/classify/_ragpt/modules.py +60 -0
- imml-0.1.0/imml/classify/_ragpt/vilt/__init__.py +3 -0
- imml-0.1.0/imml/classify/_ragpt/vilt/configuration_vilt.py +149 -0
- imml-0.1.0/imml/classify/_ragpt/vilt/image_processing_vilt.py +518 -0
- imml-0.1.0/imml/classify/_ragpt/vilt/modeling_vilt.py +1515 -0
- imml-0.1.0/imml/classify/m3care.py +342 -0
- imml-0.1.0/imml/classify/map.py +396 -0
- imml-0.1.0/imml/classify/muse.py +270 -0
- imml-0.1.0/imml/classify/ragpt.py +315 -0
- imml-0.1.0/imml/cluster/__init__.py +16 -0
- imml-0.1.0/imml/cluster/_daimc/DAIMC.m +52 -0
- imml-0.1.0/imml/cluster/_daimc/UpdateV_DAIMC.m +43 -0
- imml-0.1.0/imml/cluster/_daimc/__init__.py +0 -0
- imml-0.1.0/imml/cluster/_daimc/litekmeans.m +437 -0
- imml-0.1.0/imml/cluster/_daimc/newinit.m +43 -0
- imml-0.1.0/imml/cluster/_eeimvc/__init__.py +0 -0
- imml-0.1.0/imml/cluster/_eeimvc/algorithm2.m +12 -0
- imml-0.1.0/imml/cluster/_eeimvc/incompleteLateFusionMKCOrthHp_lambda.m +54 -0
- imml-0.1.0/imml/cluster/_eeimvc/myInitializationHp.m +17 -0
- imml-0.1.0/imml/cluster/_eeimvc/mycombFun.m +8 -0
- imml-0.1.0/imml/cluster/_eeimvc/mykernelkmeans.m +8 -0
- imml-0.1.0/imml/cluster/_eeimvc/updateBetaAbsentClustering.m +34 -0
- imml-0.1.0/imml/cluster/_eeimvc/updateHPabsentClusteringOrthHp.m +18 -0
- imml-0.1.0/imml/cluster/_eeimvc/updateWPabsentClusteringV1.m +10 -0
- imml-0.1.0/imml/cluster/_imscagl/EProjSimplex_new.m +40 -0
- imml-0.1.0/imml/cluster/_imscagl/EuDist2.m +59 -0
- imml-0.1.0/imml/cluster/_imscagl/IMSAGL.m +157 -0
- imml-0.1.0/imml/cluster/_imscagl/IMSC.m +36 -0
- imml-0.1.0/imml/cluster/_imscagl/L2_distance_1.m +23 -0
- imml-0.1.0/imml/cluster/_imscagl/__init__.py +0 -0
- imml-0.1.0/imml/cluster/_imscagl/constructW.m +526 -0
- imml-0.1.0/imml/cluster/_imscagl/solveF.m +30 -0
- imml-0.1.0/imml/cluster/_imsr/IMSC.m +36 -0
- imml-0.1.0/imml/cluster/_imsr/__init__.py +0 -0
- imml-0.1.0/imml/cluster/_imsr/baseline_spectral_onkernel.m +12 -0
- imml-0.1.0/imml/cluster/_imsr/cal_obj.m +18 -0
- imml-0.1.0/imml/cluster/_imsr/init_Z.m +19 -0
- imml-0.1.0/imml/cluster/_imsr/update_F.m +13 -0
- imml-0.1.0/imml/cluster/_imsr/update_X.m +13 -0
- imml-0.1.0/imml/cluster/_imsr/update_Z.m +48 -0
- imml-0.1.0/imml/cluster/_integrao/__init__.py +0 -0
- imml-0.1.0/imml/cluster/_integrao/_aux_integrao.py +206 -0
- imml-0.1.0/imml/cluster/_lfimvc/IncompleteMultikernelLatefusionclusteringV1Hv.m +56 -0
- imml-0.1.0/imml/cluster/_lfimvc/__init__.py +0 -0
- imml-0.1.0/imml/cluster/_lfimvc/mykernelkmeans.m +8 -0
- imml-0.1.0/imml/cluster/_lfimvc/updateHPabsentClusteringV1.m +11 -0
- imml-0.1.0/imml/cluster/_lfimvc/updateWPabsentClusteringV1.m +10 -0
- imml-0.1.0/imml/cluster/_mkkmik/DataCompletion.m +18 -0
- imml-0.1.0/imml/cluster/_mkkmik/__init__.py +0 -0
- imml-0.1.0/imml/cluster/_mkkmik/absentKernelImputation.m +26 -0
- imml-0.1.0/imml/cluster/_mkkmik/algorithm0.m +21 -0
- imml-0.1.0/imml/cluster/_mkkmik/algorithm2.m +11 -0
- imml-0.1.0/imml/cluster/_mkkmik/algorithm3.m +15 -0
- imml-0.1.0/imml/cluster/_mkkmik/algorithm4.m +20 -0
- imml-0.1.0/imml/cluster/_mkkmik/algorithm6.m +11 -0
- imml-0.1.0/imml/cluster/_mkkmik/calObjV2.m +25 -0
- imml-0.1.0/imml/cluster/_mkkmik/kcenter.m +39 -0
- imml-0.1.0/imml/cluster/_mkkmik/knorm.m +29 -0
- imml-0.1.0/imml/cluster/_mkkmik/myabsentmultikernelclustering.m +49 -0
- imml-0.1.0/imml/cluster/_mkkmik/mycombFun.m +8 -0
- imml-0.1.0/imml/cluster/_mkkmik/mykernelkmeans.m +8 -0
- imml-0.1.0/imml/cluster/_mkkmik/updateabsentkernelweightsV2.m +20 -0
- imml-0.1.0/imml/cluster/_monet/__init__.py +0 -0
- imml-0.1.0/imml/cluster/_monet/_aux_monet.py +684 -0
- imml-0.1.0/imml/cluster/_nemo/NEMO.R +134 -0
- imml-0.1.0/imml/cluster/_nemo/__init__.py +0 -0
- imml-0.1.0/imml/cluster/_omvc/ONMF_Multi_PGD_search.m +209 -0
- imml-0.1.0/imml/cluster/_omvc/__init__.py +0 -0
- imml-0.1.0/imml/cluster/_omvc/objective_ONMF_Multi.m +9 -0
- imml-0.1.0/imml/cluster/_opimc/NormalizeFea.m +55 -0
- imml-0.1.0/imml/cluster/_opimc/OPIMC.m +149 -0
- imml-0.1.0/imml/cluster/_opimc/UpdateV.m +13 -0
- imml-0.1.0/imml/cluster/_opimc/__init__.py +0 -0
- imml-0.1.0/imml/cluster/_oslfimvc/OS_LF_IMVC_alg.m +67 -0
- imml-0.1.0/imml/cluster/_oslfimvc/__init__.py +0 -0
- imml-0.1.0/imml/cluster/_oslfimvc/initializeKH.m +13 -0
- imml-0.1.0/imml/cluster/_oslfimvc/kcenter.m +39 -0
- imml-0.1.0/imml/cluster/_oslfimvc/knorm.m +30 -0
- imml-0.1.0/imml/cluster/_oslfimvc/myInitialization.m +16 -0
- imml-0.1.0/imml/cluster/_oslfimvc/myInitializationC.m +9 -0
- imml-0.1.0/imml/cluster/_oslfimvc/mySolving.m +10 -0
- imml-0.1.0/imml/cluster/_oslfimvc/mycombFun.m +8 -0
