CausalPy 0.2.2__tar.gz → 0.3.0__tar.gz
This diff represents the content of publicly available package versions that have been released to one of the supported registries. The information contained in this diff is provided for informational purposes only and reflects changes between package versions as they appear in their respective public registries.
- {CausalPy-0.2.2 → causalpy-0.3.0}/CausalPy.egg-info/PKG-INFO +17 -5
- {CausalPy-0.2.2 → causalpy-0.3.0}/CausalPy.egg-info/SOURCES.txt +1 -0
- {CausalPy-0.2.2 → causalpy-0.3.0}/CausalPy.egg-info/requires.txt +3 -2
- {CausalPy-0.2.2 → causalpy-0.3.0}/PKG-INFO +17 -5
- {CausalPy-0.2.2 → causalpy-0.3.0}/README.md +12 -1
- causalpy-0.3.0/causalpy/__init__.py +30 -0
- {CausalPy-0.2.2 → causalpy-0.3.0}/causalpy/custom_exceptions.py +13 -0
- causalpy-0.3.0/causalpy/data/__init__.py +18 -0
- {CausalPy-0.2.2 → causalpy-0.3.0}/causalpy/data/datasets.py +15 -0
- causalpy-0.3.0/causalpy/data/nhefs.csv +1567 -0
- {CausalPy-0.2.2 → causalpy-0.3.0}/causalpy/data/simulate_data.py +14 -0
- {CausalPy-0.2.2 → causalpy-0.3.0}/causalpy/data_validation.py +39 -0
- {CausalPy-0.2.2 → causalpy-0.3.0}/causalpy/plot_utils.py +13 -0
- {CausalPy-0.2.2 → causalpy-0.3.0}/causalpy/pymc_experiments.py +459 -2
- {CausalPy-0.2.2 → causalpy-0.3.0}/causalpy/pymc_models.py +127 -21
- {CausalPy-0.2.2 → causalpy-0.3.0}/causalpy/skl_experiments.py +18 -1
- {CausalPy-0.2.2 → causalpy-0.3.0}/causalpy/skl_models.py +14 -0
- {CausalPy-0.2.2 → causalpy-0.3.0}/causalpy/utils.py +14 -0
- causalpy-0.3.0/causalpy/version.py +16 -0
- {CausalPy-0.2.2 → causalpy-0.3.0}/pyproject.toml +6 -5
- CausalPy-0.2.2/causalpy/__init__.py +0 -17
- CausalPy-0.2.2/causalpy/data/__init__.py +0 -4
- CausalPy-0.2.2/causalpy/version.py +0 -2
- {CausalPy-0.2.2 → causalpy-0.3.0}/CausalPy.egg-info/dependency_links.txt +0 -0
- {CausalPy-0.2.2 → causalpy-0.3.0}/CausalPy.egg-info/top_level.txt +0 -0
- {CausalPy-0.2.2 → causalpy-0.3.0}/LICENSE +0 -0
- {CausalPy-0.2.2 → causalpy-0.3.0}/causalpy/data/AJR2001.csv +0 -0
- {CausalPy-0.2.2 → causalpy-0.3.0}/causalpy/data/GDP_in_dollars_billions.csv +0 -0
- {CausalPy-0.2.2 → causalpy-0.3.0}/causalpy/data/ancova_generated.csv +0 -0
- {CausalPy-0.2.2 → causalpy-0.3.0}/causalpy/data/banks.csv +0 -0
- {CausalPy-0.2.2 → causalpy-0.3.0}/causalpy/data/deaths_and_temps_england_wales.csv +0 -0
- {CausalPy-0.2.2 → causalpy-0.3.0}/causalpy/data/did.csv +0 -0
- {CausalPy-0.2.2 → causalpy-0.3.0}/causalpy/data/drinking.csv +0 -0
- {CausalPy-0.2.2 → causalpy-0.3.0}/causalpy/data/geolift1.csv +0 -0
- {CausalPy-0.2.2 → causalpy-0.3.0}/causalpy/data/its.csv +0 -0
- {CausalPy-0.2.2 → causalpy-0.3.0}/causalpy/data/its_simple.csv +0 -0
- {CausalPy-0.2.2 → causalpy-0.3.0}/causalpy/data/regression_discontinuity.csv +0 -0
- {CausalPy-0.2.2 → causalpy-0.3.0}/causalpy/data/synthetic_control.csv +0 -0
- {CausalPy-0.2.2 → causalpy-0.3.0}/setup.cfg +0 -0
|
@@ -1,6 +1,6 @@
|
|
|
1
1
|
Metadata-Version: 2.1
|
|
2
2
|
Name: CausalPy
|
|
3
|
-
Version: 0.
