InterSCellar 0.1.0__tar.gz
This diff represents the content of publicly available package versions that have been released to one of the supported registries. The information contained in this diff is provided for informational purposes only and reflects changes between package versions as they appear in their respective public registries.
- interscellar-0.1.0/InterSCellar.egg-info/PKG-INFO +136 -0
- interscellar-0.1.0/InterSCellar.egg-info/SOURCES.txt +20 -0
- interscellar-0.1.0/InterSCellar.egg-info/dependency_links.txt +1 -0
- interscellar-0.1.0/InterSCellar.egg-info/entry_points.txt +3 -0
- interscellar-0.1.0/InterSCellar.egg-info/requires.txt +23 -0
- interscellar-0.1.0/InterSCellar.egg-info/top_level.txt +1 -0
- interscellar-0.1.0/LICENSE +21 -0
- interscellar-0.1.0/PKG-INFO +136 -0
- interscellar-0.1.0/README.md +91 -0
- interscellar-0.1.0/pyproject.toml +86 -0
- interscellar-0.1.0/setup.cfg +4 -0
- interscellar-0.1.0/src/__init__.py +22 -0
- interscellar-0.1.0/src/api/__init__.py +14 -0
- interscellar-0.1.0/src/api/wrapper_2d.py +176 -0
- interscellar-0.1.0/src/api/wrapper_3d.py +632 -0
- interscellar-0.1.0/src/core/__init__.py +80 -0
- interscellar-0.1.0/src/core/compute_interscellar_volumes_3d.py +2221 -0
- interscellar-0.1.0/src/core/find_cell_neighbors_2d.py +997 -0
- interscellar-0.1.0/src/core/find_cell_neighbors_3d.py +1281 -0
- interscellar-0.1.0/src/visualization/__init__.py +7 -0
- interscellar-0.1.0/src/visualization/visualize_all_3d.py +177 -0
- interscellar-0.1.0/src/visualization/visualize_pair_3d.py +333 -0
|
@@ -0,0 +1,136 @@
|
|
|
1
|
+
Metadata-Version: 2.4
|
|
2
|
+
Name: InterSCellar
|
|
3
|
+
Version: 0.1.0
|
|
4
|
+
Summary: Surface-based Cell Neighbor Detection and Interscellar Volume Computation for 2D & 3D Spatial Omics
|
|
5
|
+
Author-email: Eunice Yewon Lee <yl888@duke.edu>
|
|
6
|
+
License: MIT
|
|
7
|
+
Project-URL: Homepage, https://github.com/euniceyl/InterSCellar
|
|
8
|
+
Project-URL: Repository, https://github.com/euniceyl/InterSCellar
|
|
9
|
+
Project-URL: Issues, https://github.com/euniceyl/InterSCellar/issues
|
|
10
|
+