- imml-0.1.0/imml/cluster/_oslfimvc/mykernelkmeans.m +7 -0
- imml-0.1.0/imml/cluster/_oslfimvc/updateBeta_OSLFIMVC.m +10 -0
- imml-0.1.0/imml/cluster/_oslfimvc/updateWP_OSLFIMVC.m +12 -0
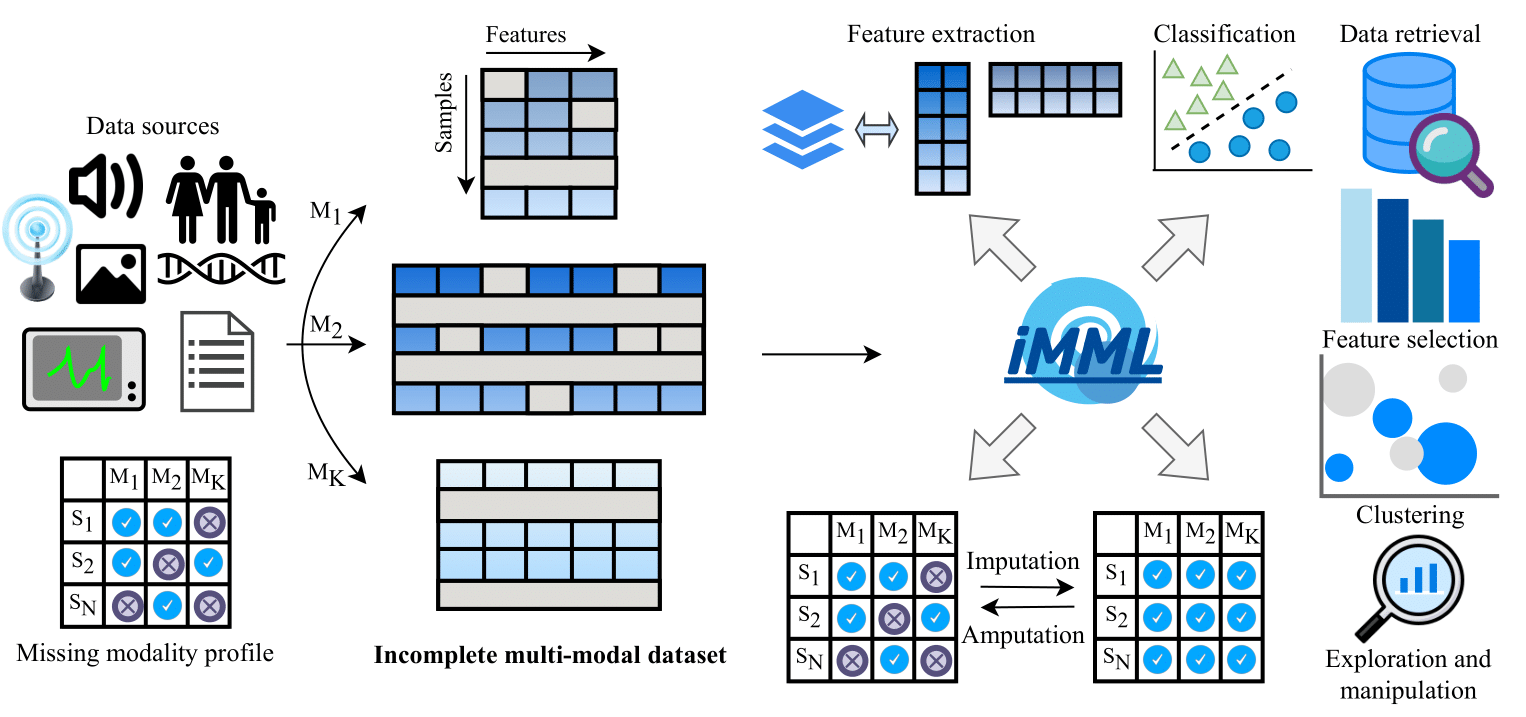
- imml-0.1.0/imml/cluster/_pimvc/EuDist2.m +59 -0
- imml-0.1.0/imml/cluster/_pimvc/PCA1.m +75 -0
- imml-0.1.0/imml/cluster/_pimvc/PIMVC.m +64 -0
- imml-0.1.0/imml/cluster/_pimvc/__init__.py +0 -0
- imml-0.1.0/imml/cluster/_pimvc/constructW.m +526 -0
- imml-0.1.0/imml/cluster/_pimvc/mySVD.m +117 -0
- imml-0.1.0/imml/cluster/_simcadc/EProjSimplex_new.m +40 -0
- imml-0.1.0/imml/cluster/_simcadc/SIMC.m +118 -0
- imml-0.1.0/imml/cluster/_simcadc/__init__.py +0 -0
- imml-0.1.0/imml/cluster/_sumo/__init__.py +0 -0
- imml-0.1.0/imml/cluster/_sumo/constants.py +70 -0
- imml-0.1.0/imml/cluster/_sumo/modes/prepare/similarity.py +160 -0
- imml-0.1.0/imml/cluster/_sumo/modes/run/solver.py +130 -0
- imml-0.1.0/imml/cluster/_sumo/modes/run/solvers/unsupervised_sumo.py +148 -0
- imml-0.1.0/imml/cluster/_sumo/modes/run/utils.py +99 -0
- imml-0.1.0/imml/cluster/_sumo/network.py +72 -0
- imml-0.1.0/imml/cluster/_sumo/utils.py +299 -0
- imml-0.1.0/imml/cluster/daimc.py +442 -0
- imml-0.1.0/imml/cluster/eeimvc.py +524 -0
- imml-0.1.0/imml/cluster/imscagl.py +223 -0
- imml-0.1.0/imml/cluster/imsr.py +454 -0
- imml-0.1.0/imml/cluster/integrao.py +424 -0
- imml-0.1.0/imml/cluster/lfimvc.py +386 -0
- imml-0.1.0/imml/cluster/mkkmik.py +223 -0
- imml-0.1.0/imml/cluster/monet.py +575 -0
- imml-0.1.0/imml/cluster/mrgcn.py +261 -0
- imml-0.1.0/imml/cluster/nemo.py +243 -0
- imml-0.1.0/imml/cluster/omvc.py +225 -0
- imml-0.1.0/imml/cluster/opimc.py +203 -0
- imml-0.1.0/imml/cluster/oslfimvc.py +213 -0
- imml-0.1.0/imml/cluster/pimvc.py +227 -0
- imml-0.1.0/imml/cluster/simcadc.py +429 -0
- imml-0.1.0/imml/cluster/sumo.py +449 -0
- imml-0.1.0/imml/decomposition/__init__.py +3 -0
- imml-0.1.0/imml/decomposition/_mofa/__init__.py +0 -0
- imml-0.1.0/imml/decomposition/_mofa/_mofax/__init__.py +0 -0
- imml-0.1.0/imml/decomposition/_mofa/_mofax/core.py +1479 -0
- imml-0.1.0/imml/decomposition/_mofa/_mofax/utils.py +259 -0
- imml-0.1.0/imml/decomposition/_mofa/build_model/__init__.py +0 -0
- imml-0.1.0/imml/decomposition/_mofa/build_model/build_model.py +380 -0
- imml-0.1.0/imml/decomposition/_mofa/build_model/init_model.py +920 -0
- imml-0.1.0/imml/decomposition/_mofa/build_model/save_model.py +757 -0
- imml-0.1.0/imml/decomposition/_mofa/build_model/train_model.py +32 -0
- imml-0.1.0/imml/decomposition/_mofa/build_model/utils.py +115 -0
- imml-0.1.0/imml/decomposition/_mofa/config.py +25 -0
- imml-0.1.0/imml/decomposition/_mofa/core/_BayesNet.py +1150 -0
- imml-0.1.0/imml/decomposition/_mofa/core/__init__.py +1 -0
- imml-0.1.0/imml/decomposition/_mofa/core/distributions/__init__.py +11 -0
- imml-0.1.0/imml/decomposition/_mofa/core/distributions/basic_distributions.py +112 -0
- imml-0.1.0/imml/decomposition/_mofa/core/distributions/bernoulli.py +55 -0
- imml-0.1.0/imml/decomposition/_mofa/core/distributions/bernoulli_gaussian.py +134 -0
- imml-0.1.0/imml/decomposition/_mofa/core/distributions/beta.py +57 -0
- imml-0.1.0/imml/decomposition/_mofa/core/distributions/binomial.py +64 -0
- imml-0.1.0/imml/decomposition/_mofa/core/distributions/gamma.py +72 -0
- imml-0.1.0/imml/decomposition/_mofa/core/distributions/multi_task_GP.py +783 -0
- imml-0.1.0/imml/decomposition/_mofa/core/distributions/multivariate_gaussian.py +407 -0
- imml-0.1.0/imml/decomposition/_mofa/core/distributions/poisson.py +54 -0
- imml-0.1.0/imml/decomposition/_mofa/core/distributions/univariate_gaussian.py +76 -0
- imml-0.1.0/imml/decomposition/_mofa/core/gp_utils.py +160 -0
- imml-0.1.0/imml/decomposition/_mofa/core/gpu_utils.py +87 -0
- imml-0.1.0/imml/decomposition/_mofa/core/nodes/Alpha_nodes.py +203 -0
- imml-0.1.0/imml/decomposition/_mofa/core/nodes/Kc_node.py +122 -0
- imml-0.1.0/imml/decomposition/_mofa/core/nodes/Kg_node.py +102 -0
- imml-0.1.0/imml/decomposition/_mofa/core/nodes/Sigma_node.py +1317 -0
- imml-0.1.0/imml/decomposition/_mofa/core/nodes/Tau_nodes.py +167 -0
- imml-0.1.0/imml/decomposition/_mofa/core/nodes/Theta_nodes.py +199 -0
- imml-0.1.0/imml/decomposition/_mofa/core/nodes/U_nodes.py +229 -0
- imml-0.1.0/imml/decomposition/_mofa/core/nodes/W_nodes.py +351 -0
- imml-0.1.0/imml/decomposition/_mofa/core/nodes/Y_nodes.py +115 -0
- imml-0.1.0/imml/decomposition/_mofa/core/nodes/Z_nodes.py +468 -0
- imml-0.1.0/imml/decomposition/_mofa/core/nodes/Z_nodes_GP.py +260 -0
- imml-0.1.0/imml/decomposition/_mofa/core/nodes/Z_nodes_GP_mv.py +220 -0