|
|
3
|
+
Version: 0.3.0
|
|
4
4
|
Summary: Causal inference for quasi-experiments in Python
|
|
5
5
|
Author-email: Ben Vincent <ben.vincent@pymc-labs.com>
|
|
6
6
|
License: Apache License
|
|
@@ -208,7 +208,7 @@ License: Apache License
|
|
|
208
208
|
Project-URL: Homepage, https://github.com/pymc-labs/CausalPy
|
|
209
209
|
Project-URL: Bug Reports, https://github.com/pymc-labs/CausalPy/issues
|
|
210
210
|
Project-URL: Source, https://github.com/pymc-labs/CausalPy
|
|
211
|
-
Requires-Python: >=3.
|
|
211
|
+
Requires-Python: >=3.10
|
|
212
212
|
Description-Content-Type: text/markdown
|
|
213
213
|
License-File: LICENSE
|
|
214
214
|
Requires-Dist: arviz>=0.14.0
|
|
@@ -218,7 +218,7 @@ Requires-Dist: matplotlib>=3.5.3
|
|
|
218
218
|
Requires-Dist: numpy<1.26.0
|
|
219
219
|
Requires-Dist: pandas
|
|
220
220
|
Requires-Dist: patsy
|
|
221
|
-
Requires-Dist: pymc>=5.
|
|
221
|
+
Requires-Dist: pymc>=5.14.0
|
|
222
222
|
Requires-Dist: scikit-learn>=1
|
|
223
223
|
Requires-Dist: scipy
|
|
224
224
|
Requires-Dist: seaborn>=0.11.2
|
|
@@ -231,8 +231,9 @@ Requires-Dist: twine; extra == "dev"
|
|
|
231
231
|
Requires-Dist: interrogate; extra == "dev"
|
|
232
232
|
Provides-Extra: docs
|
|
233
233
|
Requires-Dist: ipykernel; extra == "docs"
|
|
234
|
+
Requires-Dist: daft; extra == "docs"
|
|
234
235
|
Requires-Dist: linkify-it-py; extra == "docs"
|
|
235
|
-
Requires-Dist: myst-nb; extra == "docs"
|
|
236
|
+
Requires-Dist: myst-nb<=1.0.0; extra == "docs"
|
|
236
237
|
Requires-Dist: pathlib; extra == "docs"
|
|
237
238
|
Requires-Dist: sphinx; extra == "docs"
|
|
238
239
|
Requires-Dist: sphinx-autodoc-typehints; extra == "docs"
|
|
@@ -430,7 +431,7 @@ Interrupted time series analysis is appropriate when you have a time series of o
|
|
|
430
431
|
|
|
431
432
|
| Frequentist | Bayesian |
|
|
432
433
|
|--|--|
|
|
433
|
-
|
|
|
434
|
+
|  |  |
|
|
434
435
|
|
|
435
436
|
> The data, pre-treatment model fit, and counterfactual are plotted (top). The causal impact is shown as a blue shaded region. The Bayesian analysis shows shaded Bayesian credible regions of the model fit and counterfactual. Also shown is the causal impact (middle) and cumulative causal impact (bottom).