Keywords: spatial-omics,cell-neighbors,cell-interactions,cell-cell-communication,intercellular,single-cell,transcriptomics,proteomics,metabolomics,bioinformatics,computational-biology,visualization,3d
|
|
11
|
+
Classifier: Development Status :: 3 - Alpha
|
|
12
|
+
Classifier: Intended Audience :: Science/Research
|
|
13
|
+
Classifier: License :: OSI Approved :: MIT License
|
|
14
|
+
Classifier: Operating System :: OS Independent
|
|
15
|
+
Classifier: Programming Language :: Python :: 3
|
|
16
|
+
Classifier: Programming Language :: Python :: 3.8
|
|
17
|
+
Classifier: Programming Language :: Python :: 3.9
|
|
18
|
+
Classifier: Programming Language :: Python :: 3.10
|
|
19
|
+
Classifier: Programming Language :: Python :: 3.11
|
|
20
|
+
Classifier: Topic :: Scientific/Engineering :: Bio-Informatics
|
|
21
|
+
Requires-Python: >=3.8
|
|
22
|
+
Description-Content-Type: text/markdown
|
|
23
|
+
License-File: LICENSE
|
|
24
|
+
Requires-Dist: numpy>=1.20.0
|
|
25
|
+
Requires-Dist: pandas>=1.3.0
|
|
26
|
+
Requires-Dist: scipy>=1.7.0
|
|
27
|
+
Requires-Dist: tqdm>=4.62.0
|
|
28
|
+
Requires-Dist: opencv-python>=4.5.0
|
|
29
|
+
Requires-Dist: dask[array]>=2021.0.0
|
|
30
|
+
Requires-Dist: matplotlib>=3.4.0
|
|
31
|
+
Requires-Dist: scikit-image>=0.18.0
|
|
32
|
+
Requires-Dist: seaborn>=0.11.0
|
|
33
|
+
Requires-Dist: napari>=0.4.0
|
|
34
|
+
Requires-Dist: zarr>=2.10.0
|
|
35
|
+
Requires-Dist: anndata>=0.8.0
|
|
36
|
+
Requires-Dist: duckdb>=0.6.0
|
|
37
|
+
Provides-Extra: anndata
|
|
38
|
+
Requires-Dist: anndata>=0.8.0; extra == "anndata"
|
|
39
|
+
Provides-Extra: duckdb
|
|
40
|
+
Requires-Dist: duckdb>=0.6.0; extra == "duckdb"
|
|
41
|
+
Provides-Extra: all
|
|
42
|
+
Requires-Dist: anndata>=0.8.0; extra == "all"
|
|
43
|
+
Requires-Dist: duckdb>=0.6.0; extra == "all"
|
|
44
|
+
Dynamic: license-file
|
|
45
|
+
|
|
46
|
+
# InterSCellar
|
|
47
|
+
[](https://pypi.org/project/interscellar/)
|
|
48
|
+
[](https://opensource.org/licenses/MIT)
|
|
49
|
+
|
|
50
|
+
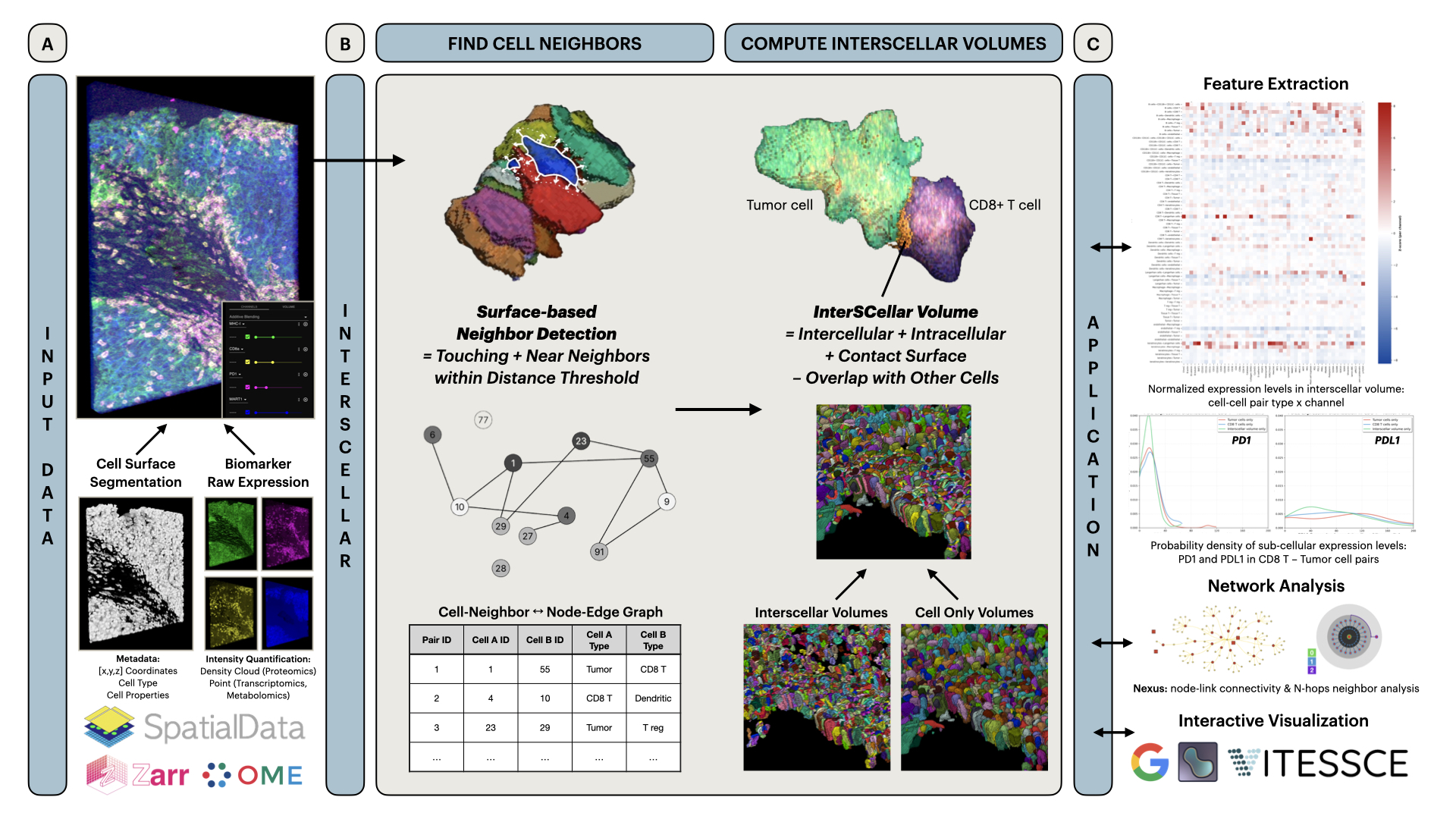
**InterSCellar** is a Python package for surface-Based cell neighborhood and interaction volume analysis in 3D spatial omics.