- imml-0.1.0/imml/decomposition/_mofa/core/nodes/ZgU_node.py +203 -0
- imml-0.1.0/imml/decomposition/_mofa/core/nodes/__init__.py +17 -0
- imml-0.1.0/imml/decomposition/_mofa/core/nodes/basic_nodes.py +135 -0
- imml-0.1.0/imml/decomposition/_mofa/core/nodes/multiview_nodes.py +187 -0
- imml-0.1.0/imml/decomposition/_mofa/core/nodes/nongaussian_nodes.py +614 -0
- imml-0.1.0/imml/decomposition/_mofa/core/nodes/variational_nodes.py +328 -0
- imml-0.1.0/imml/decomposition/_mofa/core/utils.py +143 -0
- imml-0.1.0/imml/decomposition/_mofa/run/__init__.py +0 -0
- imml-0.1.0/imml/decomposition/_mofa/run/entry_point.py +1925 -0
- imml-0.1.0/imml/decomposition/_skfusion/__init__.py +0 -0
- imml-0.1.0/imml/decomposition/_skfusion/fusion/__init__.py +2 -0
- imml-0.1.0/imml/decomposition/_skfusion/fusion/base/__init__.py +2 -0
- imml-0.1.0/imml/decomposition/_skfusion/fusion/base/base.py +251 -0
- imml-0.1.0/imml/decomposition/_skfusion/fusion/base/fusion_graph.py +566 -0
- imml-0.1.0/imml/decomposition/_skfusion/fusion/decomposition/__init__.py +2 -0
- imml-0.1.0/imml/decomposition/_skfusion/fusion/decomposition/_dfmc.py +468 -0
- imml-0.1.0/imml/decomposition/_skfusion/fusion/decomposition/_dfmf.py +454 -0
- imml-0.1.0/imml/decomposition/_skfusion/fusion/decomposition/_init.py +61 -0
- imml-0.1.0/imml/decomposition/_skfusion/fusion/decomposition/dfmc.py +115 -0
- imml-0.1.0/imml/decomposition/_skfusion/fusion/decomposition/dfmf.py +204 -0
- imml-0.1.0/imml/decomposition/dfmf.py +170 -0
- imml-0.1.0/imml/decomposition/jnmf.py +330 -0
- imml-0.1.0/imml/decomposition/mofa.py +234 -0
- imml-0.1.0/imml/explore/__init__.py +5 -0
- imml-0.1.0/imml/explore/exploring_functions.py +477 -0
- imml-0.1.0/imml/feature_selection/__init__.py +1 -0
- imml-0.1.0/imml/feature_selection/jnmf_feature_selection.py +191 -0
- imml-0.1.0/imml/impute/__init__.py +6 -0
- imml-0.1.0/imml/impute/dfmf_imputer.py +72 -0
- imml-0.1.0/imml/impute/jnmf_imputer.py +121 -0
- imml-0.1.0/imml/impute/missing_mod_indicator.py +67 -0
- imml-0.1.0/imml/impute/mofa_imputer.py +77 -0
- imml-0.1.0/imml/impute/observed_mod_indicator.py +70 -0
- imml-0.1.0/imml/impute/simple_mod_imputer.py +167 -0
- imml-0.1.0/imml/load/__init__.py +5 -0
- imml-0.1.0/imml/load/integrao_dataset.py +113 -0
- imml-0.1.0/imml/load/m3care_dataset.py +86 -0
- imml-0.1.0/imml/load/map_dataset.py +315 -0
- imml-0.1.0/imml/load/mrgcn_dataset.py +62 -0
- imml-0.1.0/imml/load/muse_dataset.py +102 -0
- imml-0.1.0/imml/load/ragpt_dataset.py +238 -0
- imml-0.1.0/imml/preprocessing/__init__.py +9 -0
- imml-0.1.0/imml/preprocessing/compose.py +302 -0
- imml-0.1.0/imml/preprocessing/multi_mod_transformer.py +102 -0
- imml-0.1.0/imml/preprocessing/normalizer_nan.py +87 -0
- imml-0.1.0/imml/preprocessing/remove_incom_samples_by_mod.py +60 -0
- imml-0.1.0/imml/preprocessing/remove_missing_samples_by_mod.py +60 -0
- imml-0.1.0/imml/preprocessing/select_complete_samples.py +62 -0
- imml-0.1.0/imml/preprocessing/select_incomplete_samples.py +62 -0
- imml-0.1.0/imml/retrieve/__init__.py +1 -0
- imml-0.1.0/imml/retrieve/mcr.py +685 -0
- imml-0.1.0/imml/statistics/__init__.py +1 -0
- imml-0.1.0/imml/statistics/pid.py +236 -0
- imml-0.1.0/imml/utils/__init__.py +3 -0
- imml-0.1.0/imml/utils/check_xs.py +87 -0
- imml-0.1.0/imml/utils/convert_dataset_format.py +44 -0
- imml-0.1.0/imml/utils/utils.py +17 -0
- imml-0.1.0/imml/visualize/__init__.py +2 -0
- imml-0.1.0/imml/visualize/plot_missing_modality.py +60 -0
- imml-0.1.0/imml/visualize/plot_pid.py +126 -0
- imml-0.1.0/imml.egg-info/PKG-INFO +237 -0
- imml-0.1.0/imml.egg-info/SOURCES.txt +243 -0
- imml-0.1.0/imml.egg-info/dependency_links.txt +1 -0
- imml-0.1.0/imml.egg-info/requires.txt +34 -0
- imml-0.1.0/imml.egg-info/top_level.txt +1 -0
- imml-0.1.0/pyproject.toml +98 -0
- imml-0.1.0/setup.cfg +4 -0
imml-0.1.0/LICENSE
ADDED
|
@@ -0,0 +1,28 @@
|
|
|
1
|
+
BSD 3-Clause License
|
|
2
|
+
|
|
3
|
+
Copyright (c) 2025, Open source contributors.
|
|
4
|
+
|
|
5
|
+
Redistribution and use in source and binary forms, with or without
|
|
6
|
+
modification, are permitted provided that the following conditions are met:
|
|
7
|
+
|
|
8
|
+
* Redistributions of source code must retain the above copyright notice, this
|
|
9
|
+
list of conditions and the following disclaimer.
|
|
10
|
+
|
|
11
|
+
* Redistributions in binary form must reproduce the above copyright notice,
|
|
12
|
+
this list of conditions and the following disclaimer in the documentation
|
|
13
|
+
and/or other materials provided with the distribution.