|
|
436
437
|
|
|
@@ -440,6 +441,17 @@ Instrumental Variable regression is an appropriate technique when you wish to es
|
|
|
440
441
|
|
|
441
442
|
|
|
442
443
|
|
|
444
|
+
|
|
445
|
+
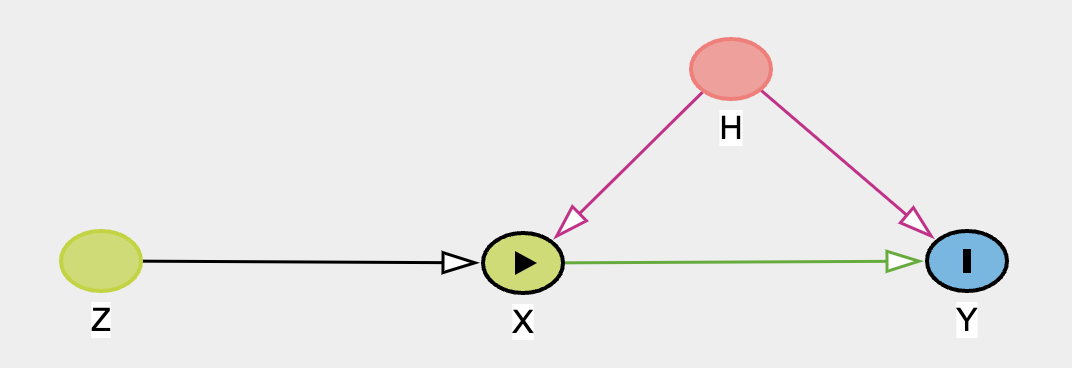
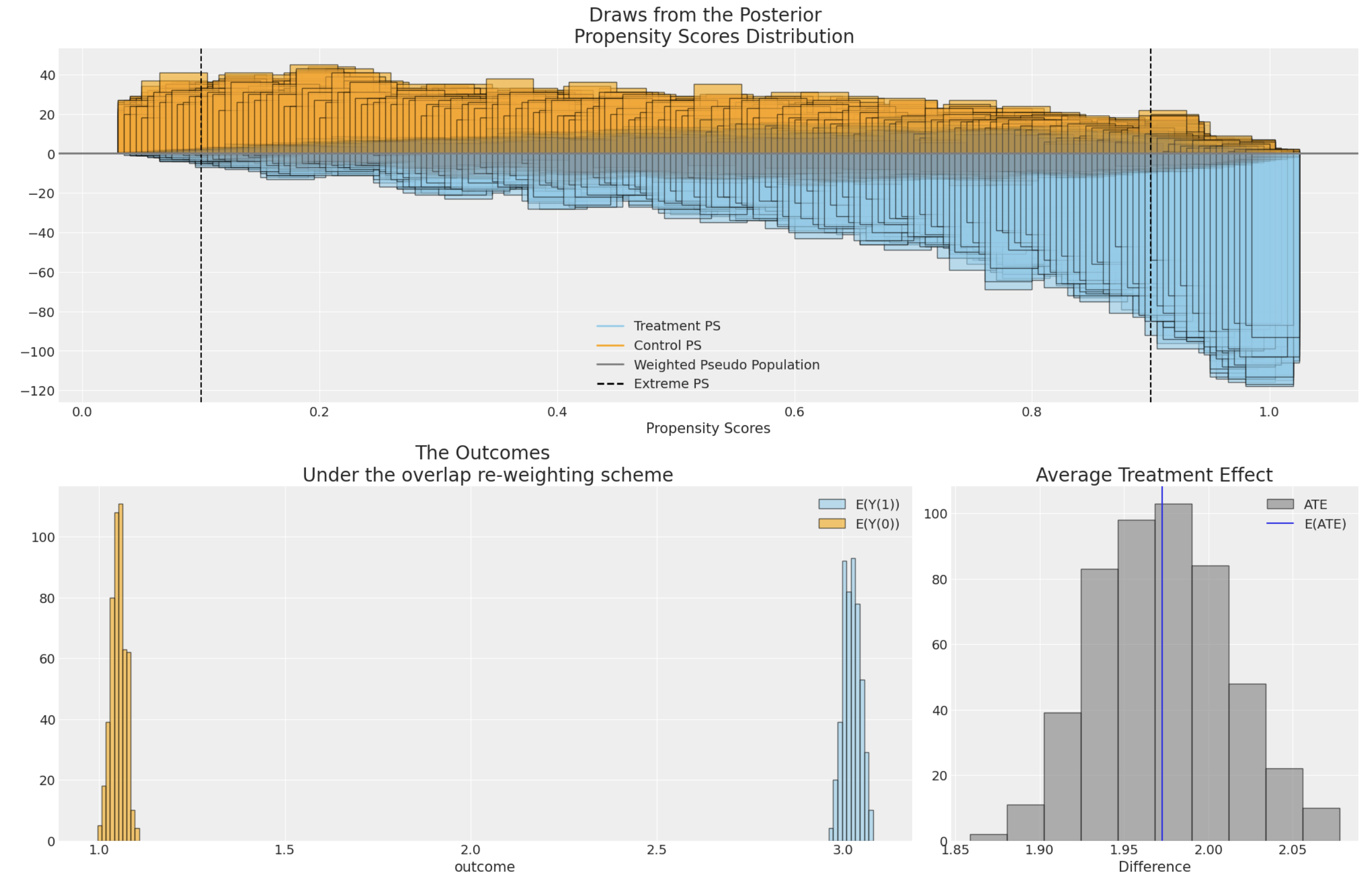
### Inverse Propensity Score Weighting
|
|
446
|
+
|
|
447
|
+
Propensity scores are often used to address the risks of bias or confounding introduced in an observational study by
|
|
448
|
+
selection effects into the treatment condition. Propensity scores can be used in a number of ways, but here we demonstrate
|
|
449
|
+
their usage within corrective weighting schemes aimed to recover as-if random allocation of subjects to the treatment condition.