|
|
51
|
+
|
|
52
|
+
|
|
53
|
+
|
|
54
|
+
## Installation
|
|
55
|
+
|
|
56
|
+
**Install package:**
|
|
57
|
+
```sh
|
|
58
|
+
pip install interscellar
|
|
59
|
+
```
|
|
60
|
+
|
|
61
|
+
## Usage
|
|
62
|
+
|
|
63
|
+
**Import:**
|
|
64
|
+
```sh
|
|
65
|
+
import interscellar
|
|
66
|
+
```
|
|
67
|
+
|
|
68
|
+
### 3D Pipeline:
|
|
69
|
+
|
|
70
|
+
**(1) Cell Neighbor Detection & Graph Construction**
|
|
71
|
+
```sh
|
|
72
|
+
neighbors_3d, adata, conn = interscellar.find_cell_neighbors_3d(
|
|
73
|
+
ome_zarr_path="data/segmentation.zarr",
|
|
74
|
+
metadata_csv_path="data/cell_metadata.csv",
|
|
75
|
+
max_distance_um=0.5,
|
|
76
|
+
voxel_size_um=(0.56, 0.28, 0.28),
|
|
77
|
+
n_jobs=4
|
|
78
|
+
)
|
|
79
|
+
```
|
|
80
|
+
|
|
81
|
+
**(2) Interscellar Volume Computation**
|
|
82
|
+
```sh
|
|
83
|
+
# Interscellar volumes
|
|
84
|
+
volumes_3d, adata, conn = interscellar.compute_interscellar_volumes_3d(
|
|
85
|
+
ome_zarr_path="data/segmentation.zarr",
|
|
86
|
+
neighbor_pairs_csv="results/neighbors_3d.csv",
|
|
87
|
+
neighbor_db_path="/results/neighbor_graph.db",
|
|
88
|
+
voxel_size_um=(0.56, 0.28, 0.28),
|
|
89
|
+
max_distance_um=3.0,
|
|
90
|
+
intracellular_threshold_um=1.0,
|
|
91
|
+
n_jobs=4
|
|
92
|
+
)
|
|
93
|
+
```
|
|
94
|
+
|
|
95
|
+
```sh
|
|
96
|
+
# Cell-only volumes
|
|
97
|
+
cellonly_3d = interscellar.compute_cell_only_volumes_3d(
|
|
98
|
+
ome_zarr_path="data/segmentation.zarr",
|
|
99
|
+
interscellar_volumes_zarr="results/interscellar_volumes.zarr"
|
|
100
|
+
)
|
|
101
|
+
```
|
|
102
|
+
|
|
103
|
+
### 2D Pipeline:
|
|
104
|
+
|
|
105
|
+
**(1) Cell Neighbor Detection & Graph Construction**
|
|
106
|
+
```sh
|
|
107
|
+
neighbors_2d, adata, conn = interscellar.find_cell_neighbors_2d(
|
|
108
|
+
polygon_json_path="data/cell_polygons.json",
|
|
109
|
+
metadata_csv_path="data/cell_metadata.csv",
|
|
110
|
+
max_distance_um=1.0,
|
|
111
|
+
pixel_size_um=0.1085,
|
|
112
|
+
n_jobs=4
|
|
113
|
+
)
|
|
114
|
+
```
|
|
115
|
+
|
|
116
|
+
### Utilities:
|
|
117
|
+
|
|
118
|
+
**Volume Visualization**
|
|
119
|
+
```sh
|
|
120
|
+
# Full dataset (Napari)
|
|
121
|
+
visualize-all-3d \
|
|
122
|
+
--cell-only-zarr "results/cell_only_volumes.zarr" \
|
|
123
|
+
--interscellar-zarr "results/interscellar_volumes.zarr" \
|
|
124
|
+
--cell-only-opacity 0.7 \
|
|
125
|
+
--interscellar-opacity 0.9
|
|
126
|
+
```
|
|
127
|
+
|
|
128
|
+
```sh
|
|
129
|
+
# Single pair (Napari)
|
|
130
|
+
visualize-pair-3d \
|
|
131
|
+
--pair-id 123 \
|
|
132
|
+
--cell-only-zarr "results/cell_only_volumes.zarr" \
|
|
133
|
+
--interscellar-zarr "results/interscellar_volumes.zarr" \
|
|
134
|
+
--pair-opacity 0.6 \
|
|
135
|
+
--cells-opacity 0.7
|
|
136
|
+
```
|
|