|
|
14
|
+
|
|
15
|
+
* Neither the name of the copyright holder nor the names of its
|
|
16
|
+
contributors may be used to endorse or promote products derived from
|
|
17
|
+
this software without specific prior written permission.
|
|
18
|
+
|
|
19
|
+
THIS SOFTWARE IS PROVIDED BY THE COPYRIGHT HOLDERS AND CONTRIBUTORS "AS IS"
|
|
20
|
+
AND ANY EXPRESS OR IMPLIED WARRANTIES, INCLUDING, BUT NOT LIMITED TO, THE
|
|
21
|
+
IMPLIED WARRANTIES OF MERCHANTABILITY AND FITNESS FOR A PARTICULAR PURPOSE ARE
|
|
22
|
+
DISCLAIMED. IN NO EVENT SHALL THE COPYRIGHT HOLDER OR CONTRIBUTORS BE LIABLE
|
|
23
|
+
FOR ANY DIRECT, INDIRECT, INCIDENTAL, SPECIAL, EXEMPLARY, OR CONSEQUENTIAL
|
|
24
|
+
DAMAGES (INCLUDING, BUT NOT LIMITED TO, PROCUREMENT OF SUBSTITUTE GOODS OR
|
|
25
|
+
SERVICES; LOSS OF USE, DATA, OR PROFITS; OR BUSINESS INTERRUPTION) HOWEVER
|
|
26
|
+
CAUSED AND ON ANY THEORY OF LIABILITY, WHETHER IN CONTRACT, STRICT LIABILITY,
|
|
27
|
+
OR TORT (INCLUDING NEGLIGENCE OR OTHERWISE) ARISING IN ANY WAY OUT OF THE USE
|
|
28
|
+
OF THIS SOFTWARE, EVEN IF ADVISED OF THE POSSIBILITY OF SUCH DAMAGE.
|
imml-0.1.0/PKG-INFO
ADDED
|
@@ -0,0 +1,237 @@
|
|
|
1
|
+
Metadata-Version: 2.1
|
|
2
|
+
Name: imml
|
|
3
|
+
Version: 0.1.0
|
|
4
|
+
Summary: A python package for multi-modal learning with incomplete data
|
|
5
|
+
Author-email: Alberto López <a.l.sanchez@medisin.uio.no>
|
|
6
|
+
Maintainer-email: Alberto López <a.l.sanchez@medisin.uio.no>
|
|
7
|
+
License: BSD 3-Clause License
|
|
8
|
+
|
|
9
|
+
Copyright (c) 2025, Open source contributors.
|
|
10
|
+
|
|
11
|
+
Redistribution and use in source and binary forms, with or without
|
|
12
|
+
modification, are permitted provided that the following conditions are met:
|
|
13
|
+
|
|
14
|
+
* Redistributions of source code must retain the above copyright notice, this
|
|
15
|
+
list of conditions and the following disclaimer.
|
|
16
|
+
|
|
17
|
+
* Redistributions in binary form must reproduce the above copyright notice,
|
|
18
|
+
this list of conditions and the following disclaimer in the documentation
|
|
19
|
+
and/or other materials provided with the distribution.
|
|
20
|
+
|
|
21
|
+
* Neither the name of the copyright holder nor the names of its
|
|
22
|
+
contributors may be used to endorse or promote products derived from
|
|
23
|
+
this software without specific prior written permission.
|
|
24
|
+
|
|
25
|
+
THIS SOFTWARE IS PROVIDED BY THE COPYRIGHT HOLDERS AND CONTRIBUTORS "AS IS"
|
|
26
|
+
AND ANY EXPRESS OR IMPLIED WARRANTIES, INCLUDING, BUT NOT LIMITED TO, THE
|
|
27
|
+
IMPLIED WARRANTIES OF MERCHANTABILITY AND FITNESS FOR A PARTICULAR PURPOSE ARE
|
|
28
|
+
DISCLAIMED. IN NO EVENT SHALL THE COPYRIGHT HOLDER OR CONTRIBUTORS BE LIABLE
|
|
29
|
+
FOR ANY DIRECT, INDIRECT, INCIDENTAL, SPECIAL, EXEMPLARY, OR CONSEQUENTIAL
|
|
30
|
+
DAMAGES (INCLUDING, BUT NOT LIMITED TO, PROCUREMENT OF SUBSTITUTE GOODS OR
|
|
31
|
+
SERVICES; LOSS OF USE, DATA, OR PROFITS; OR BUSINESS INTERRUPTION) HOWEVER
|
|
32
|
+
CAUSED AND ON ANY THEORY OF LIABILITY, WHETHER IN CONTRACT, STRICT LIABILITY,
|
|
33
|
+
OR TORT (INCLUDING NEGLIGENCE OR OTHERWISE) ARISING IN ANY WAY OUT OF THE USE
|
|
34
|
+
OF THIS SOFTWARE, EVEN IF ADVISED OF THE POSSIBILITY OF SUCH DAMAGE.
|
|
35
|
+
Project-URL: Documentation, https://imml.readthedocs.io
|
|
36
|
+
Project-URL: Source, https://github.com/ocbe-uio/imml
|
|
37
|
+
Project-URL: Download, https://github.com/ocbe-uio/imml/archive/refs/heads/main.zip
|
|
38
|
+
Project-URL: Tracker, https://github.com/ocbe-uio/imml/issues
|
|
39
|
+
Project-URL: Changelog, https://imml.readthedocs.io/stable/development/changelog.html
|
|
40
|
+
Keywords: multi-modal learning,machine learning,incomplete data,missing data
|
|
41
|
+
Classifier: Intended Audience :: Science/Research
|
|
42
|
+
Classifier: Intended Audience :: Developers
|
|
43
|
+
Classifier: Development Status :: 3 - Alpha
|
|
44
|
+
Classifier: License :: OSI Approved :: BSD License
|
|
45
|
+
Classifier: Operating System :: Microsoft :: Windows
|
|
46
|
+
Classifier: Operating System :: POSIX
|
|
47
|
+
Classifier: Operating System :: Unix
|
|
48
|
+
Classifier: Operating System :: MacOS
|
|
49
|
+
Classifier: Topic :: Software Development
|
|
50
|
+
Classifier: Topic :: Scientific/Engineering
|
|
51
|
+
Classifier: Programming Language :: Python :: 3
|
|
52
|
+
Classifier: Programming Language :: Python :: 3.10
|
|
53
|
+
Classifier: Programming Language :: Python :: 3.11
|
|
54
|
+
Classifier: Programming Language :: Python :: 3.12
|
|
55
|
+
Classifier: Programming Language :: Python :: 3.13
|
|
56
|
+
Requires-Python: >=3.10
|
|
57
|
+
Description-Content-Type: text/markdown
|
|
58
|
+
Requires-Dist: scikit-learn>=1.4.1
|
|
59
|
+
Requires-Dist: pandas>=2.2.2
|
|
60
|
+
Requires-Dist: networkx>=2.5
|
|
61
|
+
Requires-Dist: h5py>=3.9.0
|
|
62
|
+
Requires-Dist: snfpy>=0
|
|
63
|
+
Requires-Dist: control>=0.10.1
|
|
64
|
+
Requires-Dist: cvxpy>=1.7.1
|
|
65
|
+
Requires-Dist: numba>=0.58.0
|
|
66
|
+
Provides-Extra: matlab
|
|
67
|
+
Requires-Dist: oct2py>=5.8.0; extra == "matlab"
|
|
68
|
+
Provides-Extra: r
|
|
69
|
+
Requires-Dist: rpy2>=3.5.14; extra == "r"
|
|
70
|
+
Provides-Extra: deep
|
|
71
|
+
Requires-Dist: lightning>=2.2; extra == "deep"
|
|
72
|
+
Requires-Dist: transformers>=4.51.3; extra == "deep"
|
|
73
|
+
Requires-Dist: torch_geometric>=2.6.1; extra == "deep"
|
|
74
|
+
Requires-Dist: torchvision>=0.19.1; extra == "deep"
|
|
75
|
+
Provides-Extra: docs
|
|
76
|
+
Requires-Dist: lightning>=2.2; extra == "docs"
|
|
77
|
+
Requires-Dist: transformers>=4.51.3; extra == "docs"
|
|
78
|
+
Requires-Dist: ipython>=8.20.0; extra == "docs"
|
|
79
|
+
Requires-Dist: datasets==3.6.0; extra == "docs"
|
|
80
|
+
Requires-Dist: rpy2>=3.5.14; extra == "docs"
|
|
81
|
+
Requires-Dist: sphinx>=7.0.0; extra == "docs"
|
|
82
|
+
Requires-Dist: sphinx-gallery>=0.17.1; extra == "docs"
|
|
83
|
+
Requires-Dist: sphinx_rtd_theme>=1.0.0; extra == "docs"
|
|
84
|
+
Provides-Extra: tests
|
|
85
|
+
Requires-Dist: pytest>=7.1.2; extra == "tests"
|
|
86
|
+
Requires-Dist: pytest-cov>=2.9.0; extra == "tests"
|
|
87
|
+
|
|
88
|
+

|
|
89
|
+

|
|
90
|
+
[](https://imml.readthedocs.io)
|
|
91
|
+
[](https://github.com/ocbe-uio/imml/actions/workflows/ci_test.yml)
|
|
92
|
+

|
|
93
|
+