|
|
450
|
+
The technique "up-weights" or "down-weights" individual observations to better estimate a causal estimand such as the average treatment
|
|
451
|
+
effect.
|
|
452
|
+
|
|
453
|
+
|
|
454
|
+
|
|
443
455
|
## Learning resources
|
|
444
456
|
|
|
445
457
|
Here are some general resources about causal inference:
|
|
@@ -5,7 +5,7 @@ matplotlib>=3.5.3
|
|
|
5
5
|
numpy<1.26.0
|
|
6
6
|
pandas
|
|
7
7
|
patsy
|
|
8
|
-
pymc>=5.
|
|
8
|
+
pymc>=5.14.0
|
|
9
9
|
scikit-learn>=1
|
|
10
10
|
scipy
|
|
11
11
|
seaborn>=0.11.2
|
|
@@ -20,8 +20,9 @@ interrogate
|
|
|
20
20
|
|
|
21
21
|
[docs]
|
|
22
22
|
ipykernel
|
|
23
|
+
daft
|
|
23
24
|
linkify-it-py
|
|
24
|
-
myst-nb
|
|
25
|
+
myst-nb<=1.0.0
|
|
25
26
|
pathlib
|
|
26
27
|
sphinx
|
|
27
28
|
sphinx-autodoc-typehints
|
|
@@ -1,6 +1,6 @@
|
|
|
1
1
|
Metadata-Version: 2.1
|
|
2
2
|
Name: CausalPy
|
|
3
|
-
Version: 0.
|
|
3
|
+
Version: 0.3.0
|
|
4
4
|
Summary: Causal inference for quasi-experiments in Python
|
|
5
5
|
Author-email: Ben Vincent <ben.vincent@pymc-labs.com>
|
|
6
6
|
License: Apache License
|
|
@@ -208,7 +208,7 @@ License: Apache License
|
|
|
208
208
|
Project-URL: Homepage, https://github.com/pymc-labs/CausalPy
|
|
209
209
|
Project-URL: Bug Reports, https://github.com/pymc-labs/CausalPy/issues
|
|
210
210
|
Project-URL: Source, https://github.com/pymc-labs/CausalPy
|
|
211
|
-
Requires-Python: >=3.
|
|
211
|
+
Requires-Python: >=3.10
|
|
212
212
|
Description-Content-Type: text/markdown
|
|
213
213
|
License-File: LICENSE
|
|
214
214
|
Requires-Dist: arviz>=0.14.0
|
|
@@ -218,7 +218,7 @@ Requires-Dist: matplotlib>=3.5.3
|
|
|
218
218
|
Requires-Dist: numpy<1.26.0
|
|
219
219
|
Requires-Dist: pandas
|
|
220
220
|
Requires-Dist: patsy
|
|
221
|
-
Requires-Dist: pymc>=5.
|
|
221
|
+
Requires-Dist: pymc>=5.14.0
|
|
222
222
|
Requires-Dist: scikit-learn>=1
|
|
223
223
|
Requires-Dist: scipy
|
|
224
224
|
Requires-Dist: seaborn>=0.11.2
|
|
@@ -231,8 +231,9 @@ Requires-Dist: twine; extra == "dev"
|
|
|
231
231
|
Requires-Dist: interrogate; extra == "dev"
|
|
232
232
|
Provides-Extra: docs
|
|
233
233
|
Requires-Dist: ipykernel; extra == "docs"
|
|
234
|
+
Requires-Dist: daft; extra == "docs"
|
|
234
235
|
Requires-Dist: linkify-it-py; extra == "docs"
|
|
235
|
-
Requires-Dist: myst-nb; extra == "docs"
|
|
236
|
+
Requires-Dist: myst-nb<=1.0.0; extra == "docs"
|
|
236
237
|
Requires-Dist: pathlib; extra == "docs"
|
|
237
238
|
Requires-Dist: sphinx; extra == "docs"
|
|
238
239
|
Requires-Dist: sphinx-autodoc-typehints; extra == "docs"
|
|
@@ -430,7 +431,7 @@ Interrupted time series analysis is appropriate when you have a time series of o
|
|
|
430
431
|
|
|
431
432
|
| Frequentist | Bayesian |
|
|
432
433
|
|--|--|
|
|
433
|
-
|
|
|
434
|
+
|  |  |
|
|
434
435
|
|
|
435
436
|
> The data, pre-treatment model fit, and counterfactual are plotted (top). The causal impact is shown as a blue shaded region. The Bayesian analysis shows shaded Bayesian credible regions of the model fit and counterfactual. Also shown is the causal impact (middle) and cumulative causal impact (bottom).