@@ -0,0 +1,20 @@
|
|
|
1
|
+
LICENSE
|
|
2
|
+
README.md
|
|
3
|
+
pyproject.toml
|
|
4
|
+
InterSCellar.egg-info/PKG-INFO
|
|
5
|
+
InterSCellar.egg-info/SOURCES.txt
|
|
6
|
+
InterSCellar.egg-info/dependency_links.txt
|
|
7
|
+
InterSCellar.egg-info/entry_points.txt
|
|
8
|
+
InterSCellar.egg-info/requires.txt
|
|
9
|
+
InterSCellar.egg-info/top_level.txt
|
|
10
|
+
src/__init__.py
|
|
11
|
+
src/api/__init__.py
|
|
12
|
+
src/api/wrapper_2d.py
|
|
13
|
+
src/api/wrapper_3d.py
|
|
14
|
+
src/core/__init__.py
|
|
15
|
+
src/core/compute_interscellar_volumes_3d.py
|
|
16
|
+
src/core/find_cell_neighbors_2d.py
|
|
17
|
+
src/core/find_cell_neighbors_3d.py
|
|
18
|
+
src/visualization/__init__.py
|
|
19
|
+
src/visualization/visualize_all_3d.py
|
|
20
|
+
src/visualization/visualize_pair_3d.py
|
|
@@ -0,0 +1 @@
|
|
|
1
|
+
|
|
@@ -0,0 +1,23 @@
|
|
|
1
|
+
numpy>=1.20.0
|
|
2
|
+
pandas>=1.3.0
|
|
3
|
+
scipy>=1.7.0
|
|
4
|
+
tqdm>=4.62.0
|
|
5
|
+
opencv-python>=4.5.0
|
|
6
|
+
dask[array]>=2021.0.0
|
|
7
|
+
matplotlib>=3.4.0
|
|
8
|
+
scikit-image>=0.18.0
|
|
9
|
+
seaborn>=0.11.0
|
|
10
|
+
napari>=0.4.0
|
|
11
|
+
zarr>=2.10.0
|
|
12
|
+
anndata>=0.8.0
|
|
13
|
+
duckdb>=0.6.0
|
|
14
|
+
|
|
15
|
+
[all]
|
|
16
|
+
anndata>=0.8.0
|
|
17
|
+
duckdb>=0.6.0
|
|
18
|
+
|
|
19
|
+
[anndata]
|
|
20
|
+
anndata>=0.8.0
|
|
21
|
+
|
|
22
|
+
[duckdb]
|
|
23
|
+
duckdb>=0.6.0
|
|
@@ -0,0 +1 @@
|
|
|
1
|
+
interscellar
|
|
@@ -0,0 +1,21 @@
|
|
|
1
|
+
MIT License
|
|
2
|
+
|
|
3
|
+
Copyright (c) 2025 Eunice Lee
|
|
4
|
+
|
|
5
|
+
Permission is hereby granted, free of charge, to any person obtaining a copy
|
|
6
|
+
of this software and associated documentation files (the "Software"), to deal
|
|
7
|
+
in the Software without restriction, including without limitation the rights
|
|
8
|
+
to use, copy, modify, merge, publish, distribute, sublicense, and/or sell
|
|
9
|
+
copies of the Software, and to permit persons to whom the Software is
|
|
10
|
+
furnished to do so, subject to the following conditions:
|
|
11
|
+
|
|
12
|
+
The above copyright notice and this permission notice shall be included in all
|
|
13
|
+
copies or substantial portions of the Software.
|
|
14
|
+
|
|
15
|
+
THE SOFTWARE IS PROVIDED "AS IS", WITHOUT WARRANTY OF ANY KIND, EXPRESS OR
|
|
16
|
+
IMPLIED, INCLUDING BUT NOT LIMITED TO THE WARRANTIES OF MERCHANTABILITY,
|
|
17
|
+
FITNESS FOR A PARTICULAR PURPOSE AND NONINFRINGEMENT. IN NO EVENT SHALL THE
|
|
18
|
+
AUTHORS OR COPYRIGHT HOLDERS BE LIABLE FOR ANY CLAIM, DAMAGES OR OTHER
|
|
19
|
+
LIABILITY, WHETHER IN AN ACTION OF CONTRACT, TORT OR OTHERWISE, ARISING FROM,
|
|
20
|
+
OUT OF OR IN CONNECTION WITH THE SOFTWARE OR THE USE OR OTHER DEALINGS IN THE
|
|
21
|
+
SOFTWARE.