[](https://github.com/ocbe-uio/imml/actions/workflows/github-code-scanning/codeql)
|
|
94
|
+
[](https://github.com/ocbe-uio/imml/pulls)
|
|
95
|
+

|
|
96
|
+
[](https://github.com/ocbe-uio/imml/blob/main/LICENSE)
|
|
97
|
+
|
|
98
|
+
[//]: # ([![DOI]()]())
|
|
99
|
+
[//]: # ([![Paper]()]())
|
|
100
|
+
|
|
101
|
+
<p align="center">
|
|
102
|
+
<img alt="iMML Logo" src="https://raw.githubusercontent.com/ocbe-uio/imml/refs/heads/main/docs/figures/logo_imml.png">
|
|
103
|
+
</p>
|
|
104
|
+
|
|
105
|
+
Overview
|
|
106
|
+
====================
|
|
107
|
+
|
|
108
|
+
*iMML* is a Python package that provides a **robust tool-set for integrating, processing, and analyzing incomplete
|
|
109
|
+
multi-modal datasets** to support a wide range of machine learning tasks. Starting with a dataset containing N samples
|
|
110
|
+
with K modalities, *iMML* effectively handles missing data for **classification, clustering, data retrieval,
|
|
111
|
+
imputation and amputation, feature selection, feature extraction and data exploration**, hence enabling efficient
|
|
112
|
+
analysis of partially observed samples.
|
|
113
|
+
|
|
114
|
+
|
|
115
|
+
<p align="center"><strong>Overview of iMML for multi-modal learning with incomplete data.</strong></p>
|
|
116
|
+
|
|
117
|
+
|
|
118
|
+
Background
|
|
119
|
+
----------
|
|
120
|
+
|
|
121
|
+
Multi-modal learning, where diverse data types are integrated and analyzed together, has emerged as a critical field
|
|
122
|
+
in artificial intelligence. Multi-modal machine learning models that effectively integrate multiple data modalities
|
|
123
|
+
generally outperform their uni-modal counterparts by leveraging more comprehensive and complementary information.
|
|
124
|
+
However, **most algorithms in this field assume fully observed data**, an assumption that is often
|
|
125
|
+
unrealistic in real-world scenarios.
|
|
126
|
+
|
|
127
|
+
Motivation
|
|
128
|
+
----------
|
|
129
|
+
|
|
130
|
+
Learning from incomplete multi-modal data has seen an important growth last years.
|
|
131
|
+
Despite this progress, several limitations still persist.
|
|
132
|
+
The landscape of available methods is fragmented, largely due to the diversity of use cases and data modalities,
|
|
133
|
+
which complicates both their application and benchmarking.
|
|
134
|
+
Systematic use and comparison of the current methods are often hindered by practical challenges, such as
|
|
135
|
+
incompatible input data formats and conflicting software dependencies.
|
|
136
|
+
As a result, researchers and practitioners frequently face challenges in choosing a practical method and invest
|
|
137
|
+
considerable efforts into reconciling codebases, rather than addressing the core scientific questions.
|
|
138
|
+
This suggests that **the community currently lacks robust and standardized tools to effectively handle
|
|
139
|
+
incomplete multi-modal data**.
|
|
140
|
+
|
|
141
|
+
Key features
|
|
142
|
+
------------
|
|
143
|
+
|
|
144
|
+
To address this gap, we have developed *iMML*, a Python package designed for multi-modal learning with incomplete data.
|
|
145
|
+
The key features of this package are:
|
|
146
|
+
|
|
147
|
+
- **Comprehensive toolkit**: *iMML* offers a broad set of tools for integrating, processing, and analyzing
|
|
148
|
+
incomplete multi-modal datasets implemented as a single, user-friendly interface to facilitate adoption by
|
|
149
|
+
a wide community of users.
|
|
150
|
+
The package includes extensive technical testing to ensure robustness, and thorough documentation enables
|
|
151
|
+
end-users to apply its functionality effectively.
|
|
152
|
+
- **Accessible**: *iMML* makes the tools readily available to the Python community, simplifying their usage,
|
|
153
|
+
comparison, and benchmarking, and thereby addresses the current lack of resources and standardized methods
|
|
154
|
+
for handling incomplete multi-modal data.
|
|
155
|
+
- **Extensible**: *iMML* provides a common framework where researchers can contribute and
|
|
156
|
+
integrate new approaches, serving as a community platform for hosting new algorithms and methods.
|
|
157
|
+
|
|
158
|
+
|
|
159
|
+
Installation
|
|
160
|
+
--------
|
|
161
|
+
|
|
162
|
+
Run the following command to install the most recent release of *iMML* using *pip*:
|
|
163
|
+
|
|
164
|
+
```bash
|
|
165
|
+
pip install imml
|
|
166
|
+
```
|
|
167
|
+
|
|
168
|
+
Or if you prefer *uv*, use:
|
|
169
|
+
|
|
170
|
+
```bash
|
|
171
|
+
uv pip install imml
|
|
172
|
+
```
|
|
173
|
+
|
|
174
|
+
Some features of *iMML* rely on optional dependencies. To enable these additional features, ensure you install
|
|
175
|
+
the required packages as described in our documentation: https://imml.readthedocs.io/stable/main/installation.html.
|
|
176
|
+
|
|
177
|
+
|
|
178
|
+
Usage
|
|
179
|
+
--------
|
|
180
|
+
|
|
181
|
+
This package provides a user-friendly interface to apply these algorithms to user-provided data.
|
|
182
|
+
*iMML* was designed to be compatible with widely-used machine learning and data analysis tools, such as Pandas,
|
|
183
|
+
NumPy, Scikit-learn, and Lightning AI, hence allowing researchers to **apply machine learning models with
|
|
184
|
+
minimal programming effort**.
|
|
185
|
+
Moreover, it can be easily integrated into Scikit-learn pipelines for data preprocessing and modeling.
|
|
186
|
+
|
|
187
|
+
For this demonstration, we will generate a random dataset, that we have called ``Xs``, as a multi-modal dataset
|
|
188
|
+
to simulate a multi-modal scenario:
|
|
189
|
+
|
|
190
|
+
```python
|
|
191
|
+
import numpy as np
|
|
192
|
+
Xs = [np.random.random((10,5)) for i in range(3)] # or your multi-modal dataset
|
|
193
|
+
```
|
|
194
|
+
|
|
195
|
+
You can use any other complete or incomplete multi-modal dataset. Once you have your dataset ready, you can
|
|
196
|
+
leverage the *iMML* library for a wide range of machine learning tasks, such as:
|
|
197
|
+
|
|
198
|
+
- Decompose a multi-modal dataset using ``MOFA`` to capture joint information.
|
|
199
|
+
|
|
200
|
+
```python
|
|
201
|
+
from imml.decomposition import MOFA
|
|
202
|
+
transformed_Xs = MOFA().fit_transform(Xs)
|
|
203
|
+
```
|
|
204
|
+
|
|
205
|
+
- Cluster samples from a multi-modal dataset using ``NEMO`` to find hidden groups.