|
|
436
437
|
|
|
@@ -440,6 +441,17 @@ Instrumental Variable regression is an appropriate technique when you wish to es
|
|
|
440
441
|
|
|
441
442
|

|
|
442
443
|
|
|
444
|
+
|
|
445
|
+
### Inverse Propensity Score Weighting
|
|
446
|
+
|
|
447
|
+
Propensity scores are often used to address the risks of bias or confounding introduced in an observational study by
|
|
448
|
+
selection effects into the treatment condition. Propensity scores can be used in a number of ways, but here we demonstrate
|
|
449
|
+
their usage within corrective weighting schemes aimed to recover as-if random allocation of subjects to the treatment condition.
|
|
450
|
+
The technique "up-weights" or "down-weights" individual observations to better estimate a causal estimand such as the average treatment
|
|
451
|
+
effect.
|
|
452
|
+
|
|
453
|
+

|
|
454
|
+
|
|
443
455
|
## Learning resources
|
|
444
456
|
|
|
445
457
|
Here are some general resources about causal inference:
|
|
@@ -178,7 +178,7 @@ Interrupted time series analysis is appropriate when you have a time series of o
|
|
|
178
178
|
|
|
179
179
|
| Frequentist | Bayesian |
|
|
180
180
|
|--|--|
|
|
181
|
-
|
|
|
181
|
+
|  |  |
|
|
182
182
|
|
|
183
183
|
> The data, pre-treatment model fit, and counterfactual are plotted (top). The causal impact is shown as a blue shaded region. The Bayesian analysis shows shaded Bayesian credible regions of the model fit and counterfactual. Also shown is the causal impact (middle) and cumulative causal impact (bottom).
|
|
184
184
|
|
|
@@ -188,6 +188,17 @@ Instrumental Variable regression is an appropriate technique when you wish to es
|
|
|
188
188
|
|
|
189
189
|

|
|
190
190
|
|
|
191
|
+
|
|
192
|
+
### Inverse Propensity Score Weighting
|
|
193
|
+
|
|
194
|
+
Propensity scores are often used to address the risks of bias or confounding introduced in an observational study by
|
|
195
|
+
selection effects into the treatment condition. Propensity scores can be used in a number of ways, but here we demonstrate
|
|
196
|
+
their usage within corrective weighting schemes aimed to recover as-if random allocation of subjects to the treatment condition.
|
|
197
|
+
The technique "up-weights" or "down-weights" individual observations to better estimate a causal estimand such as the average treatment
|
|
198
|
+
effect.
|
|
199
|
+
|
|
200
|
+

|
|
201
|
+
|
|
191
202
|
## Learning resources
|
|
192
203
|
|
|
193
204
|
Here are some general resources about causal inference:
|
|
@@ -0,0 +1,30 @@
|
|
|
1
|
+
# Copyright 2024 The PyMC Labs Developers
|
|
2
|
+
#
|
|
3
|
+
# Licensed under the Apache License, Version 2.0 (the "License");
|
|
4
|
+
# you may not use this file except in compliance with the License.
|
|
5
|
+
# You may obtain a copy of the License at
|
|
6
|
+
#
|
|
7
|
+
# http://www.apache.org/licenses/LICENSE-2.0
|
|
8
|
+
#
|
|
9
|
+
# Unless required by applicable law or agreed to in writing, software
|
|
10
|
+
# distributed under the License is distributed on an "AS IS" BASIS,
|
|
11
|
+
# WITHOUT WARRANTIES OR CONDITIONS OF ANY KIND, either express or implied.
|
|
12
|
+
# See the License for the specific language governing permissions and
|
|
13
|
+
# limitations under the License.
|
|
14
|
+
import arviz as az
|
|
15
|
+
|
|
16
|
+
from causalpy import pymc_experiments, pymc_models, skl_experiments, skl_models
|
|
17
|
+
from causalpy.version import __version__
|
|
18
|
+
|
|
19
|
+
from .data import load_data
|
|
20
|
+
|
|
21
|
+
az.style.use("arviz-darkgrid")
|
|
22
|
+
|
|
23
|
+
__all__ = [
|
|
24
|
+
"pymc_experiments",
|
|
25
|
+
"pymc_models",
|
|
26
|
+
"skl_experiments",
|
|
27
|
+
"skl_models",
|
|
28
|
+
"load_data",
|
|
29
|
+
"__version__",
|
|
30
|
+
]
|
|
@@ -1,3 +1,16 @@
|
|
|
1
|
+
# Copyright 2024 The PyMC Labs Developers
|
|
2
|
+
#
|
|
3
|
+
# Licensed under the Apache License, Version 2.0 (the "License");
|
|
4
|
+
# you may not use this file except in compliance with the License.
|
|
5
|
+
# You may obtain a copy of the License at
|
|
6
|
+
#
|
|
7
|
+
# http://www.apache.org/licenses/LICENSE-2.0
|
|
8
|
+
#
|
|
9
|
+
# Unless required by applicable law or agreed to in writing, software
|
|
10
|
+
# distributed under the License is distributed on an "AS IS" BASIS,
|
|
11
|
+
# WITHOUT WARRANTIES OR CONDITIONS OF ANY KIND, either express or implied.
|
|
12
|
+
# See the License for the specific language governing permissions and
|
|
13
|
+
# limitations under the License.
|
|
1
14
|
"""
|
|
2
15
|
Custom Exceptions for CausalPy.
|
|
3
16
|
"""
|
|
@@ -0,0 +1,18 @@
|
|
|
1
|
+
# Copyright 2024 The PyMC Labs Developers
|
|
2
|
+
#
|
|
3
|
+
# Licensed under the Apache License, Version 2.0 (the "License");
|
|
4
|
+
# you may not use this file except in compliance with the License.
|
|
5
|
+
# You may obtain a copy of the License at
|
|
6
|
+
#
|
|
7
|
+
# http://www.apache.org/licenses/LICENSE-2.0
|
|
8
|
+
#
|
|
9
|
+
# Unless required by applicable law or agreed to in writing, software
|
|
10
|
+
# distributed under the License is distributed on an "AS IS" BASIS,
|
|
11
|
+
# WITHOUT WARRANTIES OR CONDITIONS OF ANY KIND, either express or implied.
|
|
12
|
+
# See the License for the specific language governing permissions and
|
|
13
|
+
# limitations under the License.
|
|
14
|
+
"""Code for loading datasets."""
|
|
15
|
+
|
|
16
|
+
from .datasets import load_data
|
|
17
|
+
|
|
18
|
+
__all__ = ["load_data"]
|
|
@@ -1,6 +1,20 @@
|
|
|
1
|
+
# Copyright 2024 The PyMC Labs Developers
|
|
2
|
+
#
|
|
3
|
+
# Licensed under the Apache License, Version 2.0 (the "License");
|
|
4
|
+
# you may not use this file except in compliance with the License.
|
|
5
|
+
# You may obtain a copy of the License at
|
|
6
|
+
#
|
|
7
|
+
# http://www.apache.org/licenses/LICENSE-2.0
|
|
8
|
+
#
|
|
9
|
+
# Unless required by applicable law or agreed to in writing, software
|
|
10
|
+
# distributed under the License is distributed on an "AS IS" BASIS,
|
|
11
|
+
# WITHOUT WARRANTIES OR CONDITIONS OF ANY KIND, either express or implied.
|
|
12
|
+
# See the License for the specific language governing permissions and
|
|
13
|
+
# limitations under the License.
|
|
1
14
|
"""
|
|
2
15
|
Functions to load example datasets
|
|
3
16
|
"""
|
|
17
|
+
|
|
4
18
|
import pathlib
|
|
5
19
|
|
|
6
20
|
import pandas as pd
|
|
@@ -20,6 +34,7 @@ DATASETS = {
|
|
|
20
34
|
"anova1": {"filename": "ancova_generated.csv"},
|
|
21
35
|
"geolift1": {"filename": "geolift1.csv"},
|
|
22
36
|
"risk": {"filename": "AJR2001.csv"},
|
|
37
|
+
"nhefs": {"filename": "nhefs.csv"},
|
|
23
38
|
}
|
|
24
39
|
|
|
25
40
|
|