|
|
@@ -0,0 +1,136 @@
|
|
|
1
|
+
Metadata-Version: 2.4
|
|
2
|
+
Name: InterSCellar
|
|
3
|
+
Version: 0.1.0
|
|
4
|
+
Summary: Surface-based Cell Neighbor Detection and Interscellar Volume Computation for 2D & 3D Spatial Omics
|
|
5
|
+
Author-email: Eunice Yewon Lee <yl888@duke.edu>
|
|
6
|
+
License: MIT
|
|
7
|
+
Project-URL: Homepage, https://github.com/euniceyl/InterSCellar
|
|
8
|
+
Project-URL: Repository, https://github.com/euniceyl/InterSCellar
|
|
9
|
+
Project-URL: Issues, https://github.com/euniceyl/InterSCellar/issues
|
|
10
|
+
Keywords: spatial-omics,cell-neighbors,cell-interactions,cell-cell-communication,intercellular,single-cell,transcriptomics,proteomics,metabolomics,bioinformatics,computational-biology,visualization,3d
|
|
11
|
+
Classifier: Development Status :: 3 - Alpha
|
|
12
|
+
Classifier: Intended Audience :: Science/Research
|
|
13
|
+
Classifier: License :: OSI Approved :: MIT License
|
|
14
|
+
Classifier: Operating System :: OS Independent
|
|
15
|
+
Classifier: Programming Language :: Python :: 3
|
|
16
|
+
Classifier: Programming Language :: Python :: 3.8
|
|
17
|
+
Classifier: Programming Language :: Python :: 3.9
|
|
18
|
+
Classifier: Programming Language :: Python :: 3.10
|
|
19
|
+
Classifier: Programming Language :: Python :: 3.11
|
|
20
|
+
Classifier: Topic :: Scientific/Engineering :: Bio-Informatics
|
|
21
|
+
Requires-Python: >=3.8
|
|
22
|
+
Description-Content-Type: text/markdown
|
|
23
|
+
License-File: LICENSE
|
|
24
|
+
Requires-Dist: numpy>=1.20.0
|
|
25
|
+
Requires-Dist: pandas>=1.3.0
|
|
26
|
+
Requires-Dist: scipy>=1.7.0
|
|
27
|
+
Requires-Dist: tqdm>=4.62.0
|
|
28
|
+
Requires-Dist: opencv-python>=4.5.0
|
|
29
|
+
Requires-Dist: dask[array]>=2021.0.0
|
|
30
|
+
Requires-Dist: matplotlib>=3.4.0
|
|
31
|
+
Requires-Dist: scikit-image>=0.18.0
|
|
32
|
+
Requires-Dist: seaborn>=0.11.0
|
|
33
|
+
Requires-Dist: napari>=0.4.0
|
|
34
|
+
Requires-Dist: zarr>=2.10.0
|
|
35
|
+
Requires-Dist: anndata>=0.8.0
|
|
36
|
+
Requires-Dist: duckdb>=0.6.0
|
|
37
|
+
Provides-Extra: anndata
|
|
38
|
+
Requires-Dist: anndata>=0.8.0; extra == "anndata"
|
|
39
|
+
Provides-Extra: duckdb
|
|
40
|
+
Requires-Dist: duckdb>=0.6.0; extra == "duckdb"
|
|
41
|
+
Provides-Extra: all
|
|
42
|
+
Requires-Dist: anndata>=0.8.0; extra == "all"
|
|
43
|
+
Requires-Dist: duckdb>=0.6.0; extra == "all"
|
|
44
|
+
Dynamic: license-file
|
|
45
|
+
|
|
46
|
+
# InterSCellar
|
|
47
|
+
[](https://pypi.org/project/interscellar/)
|
|
48
|
+
[](https://opensource.org/licenses/MIT)
|
|
49
|
+
|
|
50
|
+
**InterSCellar** is a Python package for surface-Based cell neighborhood and interaction volume analysis in 3D spatial omics.
|
|
51
|
+
|
|
52
|
+

|
|
53
|
+
|
|
54
|
+
## Installation
|
|
55
|
+
|
|
56
|
+
**Install package:**
|
|
57
|
+
```sh
|
|
58
|
+
pip install interscellar
|
|
59
|
+
```
|
|
60
|
+
|
|
61
|
+
## Usage
|
|
62
|
+
|
|
63
|
+
**Import:**
|
|
64
|
+
```sh
|
|
65
|
+
import interscellar
|
|
66
|
+
```
|
|
67
|
+
|
|
68
|
+
### 3D Pipeline:
|
|
69
|
+
|
|
70
|
+
**(1) Cell Neighbor Detection & Graph Construction**
|
|
71
|
+
```sh
|
|
72
|
+