|
|
206
|
+
|
|
207
|
+
```python
|
|
208
|
+
from imml.cluster import NEMO
|
|
209
|
+
labels = NEMO().fit_predict(Xs)
|
|
210
|
+
```
|
|
211
|
+
|
|
212
|
+
- Simulate incomplete multi-modal datasets for evaluation and testing purposes using ``Amputer``.
|
|
213
|
+
|
|
214
|
+
```python
|
|
215
|
+
from imml.ampute import Amputer
|
|
216
|
+
transformed_Xs = Amputer(p=0.8).fit_transform(Xs)
|
|
217
|
+
```
|
|
218
|
+
|
|
219
|
+
Free software
|
|
220
|
+
-------------
|
|
221
|
+
|
|
222
|
+
*iMML* is free software; you can redistribute it and/or modify it under the terms of the `BSD 3-Clause License`.
|
|
223
|
+
|
|
224
|
+
Contribute
|
|
225
|
+
------------
|
|
226
|
+
|
|
227
|
+
**We welcome practitioners, researchers, and the open-source community** to contribute to the *iMML* project,
|
|
228
|
+
and in doing so, helping us extend and refine the library for the community. Such a community-wide effort will
|
|
229
|
+
make *iMML* more versatile, sustainable, powerful, and accessible to the machine learning community across
|
|
230
|
+
many domains.
|
|
231
|
+
|
|
232
|
+
Project roadmap
|
|
233
|
+
------------
|
|
234
|
+
|
|
235
|
+
Our vision is to establish *iMML* as a leading and reliable library for multi-modal learning across research and
|
|
236
|
+
applied settings. Therefore, our priorities include to broaden algorithmic coverage, improve performance and
|
|
237
|
+
scalability, strengthen interoperability, and grow a healthy contributor community.
|
imml-0.1.0/README.md
ADDED
|
@@ -0,0 +1,150 @@
|
|
|
1
|
+

|
|
2
|
+

|
|
3
|
+
[](https://imml.readthedocs.io)
|
|
4
|
+
[](https://github.com/ocbe-uio/imml/actions/workflows/ci_test.yml)
|
|
5
|
+

|
|
6
|
+
[](https://github.com/ocbe-uio/imml/actions/workflows/github-code-scanning/codeql)
|
|
7
|
+
[](https://github.com/ocbe-uio/imml/pulls)
|
|
8
|
+

|
|
9
|
+
[](https://github.com/ocbe-uio/imml/blob/main/LICENSE)
|
|
10
|
+
|
|
11
|
+
[//]: # ([![DOI]()]())
|
|
12
|
+
[//]: # ([![Paper]()]())
|
|
13
|
+
|
|
14
|
+
<p align="center">
|
|
15
|
+
<img alt="iMML Logo" src="https://raw.githubusercontent.com/ocbe-uio/imml/refs/heads/main/docs/figures/logo_imml.png">
|
|
16
|
+
</p>
|
|
17
|
+
|
|
18
|
+
Overview
|
|
19
|
+
====================
|
|
20
|
+
|
|
21
|
+
*iMML* is a Python package that provides a **robust tool-set for integrating, processing, and analyzing incomplete
|
|
22
|
+
multi-modal datasets** to support a wide range of machine learning tasks. Starting with a dataset containing N samples
|
|
23
|
+
with K modalities, *iMML* effectively handles missing data for **classification, clustering, data retrieval,
|
|
24
|
+
imputation and amputation, feature selection, feature extraction and data exploration**, hence enabling efficient
|
|
25
|
+
analysis of partially observed samples.
|
|
26
|
+
|
|
27
|
+

|
|
28
|
+
<p align="center"><strong>Overview of iMML for multi-modal learning with incomplete data.</strong></p>
|
|
29
|
+
|
|
30
|
+
|
|
31
|
+
Background
|
|
32
|
+
----------
|
|
33
|
+
|
|
34
|
+
Multi-modal learning, where diverse data types are integrated and analyzed together, has emerged as a critical field
|
|
35
|
+
in artificial intelligence. Multi-modal machine learning models that effectively integrate multiple data modalities
|
|
36
|
+
generally outperform their uni-modal counterparts by leveraging more comprehensive and complementary information.
|
|
37
|
+
However, **most algorithms in this field assume fully observed data**, an assumption that is often
|
|
38
|
+
unrealistic in real-world scenarios.
|
|
39
|
+
|
|
40
|
+
Motivation
|
|
41
|
+
----------
|
|
42
|
+
|
|
43
|
+
Learning from incomplete multi-modal data has seen an important growth last years.
|
|
44
|
+
Despite this progress, several limitations still persist.
|
|
45
|
+
The landscape of available methods is fragmented, largely due to the diversity of use cases and data modalities,
|
|
46
|
+
which complicates both their application and benchmarking.
|
|
47
|
+
Systematic use and comparison of the current methods are often hindered by practical challenges, such as
|
|
48
|
+
incompatible input data formats and conflicting software dependencies.
|
|
49
|
+
As a result, researchers and practitioners frequently face challenges in choosing a practical method and invest
|
|
50
|
+
considerable efforts into reconciling codebases, rather than addressing the core scientific questions.
|
|
51
|
+
This suggests that **the community currently lacks robust and standardized tools to effectively handle
|
|
52
|
+
incomplete multi-modal data**.
|
|
53
|
+
|
|
54
|
+
Key features
|
|
55
|
+
------------
|
|
56
|
+
|
|
57
|
+
To address this gap, we have developed *iMML*, a Python package designed for multi-modal learning with incomplete data.
|
|
58
|
+
The key features of this package are:
|
|
59
|
+
|
|
60
|
+
- **Comprehensive toolkit**: *iMML* offers a broad set of tools for integrating, processing, and analyzing
|
|
61
|
+
incomplete multi-modal datasets implemented as a single, user-friendly interface to facilitate adoption by
|
|
62
|
+
a wide community of users.
|
|
63
|
+
The package includes extensive technical testing to ensure robustness, and thorough documentation enables
|
|
64
|
+
end-users to apply its functionality effectively.
|
|
65
|
+
- **Accessible**: *iMML* makes the tools readily available to the Python community, simplifying their usage,
|
|
66
|
+
comparison, and benchmarking, and thereby addresses the current lack of resources and standardized methods
|
|
67
|
+
for handling incomplete multi-modal data.
|
|
68
|
+
- **Extensible**: *iMML* provides a common framework where researchers can contribute and
|
|
69
|
+
integrate new approaches, serving as a community platform for hosting new algorithms and methods.
|
|
70
|
+
|
|
71
|
+
|
|
72
|
+
Installation
|
|
73
|
+
--------
|
|
74
|
+
|
|
75
|
+
Run the following command to install the most recent release of *iMML* using *pip*:
|
|
76
|
+
|
|
77
|
+
```bash
|
|
78
|
+
pip install imml
|
|
79
|
+
```
|
|
80
|
+
|
|
81
|
+
Or if you prefer *uv*, use:
|
|
82
|
+
|
|
83
|
+
```bash
|
|
84
|
+
uv pip install imml
|
|
85
|
+
```
|
|
86
|
+
|
|
87
|
+
Some features of *iMML* rely on optional dependencies. To enable these additional features, ensure you install
|
|
88
|
+
the required packages as described in our documentation: https://imml.readthedocs.io/stable/main/installation.html.
|
|
89
|
+
|
|
90
|
+
|
|
91
|
+
Usage
|
|
92
|
+
--------
|
|
93
|
+
|
|
94
|
+
This package provides a user-friendly interface to apply these algorithms to user-provided data.
|
|
95
|
+
*iMML* was designed to be compatible with widely-used machine learning and data analysis tools, such as Pandas,
|
|
96
|
+
NumPy, Scikit-learn, and Lightning AI, hence allowing researchers to **apply machine learning models with
|
|
97
|
+
minimal programming effort**.
|
|
98
|
+
Moreover, it can be easily integrated into Scikit-learn pipelines for data preprocessing and modeling.