neighbors_3d, adata, conn = interscellar.find_cell_neighbors_3d(
|
|
73
|
+
ome_zarr_path="data/segmentation.zarr",
|
|
74
|
+
metadata_csv_path="data/cell_metadata.csv",
|
|
75
|
+
max_distance_um=0.5,
|
|
76
|
+
voxel_size_um=(0.56, 0.28, 0.28),
|
|
77
|
+
n_jobs=4
|
|
78
|
+
)
|
|
79
|
+
```
|
|
80
|
+
|
|
81
|
+
**(2) Interscellar Volume Computation**
|
|
82
|
+
```sh
|
|
83
|
+
# Interscellar volumes
|
|
84
|
+
volumes_3d, adata, conn = interscellar.compute_interscellar_volumes_3d(
|
|
85
|
+
ome_zarr_path="data/segmentation.zarr",
|
|
86
|
+
neighbor_pairs_csv="results/neighbors_3d.csv",
|
|
87
|
+
neighbor_db_path="/results/neighbor_graph.db",
|
|
88
|
+
voxel_size_um=(0.56, 0.28, 0.28),
|
|
89
|
+
max_distance_um=3.0,
|
|
90
|
+
intracellular_threshold_um=1.0,
|
|
91
|
+
n_jobs=4
|
|
92
|
+
)
|
|
93
|
+
```
|
|
94
|
+
|
|
95
|
+
```sh
|
|
96
|
+
# Cell-only volumes
|
|
97
|
+
cellonly_3d = interscellar.compute_cell_only_volumes_3d(
|
|
98
|
+
ome_zarr_path="data/segmentation.zarr",
|
|
99
|
+
interscellar_volumes_zarr="results/interscellar_volumes.zarr"
|
|
100
|
+
)
|
|
101
|
+
```
|
|
102
|
+
|
|
103
|
+
### 2D Pipeline:
|
|
104
|
+
|
|
105
|
+
**(1) Cell Neighbor Detection & Graph Construction**
|
|
106
|
+
```sh
|
|
107
|
+
neighbors_2d, adata, conn = interscellar.find_cell_neighbors_2d(
|
|
108
|
+
polygon_json_path="data/cell_polygons.json",
|
|
109
|
+
metadata_csv_path="data/cell_metadata.csv",
|
|
110
|
+
max_distance_um=1.0,
|
|
111
|
+
pixel_size_um=0.1085,
|
|
112
|
+
n_jobs=4
|
|
113
|
+
)
|
|
114
|
+
```
|
|
115
|
+
|
|
116
|
+
### Utilities:
|
|
117
|
+
|
|
118
|
+
**Volume Visualization**
|
|
119
|
+
```sh
|
|
120
|
+
# Full dataset (Napari)
|
|
121
|
+
visualize-all-3d \
|
|
122
|
+
--cell-only-zarr "results/cell_only_volumes.zarr" \
|
|
123
|
+
--interscellar-zarr "results/interscellar_volumes.zarr" \
|
|
124
|
+
--cell-only-opacity 0.7 \
|
|
125
|
+
--interscellar-opacity 0.9
|
|
126
|
+
```
|
|
127
|
+
|
|
128
|
+
```sh
|
|
129
|
+
# Single pair (Napari)
|
|
130
|
+
visualize-pair-3d \
|
|
131
|
+
--pair-id 123 \
|
|
132
|
+
--cell-only-zarr "results/cell_only_volumes.zarr" \
|
|
133
|
+
--interscellar-zarr "results/interscellar_volumes.zarr" \
|
|
134
|
+
--pair-opacity 0.6 \
|
|
135
|
+
--cells-opacity 0.7
|
|
136
|
+
```
|
|
@@ -0,0 +1,91 @@
|
|
|
1
|
+
# InterSCellar
|
|
2
|
+
[](https://pypi.org/project/interscellar/)
|
|
3
|
+
[](https://opensource.org/licenses/MIT)
|
|
4
|
+
|
|
5
|
+
**InterSCellar** is a Python package for surface-Based cell neighborhood and interaction volume analysis in 3D spatial omics.
|
|
6
|
+
|
|
7
|
+

|
|
8
|
+
|
|
9
|
+
## Installation
|
|
10
|
+
|
|
11
|
+
**Install package:**
|
|
12
|
+
```sh
|
|
13
|
+
pip install interscellar
|
|
14
|
+
```
|
|
15
|
+
|
|
16
|
+
## Usage
|
|
17
|
+
|
|
18
|
+
**Import:**
|
|
19
|
+
```sh
|
|
20
|
+
import interscellar
|
|
21
|
+
```
|
|
22
|
+
|
|
23
|
+
### 3D Pipeline:
|
|
24
|
+
|
|
25
|
+
**(1) Cell Neighbor Detection & Graph Construction**
|
|
26
|
+
```sh
|
|
27
|
+
neighbors_3d, adata, conn = interscellar.find_cell_neighbors_3d(
|
|
28
|
+
ome_zarr_path="data/segmentation.zarr",
|
|
29
|
+
metadata_csv_path="data/cell_metadata.csv",
|
|
30
|
+
max_distance_um=0.5,
|
|
31
|
+
voxel_size_um=(0.56, 0.28, 0.28),
|
|
32
|
+
n_jobs=4
|
|
33
|
+
)
|
|
34
|
+
```
|
|
35
|
+
|
|
36
|
+
**(2) Interscellar Volume Computation**
|
|
37
|
+
```sh
|
|
38
|
+
# Interscellar volumes
|
|
39
|
+
volumes_3d, adata, conn = interscellar.compute_interscellar_volumes_3d(
|
|
40
|
+
ome_zarr_path="data/segmentation.zarr",
|
|
41
|
+
neighbor_pairs_csv="results/neighbors_3d.csv",
|
|
42
|
+
neighbor_db_path="/results/neighbor_graph.db",
|
|
43
|
+
voxel_size_um=(0.56, 0.28, 0.28),
|
|
44
|
+
max_distance_um=3.0,
|
|
45
|
+
intracellular_threshold_um=1.0,
|
|
46
|
+
n_jobs=4
|
|
47
|
+
)
|
|
48
|
+
```
|
|
49
|
+
|
|
50
|
+
```sh
|
|
51
|
+
# Cell-only volumes
|
|
52
|
+
cellonly_3d = interscellar.compute_cell_only_volumes_3d(
|
|
53
|
+
ome_zarr_path="data/segmentation.zarr",
|
|
54
|
+
interscellar_volumes_zarr="results/interscellar_volumes.zarr"
|
|
55
|
+
)
|
|
56
|
+
```
|
|
57
|
+
|
|
58
|
+
### 2D Pipeline:
|
|
59
|
+
|
|
60
|
+
**(1) Cell Neighbor Detection & Graph Construction**
|
|
61
|
+
```sh
|
|
62
|
+
neighbors_2d, adata, conn = interscellar.find_cell_neighbors_2d(
|
|
63
|
+
polygon_json_path="data/cell_polygons.json",
|
|
64
|
+
metadata_csv_path="data/cell_metadata.csv",
|
|
65
|
+
max_distance_um=1.0,
|
|
66
|
+
pixel_size_um=0.1085,
|
|
67
|
+
n_jobs=4
|
|
68
|
+
)
|
|
69
|
+
```
|
|
70
|
+
|
|
71
|
+
### Utilities:
|
|
72
|
+
|
|
73
|
+
**Volume Visualization**
|
|
74
|
+
```sh
|
|
75
|
+
# Full dataset (Napari)
|
|
76
|
+
visualize-all-3d \
|
|
77
|
+
--cell-only-zarr "results/cell_only_volumes.zarr" \
|
|
78
|
+
--interscellar-zarr "results/interscellar_volumes.zarr" \
|
|
79
|
+
--cell-only-opacity 0.7 \
|
|
80
|
+
--interscellar-opacity 0.9
|
|
81
|
+
```
|
|
82
|
+
|
|
83
|
+
```sh
|
|
84
|
+
# Single pair (Napari)
|
|
85
|
+
visualize-pair-3d \
|
|
86
|
+
--pair-id 123 \
|
|
87
|
+
--cell-only-zarr "results/cell_only_volumes.zarr" \
|
|
88
|
+
--interscellar-zarr "results/interscellar_volumes.zarr" \
|
|
89
|
+
--pair-opacity 0.6 \
|
|
90
|
+
--cells-opacity 0.7
|
|
91
|
+
```
|
|
@@ -0,0 +1,86 @@
|
|
|
1
|
+
[build-system]
|
|
2
|
+
requires = ["setuptools>=61.0", "wheel"]
|
|
3
|
+
build-backend = "setuptools.build_meta"
|
|
4
|
+
|
|
5
|
+
[project]
|
|
6
|
+
name = "InterSCellar"
|
|
7
|
+
version = "0.1.0"
|
|
8
|
+
description = "Surface-based Cell Neighbor Detection and Interscellar Volume Computation for 2D & 3D Spatial Omics"
|
|
9
|
+
readme = "README.md"
|
|
10
|
+
requires-python = ">=3.8"
|
|
11
|
+
license = {text = "MIT"}
|
|
12
|
+
authors = [
|
|
13
|
+
{name = "Eunice Yewon Lee", email = "yl888@duke.edu"}
|
|
14
|
+
]
|
|
15
|
+
keywords = [
|
|
16
|
+
"spatial-omics",
|
|
17
|
+
"cell-neighbors",
|
|
18
|
+
"cell-interactions",
|
|
19
|
+
"cell-cell-communication",
|
|
20
|
+
"intercellular",
|
|
21
|
+
"single-cell",
|
|
22
|
+
"transcriptomics",
|
|
23
|
+
"proteomics",
|
|
24
|
+
"metabolomics",
|
|
25
|
+
"bioinformatics",
|
|
26
|
+
"computational-biology",
|
|
27
|
+
"visualization",
|
|
28
|
+
"3d"
|
|
29
|
+
]
|
|
30
|