|
|
99
|
+
|
|
100
|
+
For this demonstration, we will generate a random dataset, that we have called ``Xs``, as a multi-modal dataset
|
|
101
|
+
to simulate a multi-modal scenario:
|
|
102
|
+
|
|
103
|
+
```python
|
|
104
|
+
import numpy as np
|
|
105
|
+
Xs = [np.random.random((10,5)) for i in range(3)] # or your multi-modal dataset
|
|
106
|
+
```
|
|
107
|
+
|
|
108
|
+
You can use any other complete or incomplete multi-modal dataset. Once you have your dataset ready, you can
|
|
109
|
+
leverage the *iMML* library for a wide range of machine learning tasks, such as:
|
|
110
|
+
|
|
111
|
+
- Decompose a multi-modal dataset using ``MOFA`` to capture joint information.
|
|
112
|
+
|
|
113
|
+
```python
|
|
114
|
+
from imml.decomposition import MOFA
|
|
115
|
+
transformed_Xs = MOFA().fit_transform(Xs)
|
|
116
|
+
```
|
|
117
|
+
|
|
118
|
+
- Cluster samples from a multi-modal dataset using ``NEMO`` to find hidden groups.
|
|
119
|
+
|
|
120
|
+
```python
|
|
121
|
+
from imml.cluster import NEMO
|
|
122
|
+
labels = NEMO().fit_predict(Xs)
|
|
123
|
+
```
|
|
124
|
+
|
|
125
|
+
- Simulate incomplete multi-modal datasets for evaluation and testing purposes using ``Amputer``.
|
|
126
|
+
|
|
127
|
+
```python
|
|
128
|
+
from imml.ampute import Amputer
|
|
129
|
+
transformed_Xs = Amputer(p=0.8).fit_transform(Xs)
|
|
130
|
+
```
|
|
131
|
+
|
|
132
|
+
Free software
|
|
133
|
+
-------------
|
|
134
|
+
|
|
135
|
+
*iMML* is free software; you can redistribute it and/or modify it under the terms of the `BSD 3-Clause License`.
|
|
136
|
+
|
|
137
|
+
Contribute
|
|
138
|
+
------------
|
|
139
|
+
|
|
140
|
+
**We welcome practitioners, researchers, and the open-source community** to contribute to the *iMML* project,
|
|
141
|
+
and in doing so, helping us extend and refine the library for the community. Such a community-wide effort will
|
|
142
|
+
make *iMML* more versatile, sustainable, powerful, and accessible to the machine learning community across
|
|
143
|
+
many domains.
|
|
144
|
+
|
|
145
|
+
Project roadmap
|
|
146
|
+
------------
|
|
147
|
+
|
|
148
|
+
Our vision is to establish *iMML* as a leading and reliable library for multi-modal learning across research and
|
|
149
|
+
applied settings. Therefore, our priorities include to broaden algorithmic coverage, improve performance and
|
|
150
|
+
scalability, strengthen interoperability, and grow a healthy contributor community.
|
|
@@ -0,0 +1 @@
|
|
|
1
|
+
__version__ = "0.1.0"
|
|
@@ -0,0 +1,202 @@
|
|
|
1
|
+
import copy
|
|
2
|
+
from typing import List
|
|
3
|
+
|
|
4
|
+
import numpy as np
|
|
5
|
+
import pandas as pd
|
|
6
|
+
from sklearn.base import BaseEstimator, TransformerMixin
|
|
7
|
+
|
|
8
|
+
from . import remove_mods
|
|
9
|
+
|
|
10
|
+
|
|
11
|
+
class Amputer(BaseEstimator, TransformerMixin):
|
|
12
|
+
r"""
|
|
13
|
+
Simulate an incomplete multi-modal dataset with block-wise missing data from a fully observed multi-modal dataset.
|
|
14
|
+
|
|
15
|
+
Parameters
|
|
16
|
+
----------
|
|
17
|
+
p: float, default=0.1
|
|
18
|
+
Percentage of incomplete samples.
|
|
19
|
+
mechanism: str, default="mem"
|
|
20
|
+
One of ["mem", 'mcar', 'mnar', 'pm'], corresponding to mutually exclusive missing, missing completely at random,
|
|
21
|
+
missing not at random, and partial missing, respectively.
|
|
22
|
+
weights: list, default=None
|
|
23
|
+
The probabilities associated with each number of missing modalities. If not given, the sample
|
|
24
|
+
assumes a uniform distribution. Only used if mechanism = "mnar" or mechanism = "mem".
|
|
25
|
+
random_state: int, default=None
|
|
26
|
+
If int, random_state is the seed used by the random number generator.
|
|
27
|
+
|
|
28
|
+
Example
|
|
29
|
+
--------
|
|
30
|
+
>>> import numpy as np
|
|
31
|
+
>>> import pandas as pd
|
|
32
|
+
>>> from imml.ampute import Amputer
|
|
33
|
+
>>> Xs = [pd.DataFrame(np.random.default_rng(42).random((20, 10))) for i in range(3)]
|
|
34
|
+
>>> transformer = Amputer(p= 0.2, random_state=42)
|
|
35
|
+
>>> transformer.fit_transform(Xs)
|
|
36
|
+
"""
|
|
37
|
+
|
|
38
|
+
def __init__(self, p: float = 0.1, mechanism: str = "mem", weights: List = None, random_state: int = None):
|
|
39
|
+
|
|
40
|
+
mechanisms_options = ["mem", "mcar", "mnar", "pm"]
|
|
41
|
+
if mechanism not in mechanisms_options:
|
|
42
|
+
raise ValueError(f"Invalid mechanism. Expected one of: {mechanisms_options}")
|
|
43
|
+
if p < 0 or p > 1:
|
|
44
|
+
raise ValueError(f"Invalid p. Expected between 0 and 1.")
|
|
45
|
+
|
|
46
|
+
self.mechanism = mechanism
|
|
47
|
+
self.p = p
|
|
48
|
+
self.weights = weights
|
|
49
|
+
self.random_state = random_state
|
|
50
|
+
self.rng = np.random.default_rng(self.random_state)
|
|
51
|
+
|
|
52
|
+
|
|
53
|
+
def fit(self, Xs: List, y=None):
|
|
54
|
+
r"""
|
|
55
|
+
Fit the transformer to the input data.
|
|
56
|
+
|
|
57
|
+
Parameters
|
|
58
|
+
----------
|
|
59
|
+
Xs : list of array-likes objects
|
|
60
|
+
- Xs length: n_mods
|
|
61
|
+
- Xs[i] shape: (n_samples, n_features_i)
|
|
62
|
+
|
|
63
|
+
A list of different modalities.
|
|
64
|
+
y : Ignored
|
|
65
|
+
Not used, present here for API consistency by convention.
|
|
66
|
+
|
|
67
|
+
Returns
|
|
68
|
+
-------
|
|
69
|
+
self : returns an instance of self.
|
|
70
|
+
"""
|
|
71
|
+
self.n_mods = len(Xs)
|
|
72
|
+
return self
|
|
73
|
+
|
|
74
|
+
|
|
75
|
+
def transform(self, Xs: List):
|
|
76
|
+
r"""
|
|
77
|
+
Ampute a fully observed multi-modal dataset.
|
|
78
|
+
|
|
79
|
+
Parameters
|
|
80
|
+
----------
|
|
81
|
+
Xs : list of array-likes objects
|
|
82
|
+
- Xs length: n_mods
|
|
83
|
+
- Xs[i] shape: (n_samples, n_features_i)
|
|
84
|
+
|
|
85
|
+
A list of different modalities.
|
|
86
|
+
|
|
87
|
+
Returns
|
|
88
|
+
-------
|
|
89
|
+
transformed_Xs : list of array-likes objects, shape (n_samples, n_features), length n_mods
|
|
90
|
+
The amputed multi-modal dataset.
|
|
91
|
+
"""
|
|
92
|
+
if self.p > 0:
|
|
93
|
+
pandas_format = isinstance(Xs[0], pd.DataFrame)
|
|
94
|
+
if pandas_format:
|
|
95
|
+
rownames = Xs[0].index
|
|
96
|
+
colnames = [X.columns for X in Xs]
|
|
97
|
+
Xs = [X.values for X in Xs]
|
|
98
|
+
sample_names = pd.Index(list(range(len(Xs[0]))))
|
|
99
|
+
|
|
100
|
+
if self.mechanism == "mem":
|
|
101
|
+
pseudo_observed_mod_indicator = self._mem_mask(sample_names=sample_names)
|
|
102
|
+
elif self.mechanism == "mcar":
|
|
103
|
+
pseudo_observed_mod_indicator = self._mcar_mask(sample_names=sample_names)
|
|
104
|
+
elif self.mechanism == "pm":
|
|
105
|
+
pseudo_observed_mod_indicator = self._pm_mask(sample_names=sample_names)
|
|
106
|
+
elif self.mechanism == "mnar":
|
|
107
|
+
pseudo_observed_mod_indicator = self._mnar_mask(sample_names=sample_names)
|
|
108
|
+
|
|
109
|
+
pseudo_observed_mod_indicator = pseudo_observed_mod_indicator.astype(bool)
|
|
110
|
+
transformed_Xs = remove_mods(Xs=Xs, observed_mod_indicator=pseudo_observed_mod_indicator)
|
|
111
|
+
|
|
112
|
+
if pandas_format:
|
|
113
|
+
transformed_Xs = [pd.DataFrame(X, index=rownames, columns=colnames[X_idx])
|
|
114
|
+
for X_idx, X in enumerate(transformed_Xs)]
|
|
115
|
+
else:
|
|
116
|
+
transformed_Xs = Xs
|
|
117
|
+
|
|
118
|
+
return transformed_Xs
|
|
119
|
+
|
|
120
|
+
|
|
121
|
+
def _mem_mask(self, sample_names):
|
|
122
|
+
pseudo_observed_mod_indicator = pd.DataFrame(np.ones((len(sample_names), self.n_mods)), index=sample_names)
|
|
123
|
+
common_samples = pd.Series(sample_names, index=sample_names).sample(frac=1 - self.p, replace=False,
|
|
124
|
+
random_state=self.random_state).index
|
|
125
|
+
sampled_names = copy.deepcopy(common_samples)
|
|
126
|
+
if self.weights is None:
|
|
127
|
+
n_missings = int(len(sample_names.difference(sampled_names)) / self.n_mods)
|
|
128
|
+
n_missings = [n_missings] * self.n_mods
|
|
129
|
+
else:
|
|
130
|
+
n_missings = [int(len(sample_names.difference(sampled_names)) * w) for w in self.weights]
|
|
131
|
+
for X_idx,n_missing in enumerate(n_missings):
|
|
132
|
+
x_per_mod = sample_names.difference(sampled_names)
|
|
133
|
+
if X_idx != self.n_mods - 1:
|
|
134
|
+
x_per_mod = pd.Series(x_per_mod, index=x_per_mod).sample(n=n_missing,
|
|
135
|
+
replace=False,
|
|
136
|
+
random_state=self.random_state).index
|
|
137
|
+
sampled_names = sampled_names.append(x_per_mod)
|
|
138
|
+
idxs_to_remove = common_samples.append(x_per_mod)
|
|
139
|
+
idxs_to_remove = sample_names.difference(idxs_to_remove)
|
|
140
|
+
pseudo_observed_mod_indicator.loc[idxs_to_remove, X_idx] = 0
|
|
141
|
+
return pseudo_observed_mod_indicator
|
|
142
|
+
|
|
143
|
+
|
|
144
|
+
def _mcar_mask(self, sample_names):
|
|
145
|
+
pseudo_observed_mod_indicator = pd.DataFrame(np.ones((len(sample_names), self.n_mods)), index=sample_names)
|
|
146
|
+
common_samples = pd.Series(sample_names, index=sample_names).sample(frac=1 - self.p, replace=False,
|
|
147
|
+
random_state=self.random_state).index
|
|
148
|
+
idxs_to_remove = sample_names.difference(common_samples)
|
|
149
|
+
shape = pseudo_observed_mod_indicator.loc[idxs_to_remove].shape
|
|
150
|
+
mask = self.rng.choice(2, size=shape)
|
|
151
|
+
mask = pd.DataFrame(mask, index=idxs_to_remove)
|
|
152
|
+
samples_to_fix = mask.nunique(axis=1).eq(1)
|
|
153
|
+
if samples_to_fix.any():
|
|
154
|
+
samples_to_fix = samples_to_fix[samples_to_fix]
|
|
155
|
+
mods_to_fix = self.rng.integers(low=0, high=self.n_mods, size=len(samples_to_fix))
|
|
156
|
+
for mod_idx in np.unique(mods_to_fix):
|
|
157
|
+
samples = mods_to_fix == mod_idx
|
|
158
|
+
samples = samples_to_fix[samples].index
|
|
159
|
+
mask.loc[samples, mod_idx] = np.invert(mask.loc[samples, mod_idx].astype(bool)).astype(int)
|
|
160
|
+
|
|
161
|
+
pseudo_observed_mod_indicator.loc[idxs_to_remove] = mask.astype(int)
|
|
162
|
+
return pseudo_observed_mod_indicator
|
|
163
|
+
|
|
164
|
+
|
|
165
|
+
def _mnar_mask(self, sample_names):
|
|
166
|
+
mask = pd.DataFrame(np.ones((len(sample_names), self.n_mods)), index=sample_names)
|
|
167
|
+
common_samples = pd.Series(sample_names, index=sample_names).sample(frac=1 - self.p, replace=False,
|
|
168
|
+
random_state=self.random_state).index
|
|
169
|
+
idxs_to_remove = sample_names.difference(common_samples)
|
|
170
|
+
reference_var = self.rng.choice(range(1, self.n_mods), p = self.weights, size=len(idxs_to_remove))
|
|
171
|
+
reference_var = pd.Series(reference_var, index=idxs_to_remove)
|
|
172
|
+
n_mods_to_remove = {n_mods_to_remove: self.rng.choice(self.n_mods, size=n_mods_to_remove, replace=False)
|
|
173
|
+
for n_mods_to_remove in np.unique(reference_var)}
|
|
174
|
+
for keys,values in n_mods_to_remove.items():
|
|
175
|
+
mask.loc[reference_var[reference_var == keys].index, values] = 0
|
|
176
|
+
|
|
177
|
+
return mask
|
|
178
|
+
|
|
179
|
+
|
|
180
|
+
def _pm_mask(self, sample_names):
|
|
181
|
+
pseudo_observed_mod_indicator = pd.DataFrame(np.ones((len(sample_names), self.n_mods)), index=sample_names)
|
|
182
|
+
common_samples = pd.Series(sample_names, index=sample_names).sample(frac=1 - self.p, replace=False,
|
|
183
|
+
random_state=self.random_state).index
|
|
184
|
+
idxs_to_remove = sample_names.difference(common_samples)
|
|
185
|
+
n_incomplete_modalities = self.rng.choice(np.arange(1, self.n_mods), size=1)[0]
|
|
186
|
+
if (self.n_mods == 2) or (n_incomplete_modalities == 1):
|
|
187
|
+
col = self.rng.choice(self.n_mods)
|
|
188
|
+
pseudo_observed_mod_indicator.loc[idxs_to_remove, col] = 0
|
|
189
|
+
else:
|
|
190
|
+
mask = self.rng.choice(2, size=(len(idxs_to_remove), n_incomplete_modalities))
|
|
191
|
+
mask = pd.DataFrame(mask, index=idxs_to_remove,
|
|
192
|
+
columns=self.rng.choice(self.n_mods, size=n_incomplete_modalities, replace=False))
|
|
193
|
+
samples_to_fix = mask.nunique(axis=1).eq(1)
|
|
194
|
+
if samples_to_fix.any():
|
|
195
|
+
samples_to_fix = samples_to_fix[samples_to_fix]
|
|
196
|
+
mods_to_fix = self.rng.choice(mask.columns, size=len(samples_to_fix))
|
|
197
|
+
for mod_idx in np.unique(mods_to_fix):
|
|
198
|
+
samples = mods_to_fix == mod_idx
|
|
199
|
+
samples = samples_to_fix[samples].index
|
|
200
|
+
mask.loc[samples, mod_idx] = np.invert(mask.loc[samples, mod_idx].astype(bool)).astype(int)
|
|
201
|
+
pseudo_observed_mod_indicator.loc[idxs_to_remove, mask.columns] = mask.astype(int)
|
|
202
|
+
return pseudo_observed_mod_indicator
|