+
classifiers = [
|
|
31
|
+
"Development Status :: 3 - Alpha",
|
|
32
|
+
"Intended Audience :: Science/Research",
|
|
33
|
+
"License :: OSI Approved :: MIT License",
|
|
34
|
+
"Operating System :: OS Independent",
|
|
35
|
+
"Programming Language :: Python :: 3",
|
|
36
|
+
"Programming Language :: Python :: 3.8",
|
|
37
|
+
"Programming Language :: Python :: 3.9",
|
|
38
|
+
"Programming Language :: Python :: 3.10",
|
|
39
|
+
"Programming Language :: Python :: 3.11",
|
|
40
|
+
"Topic :: Scientific/Engineering :: Bio-Informatics",
|
|
41
|
+
]
|
|
42
|
+
|
|
43
|
+
dependencies = [
|
|
44
|
+
"numpy>=1.20.0",
|
|
45
|
+
"pandas>=1.3.0",
|
|
46
|
+
"scipy>=1.7.0",
|
|
47
|
+
"tqdm>=4.62.0",
|
|
48
|
+
"opencv-python>=4.5.0",
|
|
49
|
+
"dask[array]>=2021.0.0",
|
|
50
|
+
"matplotlib>=3.4.0",
|
|
51
|
+
"scikit-image>=0.18.0",
|
|
52
|
+
"seaborn>=0.11.0",
|
|
53
|
+
"napari>=0.4.0",
|
|
54
|
+
"zarr>=2.10.0",
|
|
55
|
+
"anndata>=0.8.0",
|
|
56
|
+
"duckdb>=0.6.0",
|
|
57
|
+
]
|
|
58
|
+
|
|
59
|
+
[project.optional-dependencies]
|
|
60
|
+
anndata = [
|
|
61
|
+
"anndata>=0.8.0",
|
|
62
|
+
]
|
|
63
|
+
duckdb = [
|
|
64
|
+
"duckdb>=0.6.0",
|
|
65
|
+
]
|
|
66
|
+
all = [
|
|
67
|
+
"anndata>=0.8.0",
|
|
68
|
+
"duckdb>=0.6.0",
|
|
69
|
+
]
|
|
70
|
+
|
|
71
|
+
[project.scripts]
|
|
72
|
+
visualize-all-3d = "interscellar.visualization.visualize_all_3d:main"
|
|
73
|
+
visualize-pair-3d = "interscellar.visualization.visualize_pair_3d:main"
|
|
74
|
+
|
|
75
|
+
[project.urls]
|
|
76
|
+
Homepage = "https://github.com/euniceyl/InterSCellar"
|
|
77
|
+
Repository = "https://github.com/euniceyl/InterSCellar"
|
|
78
|
+
Issues = "https://github.com/euniceyl/InterSCellar/issues"
|
|
79
|
+
|
|
80
|
+
[tool.setuptools]
|
|
81
|
+
package-dir = {"interscellar" = "src"}
|
|
82
|
+
packages = ["interscellar", "interscellar.api", "interscellar.core", "interscellar.visualization"]
|
|
83
|
+
|
|
84
|
+
[tool.setuptools.package-data]
|
|
85
|
+
"*" = ["*.py", "*.md", "LICENSE"]
|
|
86
|
+
|
|
@@ -0,0 +1,22 @@
|
|
|
1
|
+
__version__ = "0.1.0"
|
|
2
|
+
|
|
3
|
+
from .api import (
|
|
4
|
+
find_cell_neighbors_2d,
|
|
5
|
+
find_cell_neighbors_3d,
|
|
6
|
+
compute_interscellar_volumes_3d,
|
|
7
|
+
compute_cell_only_volumes_3d
|
|
8
|
+
)
|
|
9
|
+
|
|
10
|
+
from .visualization import (
|
|
11
|
+
visualize_all_3d,
|
|
12
|
+
visualize_pair_3d
|
|
13
|
+
)
|
|
14
|
+
|
|
15
|
+
__all__ = [
|
|
16
|
+
"find_cell_neighbors_2d",
|
|
17
|
+
"find_cell_neighbors_3d",
|
|
18
|
+
"compute_interscellar_volumes_3d",
|
|
19
|
+
"compute_cell_only_volumes_3d",
|
|
20
|
+
"visualize_all_3d",
|
|
21
|
+
"visualize_pair_3d",
|
|
22
|
+
]
|
|
@@ -0,0 +1,14 @@
|
|
|
1
|
+
from .wrapper_2d import find_cell_neighbors_2d
|
|
2
|
+
from .wrapper_3d import (
|
|
3
|
+
find_cell_neighbors_3d,
|
|
4
|
+
compute_interscellar_volumes_3d,
|
|
5
|
+
compute_cell_only_volumes_3d
|
|
6
|
+
)
|
|
7
|
+
|
|
8
|
+
__all__ = [
|
|
9
|
+
"find_cell_neighbors_2d",
|
|
10
|
+
"find_cell_neighbors_3d",
|
|
11
|
+
"compute_interscellar_volumes_3d",
|
|
12
|
+
"compute_cell_only_volumes_3d",
|
|
13
|
+
]
|
|
14
|
+
|